(FAST) Is a Regulator of Alternative Splicing
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Large-Scale Analysis of Genome and Transcriptome Alterations in Multiple Tumors Unveils Novel Cancer-Relevant Splicing Networks
Downloaded from genome.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press Research Large-scale analysis of genome and transcriptome alterations in multiple tumors unveils novel cancer-relevant splicing networks Endre Sebestyén,1,5 Babita Singh,1,5 Belén Miñana,1,2 Amadís Pagès,1 Francesca Mateo,3 Miguel Angel Pujana,3 Juan Valcárcel,1,2,4 and Eduardo Eyras1,4 1Universitat Pompeu Fabra, E08003 Barcelona, Spain; 2Centre for Genomic Regulation, E08003 Barcelona, Spain; 3Program Against Cancer Therapeutic Resistance (ProCURE), Catalan Institute of Oncology (ICO), Bellvitge Institute for Biomedical Research (IDIBELL), E08908 L’Hospitalet del Llobregat, Spain; 4Catalan Institution for Research and Advanced Studies, E08010 Barcelona, Spain Alternative splicing is regulated by multiple RNA-binding proteins and influences the expression of most eukaryotic genes. However, the role of this process in human disease, and particularly in cancer, is only starting to be unveiled. We system- atically analyzed mutation, copy number, and gene expression patterns of 1348 RNA-binding protein (RBP) genes in 11 solid tumor types, together with alternative splicing changes in these tumors and the enrichment of binding motifs in the alter- natively spliced sequences. Our comprehensive study reveals widespread alterations in the expression of RBP genes, as well as novel mutations and copy number variations in association with multiple alternative splicing changes in cancer drivers and oncogenic pathways. Remarkably, the altered splicing patterns in several tumor types recapitulate those of undifferen- tiated cells. These patterns are predicted to be mainly controlled by MBNL1 and involve multiple cancer drivers, including the mitotic gene NUMA1. We show that NUMA1 alternative splicing induces enhanced cell proliferation and centrosome am- plification in nontumorigenic mammary epithelial cells. -

Overlap of Quantitative Trait Loci for Early Growth Rate, and for Body
REPRODUCTIONRESEARCH Functional reconstruction of NANOS3 expression in the germ cell lineage by a novel transgenic reporter reveals distinct subcellular localizations of NANOS3 Masashi Yamaji1, Takashi Tanaka2, Mayo Shigeta1, Shinichiro Chuma2, Yumiko Saga3 and Mitinori Saitou1,4,5 1Laboratory for Mammalian Germ Cell Biology, RIKEN Center for Developmental Biology, 2-2-3 Minatojima- Minamimachi, Chuo-ku, Kobe 650-0047, Japan, 2Department of Development and Differentiation, Institute for Frontier Medical Sciences, Kyoto University, Shogoin, Sakyo-ku, Kyoto 606-8507, Japan, 3Department of Genetics and Division of Mammalian Development, National Institute of Genetics, SOKENDAI, 1111 Yata, Mishima, Shizuoka 411-8540, Japan, 4Department of Anatomy and Cell Biology, Graduate School of Medicine, Kyoto University, Yoshida- Konoe-cho, Sakyo-ku, Kyoto 606-8501, Japan and 5JST, CREST, Yoshida-Konoe-cho, Sakyo-ku, Kyoto 606-8501, Japan Correspondence should be addressed to M Saitou at Department of Anatomy and Cell Biology, Graduate School of Medicine, Kyoto University; Email: [email protected] Abstract Mutations of RNA-binding proteins such as NANOS3, TIAL1, and DND1 in mice have been known to result in the failure of survival and/or proliferation of primordial germ cells (PGCs) soon after their fate is specified (around embryonic day (E) 8.0), leading to the infertility of these animals. However, the mechanisms of actions of these RNA-binding proteins remain largely unresolved. As a foundation to explore the role of these RNA-binding proteins in germ cells, we established a novel transgenic reporter strain that expresses NANOS3 fused with EGFP under the control of Nanos3 regulatory elements. NANOS3–EGFP exhibited exclusive expression in PGCs as early as E7.25, and continued to be expressed in female germ cells until around E14.5 and in male germ cells throughout the fetal period with declining expression levels after E16.5. -

Effects of Changes in TIA1 on Stress Granule Formation
Effects of changes in TIA1 on stress granule formation Master’s thesis Lydia Sagath University of Helsinki Faculty of Biological and Environmental Sciences Genetics August 2015 Tiedekunta – Fakultet – Faculty Laitos – Institution– Department Bio- och miljövetenskapliga fakulteten Biovetenskapliga institutionen Tekijä – Författare – Author Lydia Johanna Sagath Työn nimi – Arbetets titel – Title Effects of changes in TIA1 on stress granule formation Oppiaine – Läroämne – Subject Humangenetik Työn laji – Arbetets art – Level Aika – Datum – Month and year Sivumäärä – Sidoantal – Number of pages Pro gradu Augusti 2015 63 Tiivistelmä – Referat – Abstract Welanders distala myopati (WDM) orsakas av mutationen p.E384K i genen TIA1. Mutationen antas vara sjukdomsalstrande på grund av en ökad produktion av protein, som relaterats till formationen av stressgranuler (Hackman et al. 2013). Även omgivningsfaktorer har föreslagits verka i sjukdomens utveckling: en ökad mängd stressgranuler har observerats i celler som behandlats med köldshock jämfört med celler som förvarats i 37°C (Hofmann et al. 2012). I patienter med WDM-liknande symptom som undersökts för förändringar i TIA1 har en p.N357S-förändring noterats förrikad. Denna förändring har tidigare anmälts som en polymorfism. Förändringen i fråga ligger i samma prionliknande domän i exon 5 som WDM-orsakande förändringen p.E384K. Därmed kunde p.N357S-förändringen öka predispositionen till aggregering. Pro gradu –arbetet är uppdelat i två delar: • p.N357S-polymorfismens effekt på stressgranulsbildningen i arsenitbehandle celler • Köldshockens effekt på stressgranulsbildningen Resultaten påvisar, att förändringen p.N357S i TIA1 orsakar en förändring i det translaterade proteinets beteende. I likhet med p.E384K-förändringen orsakar även p.N357S en ökad mängd stressgranuler i arsenitbehandlade celler. Däremot tyder resultaten på att stressgranulerna återbildas snabbare i fluorescence recovery after photobleaching-studier (FRAP) I p.N357S-transfekterade celler än i celler som transfekterats med TIA1 p.E384K och vildtyp. -

A Comprehensive Analysis of 3' End Sequencing Data Sets Reveals Novel
bioRxiv preprint doi: https://doi.org/10.1101/033001; this version posted November 26, 2015. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. A comprehensive analysis of 3’ end sequencing data sets reveals novel polyadenylation signals and the repressive role of heterogenous ribonucleoprotein C on cleavage and polyadenylation 1 1 1 1 1 Andreas J. Gruber , Ralf Schmidt , Andreas R. Gruber , Georges Martin , Souvik Ghosh , 1,2 1 1,§ Manuel Belmadani , Walter Keller , Mihaela Zavolan 1 Computational and Systems Biology, Biozentrum, University of Basel, Klingelbergstrasse 5070, 4056 Basel, Switzerland 2 Current address: University of British Columbia, 177 Michael Smith Laboratories, 2185 East Mall Vancouver BC V6T 1Z4 § To whom correspondence should be addressed ([email protected]) Abstract Alternative polyadenylation (APA) is a general mechanism of transcript diversification in mammals, which has been recently linked to proliferative states and cancer. Different 3’ untranslated region (3’ UTR) isoforms interact with different RNA binding proteins (RBPs), which modify the stability, translation, and subcellular localization of the corresponding transcripts. Although the heterogeneity of premRNA 3’ end processing has been established with highthroughput approaches, the mechanisms that underlie systematic changes in 3’ UTR lengths remain to be characterized. Through a uniform analysis of a large number of 3’ end sequencing data sets we have uncovered 18 signals, 6 of which novel, whose positioning with respect to premRNA cleavage sites indicates a role in premRNA 3’ end processing in both mouse and human. -

Genetic Variant in 3' Untranslated Region of the Mouse Pycard Gene
bioRxiv preprint doi: https://doi.org/10.1101/2021.03.26.437184; this version posted March 26, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 2 3 Title: 4 Genetic Variant in 3’ Untranslated Region of the Mouse Pycard Gene Regulates Inflammasome 5 Activity 6 Running Title: 7 3’UTR SNP in Pycard regulates inflammasome activity 8 Authors: 9 Brian Ritchey1*, Qimin Hai1*, Juying Han1, John Barnard2, Jonathan D. Smith1,3 10 1Department of Cardiovascular & Metabolic Sciences, Lerner Research Institute, Cleveland Clinic, 11 Cleveland, OH 44195 12 2Department of Quantitative Health Sciences, Lerner Research Institute, Cleveland Clinic, Cleveland, OH 13 44195 14 3Department of Molecular Medicine, Cleveland Clinic Lerner College of Medicine of Case Western 15 Reserve University, Cleveland, OH 44195 16 *, These authors contributed equally to this study. 17 Address correspondence to Jonathan D. Smith: email [email protected]; ORCID ID 0000-0002-0415-386X; 18 mailing address: Cleveland Clinic, Box NC-10, 9500 Euclid Avenue, Cleveland, OH 44195, USA. 19 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.03.26.437184; this version posted March 26, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 20 Abstract 21 Quantitative trait locus mapping for interleukin-1 release after inflammasome priming and activation 22 was performed on bone marrow-derived macrophages (BMDM) from an AKRxDBA/2 strain intercross. -

Dysregulated Expression of Lipid Storage and Membrane Dynamics Factors in Tia1 Knockout Mouse Nervous Tissue
Neurogenetics DOI 10.1007/s10048-014-0397-x ORIGINAL ARTICLE Dysregulated expression of lipid storage and membrane dynamics factors in Tia1 knockout mouse nervous tissue Melanie Vanessa Heck & Mekhman Azizov & Ta n j a S t e h n i n g & Michael Walter & Nancy Kedersha & Georg Auburger Received: 28 January 2014 /Accepted: 3 March 2014 # The Author(s) 2014. This article is published with open access at Springerlink.com Abstract During cell stress, the transcription and translation decreased levels for Bid and Inca1 transcripts. Novel and sur- of immediate early genes are prioritized, while most other mes- prisingly strong expression alterations were detected for fat stor- senger RNAs (mRNAs) are stored away in stress granules or age and membrane trafficking factors, with prominent +3-fold degraded in processing bodies (P-bodies). TIA-1 is an mRNA- upregulations of Plin4, Wdfy1, Tbc1d24,andPnpla2 vs. a −2.4- binding protein that needs to translocate from the nucleus to seed fold downregulation of Cntn4 transcript, encoding an axonal the formation of stress granules in the cytoplasm. Because other membrane adhesion factor with established haploinsufficiency. stress granule components such as TDP-43, FUS, ATXN2, In comparison, subtle effects on the RNA processing machinery SMN, MAPT, HNRNPA2B1, and HNRNPA1 are crucial for included up to 1.2-fold upregulations of Dcp1b and Tial1.The the motor neuron diseases amyotrophic lateral sclerosis (ALS)/ effect on lipid dynamics factors is noteworthy, since also the gene spinal muscular atrophy (SMA) and for the frontotemporal de- deletion of Ta rd b p (encoding TDP-43) and Atxn2 ledtofat mentia (FTD), here we studied mouse nervous tissue to identify metabolism phenotypes in mouse. -
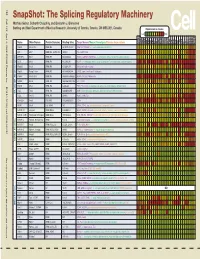
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

TIAL1 (NM 003252) Human Recombinant Protein Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TP760477 TIAL1 (NM_003252) Human Recombinant Protein Product data: Product Type: Recombinant Proteins Description: Purified recombinant protein of Human TIA1 cytotoxic granule-associated RNA binding protein-like 1 (TIAL1), transcript variant 1, full length, with N-terminal HIS tag, expressed in E.Coli, 50ug Species: Human Expression Host: E. coli Tag: N-His Predicted MW: 41.4 kDa Concentration: >50 ug/mL as determined by microplate BCA method Purity: > 80% as determined by SDS-PAGE and Coomassie blue staining Buffer: 25mM Tris, pH8.0, 150 mM NaCl, 10% glycerol, 1 % Sarkosyl. Storage: Store at -80°C. Stability: Stable for 12 months from the date of receipt of the product under proper storage and handling conditions. Avoid repeated freeze-thaw cycles. RefSeq: NP_003243 Locus ID: 7073 UniProt ID: Q01085, Q49AS9 RefSeq Size: 1436 Cytogenetics: 10q26.11 RefSeq ORF: 1125 Synonyms: TCBP; TIAR Summary: The protein encoded by this gene is a member of a family of RNA-binding proteins, has three RNA recognition motifs (RRMs), and binds adenine and uridine-rich elements in mRNA and pre-mRNAs of a wide range of genes. It regulates various activities including translational control, splicing and apoptosis. Alternate transcriptional splice variants, encoding different isoforms, have been characterized. The different isoforms have been show to function differently with respect to post-transcriptional silencing. [provided by RefSeq, Jul 2008] This product is to be used for laboratory only. -

The Devil Is in the Domain: Understanding Protein Recognition of Multiple RNA Targets
The devil is in the domain: understanding protein recognition of multiple RNA targets Glen R. Gronland1 and Andres Ramos1 1. Institute of Structural and Molecular Biology, University College London, London WC1E 6XA, UK Correspondence to Andres Ramos: [email protected] Abstract: RNA regulation provides a finely-tuned and highly-coordinated control of gene expression. Regulation is mediated by hundreds to thousands of multi-functional RNA-binding proteins which often interact with large sets of RNAs. In this brief review, we focus on recent work that highlights how the proteins use multiple RNA-binding domains to interact selectively with the different RNA targets. De-convoluting the molecular complexity of the RNA regulatory network is essential to understanding cell differentiation and function, and requires accurate models for protein-RNA recognition and protein target selectivity. We discuss that the structural and molecular understanding of the key determinant of recognition, together with the availability of methods to examine protein-RNA interactions at the transcriptome level, may provide an avenue to establish these models. Abbreviations list: CSD, Cold shock domain dsRBD, Double-stranded RNA-binding domain KH, hnRNP K-homology mRNA, Messenger RNA ncRNA, Non-coding RNA RBD, RNA-binding domain RBP, RNA-binding protein RRM, RNA-recognition motif ZnF, Zinc finger Introduction The combined regulation of the various steps in the metabolism and transport of messenger RNAs (mRNAs) and non-coding RNAs (ncRNAs) multiplies genomic potential, and allows cellular differentiation and the development of complex organisms. In the cell, functional RNAs are associated with a fluctuating assortment of multi-domain RNA-binding proteins (RBPs), forming integrated complexes, whose composition directs the fate of the transcript1,2. -

Anti-TIA1 Antibody (ARG58772)
Product datasheet [email protected] ARG58772 Package: 100 μl anti-TIA1 antibody Store at: -20°C Summary Product Description Rabbit Polyclonal antibody recognizes TIA1 Tested Reactivity Hu, Ms Tested Application IHC-P, WB Host Rabbit Clonality Polyclonal Isotype IgG Target Name TIA1 Antigen Species Human Immunogen Synthetic peptide from Human TIA1. Conjugation Un-conjugated Alternate Names T-cell-restricted intracellular antigen-1; WDM; TIA-1; RNA-binding protein TIA-1; p40-TIA-1; Nucleolysin TIA-1 isoform p40 Application Instructions Application table Application Dilution IHC-P 1:50 - 1:200 WB 1:500 - 1:2000 Application Note IHC-P: Antigen Retrieval: Heat mediated. * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. Calculated Mw 43 kDa Observed Size ~ 45 kDa Properties Form Liquid Purification Affinity purified. Buffer PBS (pH 7.4), 150mM NaCl, 0.02% Sodium azide and 50% Glycerol. Preservative 0.02% Sodium azide Stabilizer 50% Glycerol Storage instruction For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot and store at -20°C. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. www.arigobio.com 1/2 Note For laboratory research only, not for drug, diagnostic or other use. Bioinformation Gene Symbol TIA1 Gene Full Name TIA1 cytotoxic granule-associated RNA binding protein Background The product encoded by this gene is a member of a RNA-binding protein family and possesses nucleolytic activity against cytotoxic lymphocyte (CTL) target cells. -

MOCHI Enables Discovery of Heterogeneous Interactome Modules in 3D Nucleome
Downloaded from genome.cshlp.org on October 4, 2021 - Published by Cold Spring Harbor Laboratory Press MOCHI enables discovery of heterogeneous interactome modules in 3D nucleome Dechao Tian1,# , Ruochi Zhang1,# , Yang Zhang1, Xiaopeng Zhu1, and Jian Ma1,* 1Computational Biology Department, School of Computer Science, Carnegie Mellon University, Pittsburgh, PA 15213, USA #These two authors contributed equally *Correspondence: [email protected] Contact To whom correspondence should be addressed: Jian Ma School of Computer Science Carnegie Mellon University 7705 Gates-Hillman Complex 5000 Forbes Avenue Pittsburgh, PA 15213 Phone: +1 (412) 268-2776 Email: [email protected] 1 Downloaded from genome.cshlp.org on October 4, 2021 - Published by Cold Spring Harbor Laboratory Press Abstract The composition of the cell nucleus is highly heterogeneous, with different constituents forming complex interactomes. However, the global patterns of these interwoven heterogeneous interactomes remain poorly understood. Here we focus on two different interactomes, chromatin interaction network and gene regulatory network, as a proof-of-principle, to identify heterogeneous interactome modules (HIMs), each of which represents a cluster of gene loci that are in spatial contact more frequently than expected and that are regulated by the same group of transcription factors. HIM integrates transcription factor binding and 3D genome structure to reflect “transcriptional niche” in the nucleus. We develop a new algorithm MOCHI to facilitate the discovery of HIMs based on network motif clustering in heterogeneous interactomes. By applying MOCHI to five different cell types, we found that HIMs have strong spatial preference within the nucleus and exhibit distinct functional properties. Through integrative analysis, this work demonstrates the utility of MOCHI to identify HIMs, which may provide new perspectives on the interplay between transcriptional regulation and 3D genome organization. -

TIAL1 Rabbit Pab
Leader in Biomolecular Solutions for Life Science TIAL1 Rabbit pAb Catalog No.: A6075 Basic Information Background Catalog No. The protein encoded by this gene is a member of a family of RNA-binding proteins, has A6075 three RNA recognition motifs (RRMs), and binds adenine and uridine-rich elements in mRNA and pre-mRNAs of a wide range of genes. It regulates various activities including Observed MW translational control, splicing and apoptosis. Alternate transcriptional splice variants, 47kDa encoding different isoforms, have been characterized. The different isoforms have been show to function differently with respect to post-transcriptional silencing. Calculated MW 41kDa/43kDa Category Primary antibody Applications WB, IHC Cross-Reactivity Human, Mouse, Rat Recommended Dilutions Immunogen Information WB 1:500 - 1:2000 Gene ID Swiss Prot 7073 Q01085 IHC 1:100 - 1:200 Immunogen Recombinant fusion protein containing a sequence corresponding to amino acids 266-375 of human TIAL1 (NP_003243.1). Synonyms TIAL1;TCBP;TIAR Contact Product Information www.abclonal.com Source Isotype Purification Rabbit IgG Affinity purification Storage Store at -20℃. Avoid freeze / thaw cycles. Buffer: PBS with 0.02% sodium azide,50% glycerol,pH7.3. Validation Data Western blot analysis of extracts of various cell lines, using TIAL1 antibody (A6075) at 1:3000 dilution. Secondary antibody: HRP Goat Anti-Rabbit IgG (H+L) (AS014) at 1:10000 dilution. Lysates/proteins: 25ug per lane. Blocking buffer: 3% nonfat dry milk in TBST. Detection: ECL Basic Kit (RM00020). Exposure time: 90s. Immunohistochemistry of paraffin- Immunohistochemistry of paraffin- Immunohistochemistry of paraffin- embedded rat lung using TIAL1 antibody embedded human appendix using TIAL1 embedded mouse spleen using TIAL1 (A6075) at dilution of 1:100 (40x lens).