Streptomyces Coelicolor: Genetic and Physiological Changes Induced by a Chemical Probe
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

UC Berkeley UC Berkeley Electronic Theses and Dissertations
UC Berkeley UC Berkeley Electronic Theses and Dissertations Title Analyzing Microbial Physiology and Nutrient Transformation in a Model, Acidophilic Microbial Community using Integrated `Omics' Technologies Permalink https://escholarship.org/uc/item/259113st Author Justice, Nicholas Bruce Publication Date 2013 Supplemental Material https://escholarship.org/uc/item/259113st#supplemental Peer reviewed|Thesis/dissertation eScholarship.org Powered by the California Digital Library University of California Analyzing Microbial Physiology and Nutrient Transformation in a Model, Acidophilic Microbial Community using Integrated ‘Omics’ Technologies By Nicholas Bruce Justice A dissertation submitted in partial satisfaction of the requirements for the degree of Doctor of Philosophy in Microbiology in the Graduate Division of the University of California, Berkeley Committee in charge: Professor Jillian Banfield, Chair Professor Mary Firestone Professor Mary Power Professor John Coates Fall 2013 Abstract Analyzing Microbial Physiology and Nutrient Transformation in a Model, Acidophilic Microbial Community using Integrated ‘Omics’ Technologies by Nicholas Bruce Justice Doctor of Philosophy in Microbiology University of California, Berkeley Professor Jillian F. Banfield, Chair Understanding how microorganisms contribute to nutrient transformations within their community is critical to prediction of overall ecosystem function, and thus is a major goal of microbial ecology. Communities of relatively tractable complexity provide a unique opportunity to study the distribution of metabolic characteristics amongst microorganisms and how those characteristics subscribe diverse ecological functions to co-occurring, and often closely related, species. The microbial communities present in the low-pH, metal-rich environment of the acid mine drainage (AMD) system in Richmond Mine at Iron Mountain, CA constitute a model microbial community due to their relatively low diversity and extensive characterization over the preceding fifteen years. -

Product Sheet Info
Master Clone List for NR-19279 ® Vibrio cholerae Gateway Clone Set, Recombinant in Escherichia coli, Plates 1-46 Catalog No. NR-19279 Table 1: Vibrio cholerae Gateway® Clones, Plate 1 (NR-19679) Clone ID Well ORF Locus ID Symbol Product Accession Position Length Number 174071 A02 367 VC2271 ribD riboflavin-specific deaminase NP_231902.1 174346 A03 336 VC1877 lpxK tetraacyldisaccharide 4`-kinase NP_231511.1 174354 A04 342 VC0953 holA DNA polymerase III, delta subunit NP_230600.1 174115 A05 388 VC2085 sucC succinyl-CoA synthase, beta subunit NP_231717.1 174310 A06 506 VC2400 murC UDP-N-acetylmuramate--alanine ligase NP_232030.1 174523 A07 132 VC0644 rbfA ribosome-binding factor A NP_230293.2 174632 A08 322 VC0681 ribF riboflavin kinase-FMN adenylyltransferase NP_230330.1 174930 A09 433 VC0720 phoR histidine protein kinase PhoR NP_230369.1 174953 A10 206 VC1178 conserved hypothetical protein NP_230823.1 174976 A11 213 VC2358 hypothetical protein NP_231988.1 174898 A12 369 VC0154 trmA tRNA (uracil-5-)-methyltransferase NP_229811.1 174059 B01 73 VC2098 hypothetical protein NP_231730.1 174075 B02 82 VC0561 rpsP ribosomal protein S16 NP_230212.1 174087 B03 378 VC1843 cydB-1 cytochrome d ubiquinol oxidase, subunit II NP_231477.1 174099 B04 383 VC1798 eha eha protein NP_231433.1 174294 B05 494 VC0763 GTP-binding protein NP_230412.1 174311 B06 314 VC2183 prsA ribose-phosphate pyrophosphokinase NP_231814.1 174603 B07 108 VC0675 thyA thymidylate synthase NP_230324.1 174474 B08 466 VC1297 asnS asparaginyl-tRNA synthetase NP_230942.2 174933 B09 198 -

Comparative Genomics Study of Polyhydroxyalkanoates (PHA) and Ectoine Relevant Genes from Halomonas Sp
Cai et al. Microbial Cell Factories 2011, 10:88 http://www.microbialcellfactories.com/content/10/1/88 RESEARCH Open Access Comparative genomics study of polyhydroxyalkanoates (PHA) and ectoine relevant genes from Halomonas sp. TD01 revealed extensive horizontal gene transfer events and co-evolutionary relationships Lei Cai1†, Dan Tan1†, Gulsimay Aibaidula2, Xin-Ran Dong3, Jin-Chun Chen1, Wei-Dong Tian3* and Guo-Qiang Chen1* Abstract Background: Halophilic bacteria have shown their significance in industrial production of polyhydroxyalkanoates (PHA) and are gaining more attention for genetic engineering modification. Yet, little information on the genomics and PHA related genes from halophilic bacteria have been disclosed so far. Results: The draft genome of moderately halophilic bacterium, Halomonas sp. TD01, a strain of great potential for industrial production of short-chain-length polyhydroxyalkanoates (PHA), was analyzed through computational methods to reveal the osmoregulation mechanism and the evolutionary relationship of the enzymes relevant to PHA and ectoine syntheses. Genes involved in the metabolism of PHA and osmolytes were annotated and studied in silico. Although PHA synthase, depolymerase, regulator/repressor and phasin were all involved in PHA metabolic pathways, they demonstrated different horizontal gene transfer (HGT) events between the genomes of different strains. In contrast, co-occurrence of ectoine genes in the same genome was more frequently observed, and ectoine genes were more likely under coincidental horizontal gene transfer than PHA related genes. In addition, the adjacent organization of the homologues of PHA synthase phaC1 and PHA granule binding protein phaP was conserved in the strain TD01, which was also observed in some halophiles and non-halophiles exclusively from g- proteobacteria. -

Systems Metabolic Engineering of Basfia Succiniciproducens For
Systems metabolic engineering of Corynebacterium glutamicum for production of L-lysine and ectoine Dissertation zur Erlangung des Grades des Doktors der Ingenieurwissenschaften der Naturwissenschaftlich-Technischen Fakultät der Universität des Saarlandes von Gideon Gießelmann Saarbrücken 2019 Tag des Kolloquiums: 16.7.2019 Dekan: Prof. Guido Kickelbick Berichterstatter 1: Prof. Christoph Wittmann Berichterstatter 2: Prof. Gert-Wieland Kohring Vorsitz: Prof. Bruce Morgan Akad. Mitarbeiter: Dr. Björn Becker Publications Partial results of this work have been published previously. This was authorized by the Institute of Systems Biotechnology, represented by Prof. Dr. Christoph Wittmann. Peer-reviewed articles: Gießelmann G, D. Dietrich, L. Jungmann, M. Kohlstedt, E. Jung Jeon, S. Sun Yim, F. Sommer, D. Zimmer, T. Mühlhaus, M. Schroda, K. Jun Jeong, J. Becker, and C. Wittmann, 2019. Metabolic engineering of Corynebacterium glutamicum for high-level ectoine production – design, combinatorial assembly and implementation of a transcriptionally balanced heterologous ectoine pathway. Biotechnology Journal, under revision Becker J, G. Gießelmann, SL. Hoffmann, C. Wittmann, 2016: Corynebacterium glutamicum for sustainable bioproduction: from metabolic physiology to systems metabolic engineering. In: Zhao H., Zeng AP. (eds) Synthetic Biology – Metabolic Engineering. Advances in Biochemical Engineering/Biotechnology, vol 162. Springer, Cham The following peer-reviewed articles were published during this work, but are not part of the dissertation: Rohles CM, L. Gläser, M. Kohlstedt, G. Gießelmann, S. Pearson, A. del Campo, J. Becker, C. Wittmann, 2018. A bio-based route to the carbon-5 chemical glutaric acid and to bionylon-6,5 using metabolically engineered Corynebacterium glutamicum, Green Chemistry 20:4662-4674. Vassilev I, G. Gießelmann, SK. Schwechheimer, C. Wittmann, B Virdis, JO. -

Metabolic Engineering of Escherichia Coli for Ectoine Production with the Fermentation Strategy of Supplementing Amino Donor
Metabolic Engineering of Escherichia Coli for Ectoine Production With the Fermentation Strategy of Supplementing Amino Donor Hao Zhang Nanjing Tech University Yingying Cao Nanjing Tech University Yingsheng Dong Nanjing Tech University Ming Zhao Nanjing Tech University Yanqin Ma Nanjing Tech University Zhengshan Luo Nanjing Tech University Sha Li ( [email protected] ) Nanjing Tech University https://orcid.org/0000-0001-8256-1263 Hong Xu Nanjing Tech University Research Keywords: Ectoine, Escherichia coli, Metabolic engineering, Medium optimization, Amino donor, RT qPCR Posted Date: December 28th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-134030/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/17 Abstract Background: Ectoine, a compatible solute, has broad application prospects in food biotechnology, agriculture, medicine, and cosmetics because of its protective action on biological compounds. Industrially, ectoine is produced by halophilic bacteria in a complex process. Recently, various works focus on improving ectoine production by using engineered strains, but there are still problems of low yield and low ectoine production eciency. Results: To overcome the drawback, a nal metabolic engineered strain E. coli ET08 was constructed by eliminating lysine synthesis branch and by-product metabolic pathways, and ectoine production reached 10.2 g/L through culture medium optimization. Compared with nitrate, addition of ammonium salt contributed more to the ectoine synthesis. Furthermore, the ammonium sulphate boosted more ectoine titers than ammonium chloride and sodium glutamate. The analysis of transcriptional levels revealed that the ammonium sulfate enhanced ectoine biosynthesis by enhancing metabolic ux toward ectoine biosynthesis and providing auent synthetic precursors. -

Pseudomonas/Brachypodium As a Model System for Studying Rhizosphere Plant Microbe Interactions Under Water Stress
The University of Southern Mississippi The Aquila Digital Community Master's Theses Fall 12-1-2018 Pseudomonas/Brachypodium as a Model System for Studying Rhizosphere Plant Microbe Interactions Under Water Stress Janiece McWilliams University of Southern Mississippi Follow this and additional works at: https://aquila.usm.edu/masters_theses Recommended Citation McWilliams, Janiece, "Pseudomonas/Brachypodium as a Model System for Studying Rhizosphere Plant Microbe Interactions Under Water Stress" (2018). Master's Theses. 602. https://aquila.usm.edu/masters_theses/602 This Masters Thesis is brought to you for free and open access by The Aquila Digital Community. It has been accepted for inclusion in Master's Theses by an authorized administrator of The Aquila Digital Community. For more information, please contact [email protected]. PSEUDOMONAS/BRACHYPODIUM AS A MODEL SYSTEM FOR STUDYING RHIZOSPHERE PLANT-MICROBE INTERACTIONS UNDER WATER STRESS by Janiece Hutchins McWilliams A Thesis Submitted to the Graduate School, the College of Arts and Sciences and the School of Biological, Environmental, and Earth Sciences at The University of Southern Mississippi in Partial Fulfillment of the Requirements for the Degree of Master of Science Approved by: Dr. Dmitri Mavrodi, Committee Chair Dr. Kevin Kuehn Dr. Micheal Davis ____________________ ____________________ ____________________ Dr. Dmitri Mavrodi Dr. Jake Schaefer Dr. Karen S. Coats Committee Chair Director of School Dean of the Graduate School December 2018 ABSTRACT In contrast to well-studied mechanisms of drought tolerance in plants, the interactions between plants and their microbiome during water stress are still poorly understood. This missing knowledge is crucial for the exploitation of beneficial microbial communities to improve the sustainability of agriculture under changing climatic conditions. -

Cellular Adjustments of Bacillus Subtilis and Other Bacilli to Fluctuating Salinities
Chapter 15 Cellular Adjustments of Bacillus subtilis and Other Bacilli to Fluctuating Salinities Marco Pittelkow and Erhard Bremer 15.1 The Osmotic Challenge Bacilli are exposed, either suddenly or on a sustained basis, to changes in the osmotic conditions of the varied ecological niches that they can colonize. This can be vividly visualized by considering the main habitat of Bacillus subtilis, the upper layers of the soil (Earl et al. 2008). Rainfall and drying of the soil cause fluctuations in water availability and in the osmotic potential of this ecological niche. Such changes pose a considerable challenge to the microbial cell since they elicit water fluxes across the cytoplasmic membrane. These water fluxes are driven by the differential in the osmotic potential between the cell’s interior and that of the surrounding microenvironment (Bremer and Kr€amer 2000). Water entry at low osmolarity can increase turgor to such an extent that the elastic and stress-bearing peptidoglycan sacculus can no longer cope with it and the cell will rupture. Water efflux at high osmolarity will trigger dehydration of the cytoplasm and the ensuing reduction or even collapse of turgor will cause growth arrest or even cell death. Turgor, an intracellular hydrostatic pressure considered to be essential for cell expansion and growth, is difficult to determine experimentally but has been estimated at 1.9 MPa (19.37 atm) for B. subtilis (Whatmore and Reed 1990), a pressure that is close to ten times the pressure present in a standard car tire. To maintain turgor within physiologically acceptable boundaries, the B. subtilis cell needs to take active countermeasures to redirect the flow of water in or out of the cell when it faces decreases or increases in the external osmolarity. -

"Ectoine Synthase: an Iron-Dependent Member of the Cupin Superfamily" In
Ectoine synthase: an iron-dependent member of the cupin superfamily Laura Czech†,‡, Astrid Hoeppner§, Sander HJ Smits§,¶, and Erhard Bremer†,‡ †Department of Biology, Laboratory for Microbiology, Philipps-University Marburg, Marburg, Germany ‡SYNMIKRO Research Center, Philipps-University Marburg, Marburg, Germany §Center for Structural Studies, Heinrich-Heine University Düsseldorf, Düsseldorf, Germany ¶Institute of Biochemistry, Heinrich-Heine University Düsseldorf, Düsseldorf, Germany FUNCTIONAL CLASS harbors a catalytically critical Fe2+ ion. The three residues Enzyme; ectoine synthase (EctC), EC 4.2.1.108. EctC coordinating this metal protrude into the lumen of the belongs to the carbon–oxygen lyases (EC 4.2) and cupin barrel in which the EctC-promoted enzyme reaction, more specifically to the hydro-lyases (EC 4.2.1), a a water elimination, is catalyzed, using N-γ-ADABA as the 1 group of enzymes that cleaves carbon–oxygen bonds. substrate. The EctC enzyme catalyzes in an Fe2+-dependent fashion the last step of ectoine [(S)-2-methyl-1,4,5,6- tetrahydropyrimidine-4-carboxylic acid] synthesis through OCCURENCE cyclocondensation of the EctA-formed substrate N- γ-acetyl-L-2,4-diaminobutyric acid (N-γ-ADABA) via a EctC is the diagnostic enzyme of the ectoine biosynthetic 7–10 water-elimination reaction. The reaction product, ectoine, route, a set of three enzymes that produces this effective 11–13 is a member of the so-called compatible solutes and microbial stress-protectant and chemical chaperone. 14 serves as a cytoprotectant against osmotic stress and Biosynthesis of ectoine initiates from the central micro- 9,10,15–17 growth temperature extremes. It also functions as a bial metabolite L-aspartate-β-semialdehyde. -
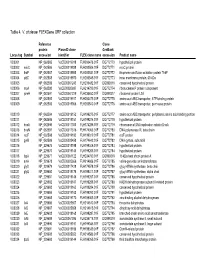
Table 4. V. Cholerae Flexgene ORF Collection
Table 4. V. cholerae FLEXGene ORF collection Reference Clone protein PlasmID clone GenBank Locus tag Symbol accession identifier FLEX clone name accession Product name VC0001 NP_062585 VcCD00019918 FLH200476.01F DQ772770 hypothetical protein VC0002 mioC NP_062586 VcCD00019938 FLH200506.01F DQ772771 mioC protein VC0003 thdF NP_062587 VcCD00019958 FLH200531.01F DQ772772 thiophene and furan oxidation protein ThdF VC0004 yidC NP_062588 VcCD00019970 FLH200545.01F DQ772773 inner membrane protein, 60 kDa VC0005 NP_062589 VcCD00061243 FLH236482.01F DQ899316 conserved hypothetical protein VC0006 rnpA NP_062590 VcCD00025697 FLH214799.01F DQ772774 ribonuclease P protein component VC0007 rpmH NP_062591 VcCD00061229 FLH236450.01F DQ899317 ribosomal protein L34 VC0008 NP_062592 VcCD00019917 FLH200475.01F DQ772775 amino acid ABC transporter, ATP-binding protein VC0009 NP_062593 VcCD00019966 FLH200540.01F DQ772776 amino acid ABC transproter, permease protein VC0010 NP_062594 VcCD00019152 FLH199275.01F DQ772777 amino acid ABC transporter, periplasmic amino acid-binding portion VC0011 NP_062595 VcCD00019151 FLH199274.01F DQ772778 hypothetical protein VC0012 dnaA NP_062596 VcCD00017363 FLH174286.01F DQ772779 chromosomal DNA replication initiator DnaA VC0013 dnaN NP_062597 VcCD00017316 FLH174063.01F DQ772780 DNA polymerase III, beta chain VC0014 recF NP_062598 VcCD00019182 FLH199319.01F DQ772781 recF protein VC0015 gyrB NP_062599 VcCD00025458 FLH174642.01F DQ772782 DNA gyrase, subunit B VC0016 NP_229675 VcCD00019198 FLH199346.01F DQ772783 hypothetical protein -

Metabolic Engineering of Corynebacterium Glutamicum for Production of the Chemical Chaperone Ectoine
Metabolic engineering of Corynebacterium glutamicum for the production of L-aspartate and its derivatives β-alanine and ectoine Dissertation zur Erlangung des akademischen Grades Doktor der Naturwissenschaften (Dr. rer. nat.) Fakultät für Biologie Universität Bielefeld vorgelegt von Christian Ziert aus Fulda Ebersberg, April 2014 Table of contents Table of figures .................................................................................................8 List of tables ......................................................................................................9 Abbreviations .................................................................................................. 10 Summary ......................................................................................................... 11 Zusammenfassung .......................................................................................... 12 1 Introduction .............................................................................................. 13 1.1 Characteristics of Corynebacterium glutamicum and its application in biotechnology ........... 13 1.2 The central metabolism of C. glutamicum ............................................................................... 14 1.2.1 Carbon source uptake ...................................................................................................... 16 1.2.2 Glycolysis .......................................................................................................................... 17 1.2.3 Pentose -

Biosynthesis of the Acetyl-Coa Carboxylase-Inhibiting Antibiotic, Andrimid, in Serratia Is Regulated by Hfq and The
CORE Metadata, citation and similar papers at core.ac.uk Provided by Apollo 1 Biosynthesis of the acetyl-CoA carboxylase-inhibiting 2 antibiotic, andrimid, in Serratia is regulated by Hfq and the 3 LysR-type transcriptional regulator, AdmX. 4 Miguel A. Matillaab*, Veronika Nogellovaa, Bertrand Morelb, Tino Krellb and 5 George P.C. Salmonda* 6 7 aDepartment of Biochemistry, University of Cambridge, Tennis Court Road, 8 Cambridge, UK, CB2 1QW. 9 bDepartment of Environmental Protection, Estación Experimental del Zaidín, 10 Consejo Superior de Investigaciones Científicas, Prof. Albareda 1, Granada, 11 Spain, 18008 12 13 Running title: Regulation of andrimid biosynthesis 14 15 *Address correspondence to George P.C. Salmond, Department of 16 Biochemistry, University of Cambridge, Tennis Court Road, Cambridge, UK, 17 CB2 1QW. Tel: +44 (0)1223 333650; Fax: +44 (0)1223 766108; E-mail: 18 [email protected] *Address correspondence to Miguel A. Matilla, Department of Environmental Protection, Estación Experimental del Zaidín, Consejo Superior de Investigaciones Científicas, Prof. Albareda 1, Granada, Spain, 18008. Tel: +34 958 181600; Fax: + 34 958 135740; E-mail: [email protected] 1 1 SUMMARY 2 Infections due to multidrug-resistant bacteria represent a major global health challenge. 3 To combat this problem, new antibiotics are urgently needed and some plant- 4 associated bacteria are a promising source. The rhizobacterium Serratia plymuthica 5 A153 produces several bioactive secondary metabolites, including the anti-oomycete 6 and antifungal haterumalide, oocydin A, and the broad spectrum polyamine antibiotic, 7 zeamine. In this study, we show that A153 produces a second broad spectrum 8 antibiotic, andrimid. -

Uncovering the Mechanisms of Halotolerance in the Extremely Acidophilic Members of the Acidihalobacter Genus Through Comparative Genome Analysis
fmicb-10-00155 February 8, 2019 Time: 16:55 # 1 ORIGINAL RESEARCH published: 08 February 2019 doi: 10.3389/fmicb.2019.00155 Uncovering the Mechanisms of Halotolerance in the Extremely Acidophilic Members of the Acidihalobacter Genus Through Comparative Genome Analysis Himel N. Khaleque1,2, Carolina González3, Raihan Shafique4, Anna H. Kaksonen2, David S. Holmes3,5 and Elizabeth L. J. Watkin1* 1 School of Pharmacy and Biomedical Sciences, Curtin Health Innovation Research Institute, Curtin University, Perth, WA, Australia, 2 CSIRO Land and Water, Floreat, WA, Australia, 3 Center for Bioinformatics and Genome Biology, Science for Life Foundation, Santiago, Chile, 4 Sodexo Australia, Perth, WA, Australia, 5 Centro de Genómica y Bioinformática, Facultad de Ciencias, Universidad Mayor, Santiago, Chile There are few naturally occurring environments where both acid and salinity stress exist Edited by: together, consequently, there has been little evolutionary pressure for microorganisms to Claudia P. Saavedra, develop systems that enable them to deal with both stresses simultaneously. Members Universidad Andrés Bello, Chile of the genus Acidihalobacter are iron- and sulfur-oxidizing, halotolerant acidophiles that Reviewed by: Mario A. Vera, have developed the ability to tolerate acid and saline stress and, therefore, have the Pontificia Universidad Católica potential to bioleach ores with brackish or saline process waters under acidic conditions. de Chile, Chile T Nicolas Guiliani, The genus consists of four members, A. prosperus DSM 5130 , A. prosperus DSM Universidad de Chile, Chile 14174, A. prosperus F5 and “A. ferrooxidans” DSM 14175. An in depth genome *Correspondence: comparison was undertaken in order to provide a more comprehensive description Elizabeth L. J.