A Population-Based Atlas of the Human Pyramidal Tract in 410
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Network of Genetic Repression and Derepression Specifies Projection
A network of genetic repression and derepression INAUGURAL ARTICLE specifies projection fates in the developing neocortex Karpagam Srinivasana,1, Dino P. Leonea, Rosalie K. Batesona, Gergana Dobrevab, Yoshinori Kohwic, Terumi Kohwi-Shigematsuc, Rudolf Grosschedlb, and Susan K. McConnella,2 aDepartment of Biology, Stanford University, Stanford, CA 94305; bMax-Planck Institute of Immunobiology and Epigenetics, 79108 Freiburg, Germany; and cLawrence Berkeley National Laboratory, Berkeley, CA 94720 This contribution is part of the special series of Inaugural Articles by members of the National Academy of Sciences elected in 2011. Contributed by Susan K. McConnell, September 28, 2012 (sent for review August 24, 2012) Neurons within each layer in the mammalian cortex have stereotypic which results in a suppression of corticothalamic and callosal projections. Four genes—Fezf2, Ctip2, Tbr1, and Satb2—regulate fates, respectively, and axon extension along the corticospinal these projection identities. These genes also interact with each tract (CST). Fezf2 mutant neurons fail to repress Satb2 and Tbr1, other, and it is unclear how these interactions shape the final pro- thus their axons cross the CC and/or innervate the thalamus jection identity. Here we show, by generating double mutants of inappropriately. Fezf2, Ctip2, and Satb2, that cortical neurons deploy a complex In callosal projection neurons, Satb2 represses the expression Ctip2 Bhlhb5 genetic switch that uses mutual repression to produce subcortical of and , leading to a repression of subcerebral fates. Satb2 Tbr1 or callosal projections. We discovered that Tbr1, EphA4, and Unc5H3 Interestingly, we found that promotes expression in fi upper layer callosal neurons, and Tbr1 expression in these neu- are critical downstream targets of Satb2 in callosal fate speci ca- fi tion. -

Spinal Tracts.Pdf
Spinal Tracts Andreas Talgø Lie Illustrations by: Peder Olai Skjeflo Holman Previous material: Maja Solbakken Definitions to bring home Nerve Ganglion Neuron Nucleus Tract Collection neurons Collection of nerve A single cell Collection of Collection of axons that transmits cell bodies in the transmitting nerve cell bodies in traveling up or sensation or motor PNS, typically linked electrical impluses. the CNS, typically down the spinal impulses depending by synapses. linked by synapses. cord, depending on the function and Location: both on function destination. Location: PNS Location: CNS and destination. Location: PNS Location: CNS - does it matter? Tract - highway to pass anatomy exam? • Highway = Tract • Lane = Neuron • Car = Signal Slow... Fast!! Just to make sure... • Ipsilateral or contralateral • Ventral = anterior • Dorsal = posterior High yield points to understand • What does the tract transmit - Motor or sensory? If sensory: which sensation? • Where does the neurones synapse - 1.st, 2nd and 3rd order neron? Which ganglion/nuclei? • Where there are decussations - If there are any decussations at all? ASCENDING / SENSORY TRACTS Sensations * Temperature NOT transmitted by the tracts: * Pressure * Visualization * Pain * Audition * Fine touch * Olfaction * Crude touch * Gustation * Proprioception * Vibration Sensations Precise sensation Primitive sensation Fine touch Crude touch Pressure Pain Vibration Temperature Proprioception Other: sexual, itching, tickling Ascending tracts / Sensory tracts Dorsal coulmn Lat. Spinothalamic Ant. -

The Superior and Inferior Colliculi of the Mole (Scalopus Aquaticus Machxinus)
THE SUPERIOR AND INFERIOR COLLICULI OF THE MOLE (SCALOPUS AQUATICUS MACHXINUS) THOMAS N. JOHNSON' Laboratory of Comparative Neurology, Departmmt of Amtomy, Un&versity of hfiehigan, Ann Arbor INTRODUCTION This investigation is a study of the afferent and efferent connections of the tectum of the midbrain in the mole (Scalo- pus aquaticus machrinus). An attempt is made to correlate these findings with the known habits of the animal. A subterranean animal of the middle western portion of the United States, Scalopus aquaticus machrinus is the largest of the genus Scalopus and its habits have been more thor- oughly studied than those of others of this genus according to Jackson ('15) and Hamilton ('43). This animal prefers a well-drained, loose soil. It usually frequents open fields and pastures but also is found in thin woods and meadows. Following a rain, new superficial burrows just below the surface of the ground are pushed in all directions to facili- tate the capture of worms and other soil life. Ten inches or more below the surface the regular permanent highway is constructed; the mole retreats here during long periods of dry weather or when frost is in the ground. The principal food is earthworms although, under some circumstances, larvae and adult insects are the more usual fare. It has been demonstrated conclusively that, under normal conditions, moles will eat vegetable matter. It seems not improbable that they may take considerable quantities of it at times. A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the University of Michigan. -
L4-Physiology of Motor Tracts.Pdf
: chapter 55 page 667 Objectives (1) Describe the upper and lower motor neurons. (2) Understand the pathway of Pyramidal tracts (Corticospinal & corticobulbar tracts). (3) Understand the lateral and ventral corticospinal tracts. (4) Explain functional role of corticospinal & corticobulbar tracts. (5) Describe the Extrapyramidal tracts as Rubrospinal, Vestibulospinal, Reticulospinal and Tectspinal Tracts. The name of the tract indicate its pathway, for example Corticobulbar : Terms: - cortico: cerebral cortex. Decustation: crossing. - Bulbar: brainstem. Ipsilateral : same side. *So it starts at cerebral cortex and Contralateral: opposite side. terminate at the brainstem. CNS influence the activity of skeletal muscle through two set of neurons : 1- Upper motor neurons (UMN) 2- lower motor neuron (LMN) They are neurons of motor cortex & their axons that pass to brain stem and They are Spinal motor neurons in the spinal spinal cord to activate: cord & cranial motor neurons in the brain • cranial motor neurons (in brainstem) stem which innervate muscles directly. • spinal motor neurons (in spinal cord) - These are the only neurons that innervate - Upper motor neurons (UMN) are the skeletal muscle fibers, they function as responsible for conveying impulses for the final common pathway, the final link voluntary motor activity through between the CNS and skeletal muscles. descending motor pathways that make up by the upper motor neurons. Lower motor neurons are classified based on the type of muscle fiber the innervate: There are two UMN Systems through which 1- alpha motor neurons (UMN) control (LMN): 2- gamma motor neurons 1- Pyramidal system (corticospinal tracts ). 2- Extrapyramidal system The activity of the lower motor neuron (LMN, spinal or cranial) is influenced by: 1. -

(Corticobulbar) Tract
Cor$cospinal (cor$cobulbar) Tract 6/18/12 1. Appreciate the functions of the corticospinal & corticobulbar tracts. 2. Distinguish corticospinal and corticobulbar tracts. Theodore Tzavaras MD2015 3. Identify the locations of both Laurie L. Wellman Ph.D. tracts through the forebrain, Eastern Virginia Medical School brainstem and spinal cord. 4. Identify the level of decussation of the corticospinal tract, and it’s clinical importance. Dr. Craig Goodmurphy Anatomy Guy Pathway Overview Pathway Overview Corticospinal Corticobulbar ² Begin in primary motor cortex. ² Begin in primary motor cortex. ² Upper ² Travel through the posterior limb Travel through the posterior limb Upper of the internal capsule (somatotopic). Motor of the internal capsule (somatotopic). Motor ² Descend through the middle 3/5 of Neuron ² Descend through the middle 3/5 of the crus cerebri (somatotopic). the crus cerebri (somatotopic). Neuron ² Travel through the brainstem as ² Travel through the brainstem and the descending pyramidal system. synapse on brainstem nuclei. Crosses Lower ² Decussate at the caudal medulla in Midline ² α-Motor neurons project along the pyramidal decussation. cranial nerves for facial movements Motor ² Descend in the spinal cord as the and voice production. Neuron corticospinal tract Lower ² All cranial nerve nuclei receive ² Synapse on α-motor neurons Motor bilateral UMN input* ² Exit spinal cord in ventral rami Neuron *Part of CN VI is the exception AnatomyGuy.com 1 Cor$cospinal (cor$cobulbar) Tract 6/18/12 Overview Corticospinal ² Primary motor cortex Don’t ü ² Try to learn all the details on your first Posterior limb of the internal pass. capsule (somatotopic). Do ü Try to sketch out each brain, ² Middle 3/5 of the crus cerebri brainstem, and cord level we showed (somatotopic). -

Identification of the Corticobulbar Tracts of the Tongue and Face Using Deterministic and Probabilistic DTI Fiber Tracking in Pa
Published August 6, 2015 as 10.3174/ajnr.A4430 ORIGINAL RESEARCH ADULT BRAIN Identification of the Corticobulbar Tracts of the Tongue and Face Using Deterministic and Probabilistic DTI Fiber Tracking in Patients with Brain Tumor M. Jenabi, X K.K. Peck, X R.J. Young, N. Brennan, and X A.I. Holodny ABSTRACT BACKGROUND AND PURPOSE: The corticobulbar tract of the face and tongue, a critical white matter tract connecting the primary motor cortex and the pons, is rarely detected by deterministic DTI fiber tractography. Detection becomes even more difficult in the presence of a tumor. The purpose of this study was to compare identification of the corticobulbar tract by using deterministic and probabilistic tractography in patients with brain tumor. MATERIALS AND METHODS: Fifty patients with brain tumor who underwent DTI were studied. Deterministic tractography was per- formed by using the fiber assignment by continuous tractography algorithm. Probabilistic tractography was performed by using a Monte Carlo simulation method. ROIs were drawn of the face and tongue motor homunculi and the pons in both hemispheres. RESULTS: In all subjects, fiber assignment by continuous tractography was ineffectual in visualizing the entire course of the corticobulbar tract between the face and tongue motor cortices and the pons on either side. However, probabilistic tractography successfully visualized the corticobulbar tract from the face and tongue motor cortices in all patients on both sides. No significant difference (P Ͻ .08) was found between both sides in terms of the number of voxels or degree of connectivity. The fractional anisotropy of both the face and tongue was significantly lower on the tumor side (P Ͻ .03). -

Do the Corticospinal and Corticobulbar Tracts Mediate Functions in the Human Newborn?
LE JOURNAL CANAD1EN DES SCIENCES NEUROLOGIQUES HYPOTHESIS Do the Corticospinal and Corticobulbar Tracts Mediate Functions in the Human Newborn? Harvey B. Sarnat ABSTRACT: Unlike the numerous dispersed bulbospinal pathways that are already well myelinated at term, the more compact corticospinal and corticobulbar tracts are only beginning their myelination cycle in late gestation and do not complete it until two years of age. During this same period, these pathways also develop extensive ramification of ter minal axonal segments, growth of collateral axons, and proliferation of synapses. Despite their immaturity in the full- term human newborn, several proposed functions may be attributed to the descending pathways from the neonatal cerebral cortex: a) a contribution to the differential development of passive muscle tone and resting postures; in general they function as an antagonist to the "subcorticospinal pathways" in mediating proximal flexion and distal extension, except for the rubrospinal tract which is probably synergistic with the corticospinal tract; b) enhancement of tactile reflexes originating in the brainstem and spinal cord, including suck and swallow; c) relay of epileptic activity of corti cal origin; d) inhibition of complex stereotyped motor reflexes including many phenomena formerly termed "subtle seizures"; e) a possible influence on muscle maturation, particularly in relaying cerebellar impulses that modify the histochemical differentiation of myofibres. However, the bulbospinal tracts are probably more influential -

Brainstem Dysfunction in Critically Ill Patients
Benghanem et al. Critical Care (2020) 24:5 https://doi.org/10.1186/s13054-019-2718-9 REVIEW Open Access Brainstem dysfunction in critically ill patients Sarah Benghanem1,2 , Aurélien Mazeraud3,4, Eric Azabou5, Vibol Chhor6, Cassia Righy Shinotsuka7,8, Jan Claassen9, Benjamin Rohaut1,9,10† and Tarek Sharshar3,4*† Abstract The brainstem conveys sensory and motor inputs between the spinal cord and the brain, and contains nuclei of the cranial nerves. It controls the sleep-wake cycle and vital functions via the ascending reticular activating system and the autonomic nuclei, respectively. Brainstem dysfunction may lead to sensory and motor deficits, cranial nerve palsies, impairment of consciousness, dysautonomia, and respiratory failure. The brainstem is prone to various primary and secondary insults, resulting in acute or chronic dysfunction. Of particular importance for characterizing brainstem dysfunction and identifying the underlying etiology are a detailed clinical examination, MRI, neurophysiologic tests such as brainstem auditory evoked potentials, and an analysis of the cerebrospinal fluid. Detection of brainstem dysfunction is challenging but of utmost importance in comatose and deeply sedated patients both to guide therapy and to support outcome prediction. In the present review, we summarize the neuroanatomy, clinical syndromes, and diagnostic techniques of critical illness-associated brainstem dysfunction for the critical care setting. Keywords: Brainstem dysfunction, Brain injured patients, Intensive care unit, Sedation, Brainstem -

1 Physiology Week 4 – Reflexes
Physiology week 4 – Reflexes Reflexes Reflex arc Sensory organ Afferent neuron (enters cord via dorsal roots or cranial nerves) One or more synapses Efferent neuron (leaves the cord via ventral roots or cranial nerves) Effector Bell-Magendie law Spinal cord dorsal roots are sensory Spinal cord ventral roots are motor Monosynaptic reflexes Stretch reflex eg. knee jerk, ankle jerk Sense organ (muscle spindle) Ia fibre, afferent neuron (same as efferent nerve supply) One synapse (neurotransmitter is glutamate) Efferent neuron (motor neurone) Effector (muscle) Reciprocal innervation of muscle antagonist Muscle spindle fxn - parallel intrafusal fibres respond to stretch with different dynamic/static responses. Inverse stretch reflex Stretch can increase to a maximum, but once this point is reached, the muscle relaxes (following prolonged stretch or muscle contraction the contracted muscles suddenly relax) Sense organ (golgi apparatus) Afferent neuron (same as efferent nerve supply) One synapses (neurotransmitter is glutamate) Efferent neuron Effector (muscle) Clonus ‘regular rhythmic contraction of a muscle that is subjected to sudden maintained stretch’ Caused by increased gamma efferent discharge in response to hyperactive muscle spindles Polysynaptic reflexes Polysynaptic reflexes branch in a complex fashion One or more interneurons, interposed between the afferent and efferent neurones Number of synapses is variable Withdrawal reflex Sense organ (noxious stimulus skin or subcut) Afferent neuron (same as efferent nerve supply) Multiple synapses (neurotransmitter is glutamate) 1 Efferent neuron Effector (ipsilateral flexor contraction/extensor relaxation + crossed extensor response of opposite limb) Results in withdrawal of limb from stimulus Cross extensor response opposite limb Prepotency of the withdrawal reflex the reflex pre-empts spinal pathways from any other reflex activity occurring at the same time. -
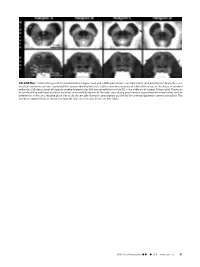
ON-LINE FIG 1. Selected Images of the Caudal Midbrain (Upper Row
ON-LINE FIG 1. Selected images of the caudal midbrain (upper row) and middle pons (lower row) from 4 of 13 total postmortem brains illustrate excellent anatomic contrast reproducibility across individual datasets. Subtle variations are present. Note differences in the shape of cerebral peduncles (24), decussation of superior cerebellar peduncles (25), and spinothalamic tract (12) in the midbrain of subject D (top right). These can be attributed to individual anatomic variation, some mild distortion of the brain stem during procurement at postmortem examination, and/or differences in the axial imaging plane not easily discernable during its prescription parallel to the anterior/posterior commissure plane. The numbers in parentheses in the on-line legends refer to structures in the On-line Table. AJNR Am J Neuroradiol ●:●●2019 www.ajnr.org E1 ON-LINE FIG 3. Demonstration of the dentatorubrothalamic tract within the superior cerebellar peduncle (asterisk) and rostral brain stem. A, Axial caudal midbrain image angled 10° anterosuperior to posteroinferior relative to the ACPC plane demonstrates the tract traveling the midbrain to reach the decussation (25). B, Coronal oblique image that is perpendicular to the long axis of the hippocam- pus (structure not shown) at the level of the ventral superior cerebel- lar decussation shows a component of the dentatorubrothalamic tract arising from the cerebellar dentate nucleus (63), ascending via the superior cerebellar peduncle to the decussation (25), and then enveloping the contralateral red nucleus (3). C, Parasagittal image shows the relatively long anteroposterior dimension of this tract, which becomes less compact and distinct as it ascends toward the thalamus. ON-LINE FIG 2. -

Brainstem Dysfunction in Critically Ill Patients
Benghanem et al. Critical Care (2020) 24:5 https://doi.org/10.1186/s13054-019-2718-9 REVIEW Open Access Brainstem dysfunction in critically ill patients Sarah Benghanem1,2 , Aurélien Mazeraud3,4, Eric Azabou5, Vibol Chhor6, Cassia Righy Shinotsuka7,8, Jan Claassen9, Benjamin Rohaut1,9,10† and Tarek Sharshar3,4*† Abstract The brainstem conveys sensory and motor inputs between the spinal cord and the brain, and contains nuclei of the cranial nerves. It controls the sleep-wake cycle and vital functions via the ascending reticular activating system and the autonomic nuclei, respectively. Brainstem dysfunction may lead to sensory and motor deficits, cranial nerve palsies, impairment of consciousness, dysautonomia, and respiratory failure. The brainstem is prone to various primary and secondary insults, resulting in acute or chronic dysfunction. Of particular importance for characterizing brainstem dysfunction and identifying the underlying etiology are a detailed clinical examination, MRI, neurophysiologic tests such as brainstem auditory evoked potentials, and an analysis of the cerebrospinal fluid. Detection of brainstem dysfunction is challenging but of utmost importance in comatose and deeply sedated patients both to guide therapy and to support outcome prediction. In the present review, we summarize the neuroanatomy, clinical syndromes, and diagnostic techniques of critical illness-associated brainstem dysfunction for the critical care setting. Keywords: Brainstem dysfunction, Brain injured patients, Intensive care unit, Sedation, Brainstem -

Cerebral Cortex
Cerebral Cortex • Research on the structure and function of the brain reveals that there are both specialized and diffuse areas of function • Motor and sensory areas are localized in discrete cortical areas called domains • Many higher mental functions such as memory and language appear to have overlapping domains and are more diffusely located • Broadmann areas are areas of localized function Cerebral Cortex - Generalizations • The cerebral cortex has three types of functional areas – Motor areas / control voluntary motor function – Sensory areas / provide conscious awareness of sensation – Association areas / act mainly to integrate diverse information for purposeful action • Each hemisphere is chiefly concerned with the sensory and motor functions of the opposite (contralateral) side of the body Motor Areas • Cortical areas controlling motor functions lie in the posterior part of the frontal lobes • Motor areas include the primary motor cortex, the premotor cortex, Broca’s area, and the front eye field Primary Motor Cortex • The primary motor cortex is located in the precentral gyrus of the frontal lobe of each hemisphere • Large neurons (pyramidal cells) in these gyri allow us to consciously control the precise or skill voluntary movements of our skeletal muscles Pyramidal cells • These long axons, which project to the spinal cord, form the massive Dendrites voluntary motor tracts called the pyramidal, or corticospinal tracts • All other descending motor tracts issue from brain stem nuclei and consists of chains of two, three, or