Molecular and Functional Evolution of the Spermatophyte Sesquiterpene Synthases
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gymnosperms the MESOZOIC: ERA of GYMNOSPERM DOMINANCE
Chapter 24 Gymnosperms THE MESOZOIC: ERA OF GYMNOSPERM DOMINANCE THE VASCULAR SYSTEM OF GYMNOSPERMS CYCADS GINKGO CONIFERS Pinaceae Include the Pines, Firs, and Spruces Cupressaceae Include the Junipers, Cypresses, and Redwoods Taxaceae Include the Yews, but Plum Yews Belong to Cephalotaxaceae Podocarpaceae and Araucariaceae Are Largely Southern Hemisphere Conifers THE LIFE CYCLE OF PINUS, A REPRESENTATIVE GYMNOSPERM Pollen and Ovules Are Produced in Different Kinds of Structures Pollination Replaces the Need for Free Water Fertilization Leads to Seed Formation GNETOPHYTES GYMNOSPERMS: SEEDS, POLLEN, AND WOOD THE ECOLOGICAL AND ECONOMIC IMPORTANCE OF GYMNOSPERMS The Origin of Seeds, Pollen, and Wood Seeds and Pollen Are Key Reproductive SUMMARY Innovations for Life on Land Seed Plants Have Distinctive Vegetative PLANTS, PEOPLE, AND THE Features ENVIRONMENT: The California Coast Relationships among Gymnosperms Redwood Forest 1 KEY CONCEPTS 1. The evolution of seeds, pollen, and wood freed plants from the need for water during reproduction, allowed for more effective dispersal of sperm, increased parental investment in the next generation and allowed for greater size and strength. 2. Seed plants originated in the Devonian period from a group called the progymnosperms, which possessed wood and heterospory, but reproduced by releasing spores. Currently, five lineages of seed plants survive--the flowering plants plus four groups of gymnosperms: cycads, Ginkgo, conifers, and gnetophytes. Conifers are the best known and most economically important group, including pines, firs, spruces, hemlocks, redwoods, cedars, cypress, yews, and several Southern Hemisphere genera. 3. The pine life cycle is heterosporous. Pollen strobili are small and seasonal. Each sporophyll has two microsporangia, in which microspores are formed and divide into immature male gametophytes while still retained in the microsporangia. -

The Glycosyltransferase Repertoire of the Spikemoss Selaginella Moellendorffiind a a Comparative Study of Its Cell Wall
Purdue University Purdue e-Pubs Department of Botany and Plant Pathology Faculty Department of Botany and Plant Pathology Publications 5-2-2012 The Glycosyltransferase Repertoire of the Spikemoss Selaginella moellendorffiind a a Comparative Study of Its Cell Wall. Jesper Harholt Iben Sørensen Cornell University Jonatan Fangel Alison Roberts University of Rhode Island William G.T. Willats See next page for additional authors Follow this and additional works at: http://docs.lib.purdue.edu/btnypubs Recommended Citation Harholt, Jesper; Sørensen, Iben; Fangel, Jonatan; Roberts, Alison; Willats, William G.T.; Scheller, Henrik Vibe; Petersen, Bent Larsen; Banks, Jo Ann; and Ulvskov, Peter, "The Glycosyltransferase Repertoire of the Spikemoss Selaginella moellendorffii and a Comparative Study of Its Cell Wall." (2012). Department of Botany and Plant Pathology Faculty Publications. Paper 15. http://dx.doi.org/10.1371/journal.pone.0035846 This document has been made available through Purdue e-Pubs, a service of the Purdue University Libraries. Please contact [email protected] for additional information. Authors Jesper Harholt, Iben Sørensen, Jonatan Fangel, Alison Roberts, William G.T. Willats, Henrik Vibe Scheller, Bent Larsen Petersen, Jo Ann Banks, and Peter Ulvskov This article is available at Purdue e-Pubs: http://docs.lib.purdue.edu/btnypubs/15 The Glycosyltransferase Repertoire of the Spikemoss Selaginella moellendorffii and a Comparative Study of Its Cell Wall Jesper Harholt1, Iben Sørensen1,2, Jonatan Fangel1, Alison Roberts3, William G. -
Plant Classification
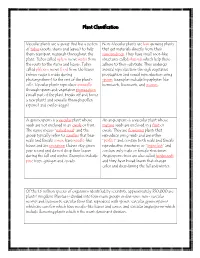
Plant Classification Vascular plants are a group that has a system Non-Vascular plants are low growing plants of tubes (roots, stems and leaves) to help that get materials directly from their them transport materials throughout the surroundings. They have small root-like plant. Tubes called xylem move water from structures called rhizoids which help them the roots to the stems and leaves. Tubes adhere to their substrate. They undergo called phloem move food from the leaves asexual reproduction through vegetative (where sugar is made during propagation and sexual reproduction using photosynthesis) to the rest of the plant’s spores. Examples include bryophytes like cells. Vascular plants reproduce asexually hornworts, liverworts, and mosses. through spores and vegetative propagation (small part of the plant breaks off and forms a new plant) and sexually through pollen (sperm) and ovules (eggs). A gymnosperm is a vascular plant whose An angiosperm is a vascular plant whose seeds are not enclosed in an ovule or fruit. mature seeds are enclosed in a fruit or The name means “naked seed” and the ovule. They are flowering plants that group typically refers to conifers that bear reproduce using seeds and are either male and female cones, have needle-like “perfect” and contain both male and female leaves and are evergreen (leaves stay green reproductive structures or “imperfect” and year round and do not drop their leaves contain only male or female structures. during the fall and winter. Examples include Angiosperm trees are also called hardwoods pine trees, ginkgos and cycads. and they have broad leaves that change color and drop during the fall and winter. -

B.Sc Botany (Sub.) I Group: a General Characters of Gymnosperm
1 B.Sc Botany (Sub.) I Group: A General Characters of Gymnosperm Gymnosperms are a small group of plants comprising only 70 genera and 725 living species. The word Gymnosperm was used by the Greek botanists Theophrastus in 300 B.C. for the plants with unprotected seeds. Gymnosperms are naked seeded plants. The gymnosperms are characterized by the following features: i. It shows the distinct alternation of generations. ii. Sporophytic generation is the dormant phase of the life cycle. The main plants are sporophyte and the gametophytes are dependent on it throughout. iii. The sporophytic plants are usually tall, woody, perennial trees or shrubs and differentiated into root, stem and leaves. iv. Root- Root is usually well developed tap root system. The stele in root is diarch or polyarch. v. Stem- stems are usually branched but unbranched in Cycas. vi. The leaves may be compound as in Cycas or simple as in Pinus. Internal structure: i. The vascular bundles in stem are arranged in a ring. The bundles are conjoint, collateral and open. ii. The secondary growth is effected by a cambial layer which produces secondary xylem and secondary phloem on the inner and outer side respectively. iii. The secondary wood forming distinct annual ring. Spring wood: It is made up of layer tracheids with the walls and layer lumen. Autumn wood: It is made up of compactly arranged smaller tracheids with thick walls and smaller lumen. iv. The wood may be manoxylic or pycnoxylic and may be monoxylic or polyxylic. Dr. Sanjeev Kumar Vidyarthi, dept. of Botany, Dr. L.K.V.D. -

A Global Analysis of the Distribution and Conservation Status Of
Journal of Biogeography (J. Biogeogr.) (2015) 42, 809–820 SYNTHESIS Fighting their last stand? A global analysis of the distribution and conservation status of gymnosperms Yann Fragniere1,Sebastien Betrisey2,3,Leonard Cardinaux1, Markus Stoffel4,5 and Gregor Kozlowski1,2* 1Natural History Museum Fribourg, CH-1700 ABSTRACT Fribourg, Switzerland, 2Department of Biology Aim Gymnosperms are often described as a marginal and threatened group, and Botanic Garden, University of Fribourg, members of which tend to be out-competed by angiosperms and which therefore CH-1700 Fribourg, Switzerland, 3Conservation Biogeography, Department of preferentially persist at higher latitudes and elevations. The aim of our synthesis Geosciences, University of Fribourg, CH-1700 was to test these statements by investigating the global latitudinal and elevational Fribourg, Switzerland, 4Dendrolab.ch, distribution of gymnosperms, as well as their conservation status, using all extant Institute of Geological Sciences, University of gymnosperm groups (cycads, gnetophytes, ginkgophytes and conifers). 5 Bern, CH-3012 Bern, Switzerland, Institute Location Worldwide. for Environmental Sciences, Climatic Change and Climate Impacts, University of Geneva, Methods We developed a database of 1014 species of gymnosperms containing CH-1227 Carouge, Switzerland latitudinal and elevational distribution data, as well as their global conservation status, as described in the literature. The 1014 species comprised 305 cycads, 101 gnetophytes, the only living representative of ginkgophytes, and 607 conifers. Generalized additive models, frequency histograms, kernel density estimations and distribution maps based on Takhtajan’s floristic regions were used. Results Although the diversity of gymnosperms decreases at equatorial lati- tudes, approximately 50% of the extant species occur primarily between the tropics. More than 43% of gymnosperms can occur at very low elevations (≤ 200 m a.s.l.). -

Name That Gymnosperm
A B C D This tree is found frequently This evergreen is found throughout This species is found at drier and This Wyoming tree is a remnant of throughout the eastern half of the state. The species is known for lower elevations compared to some the last ice age and is found exclu- Wyoming ranging from the Black often having serotinous cones that of the other trees pictured. This tree sively in the Black Hills of Wyoming Hills to the Laramie Mountains and only open to release seeds when the shares its name with the town in and South Dakota. It grows at higher the Bighorns. The tree is known cone is heated. Once the limbs are central Wyoming. elevations along riparian or wet areas for its ability to withstand the heat removed, this tree makes an excel- in its native habitat. of wildfires because of its thick, lent pole for teepees because of its reddish-colored bark. long, slender trunk. NAME THAT GYMNOSPERM Wyoming is host to many conifer (gymnosperm or “naked in hot, dry, and low elevations while others are found in cold and seed” plants including conifer, cycads, and ginkos) tree species. high elevations. Cones are an excellent tool that can be used for The cones in this quiz and their parent trees are found at different identification of evergreens. Match the tree to the photo. Good geographical locations across the Cowboy State. Some are found luck and keep an eye out for these trees this year! E F G H Known for having extremely flexible This species is probably better known This tree is found in most high- This tree grows at high elevations limbs, this tree is often found brav- as an ornamental in Wyoming but is elevation forests of Wyoming. -

Spermatophyte Flora of Liangzi Lake Wetland Nature Reserve
E3S Web of Conferences 143, 02040 (2 020) https://doi.org/10.1051/e3sconf/20 2014302040 ARFEE 2019 Spermatophyte Flora of Liangzi Lake Wetland Nature Reserve Xinyang Zhang, Shijing He*, Rong Tao, and Huan Dai Wuhan Institute of Design and Sciences, Wuhan 430205, China Abstract. Based on route and sample-plot survey, plant resources of Liangzi Lake Wetland Nature Reserve were investigated. The result showed that there were 503 species of spermatophyte belonging to 296 genera of 86 families. There were 5 species under national first and second level protection. The dominant families of spermatophyte contained 20 species and above. The dominant genera of spermatophyte contained 4 species and below. The 86 families of spermatophyte can be divided into 7 distribution types and 4 variants. Tropic distribution type was dominant, accounting for 70.83% in total (excluding cosmopolitans). The 296 genera of spermatophyte can be divided into 14 distribution types and 9 variants. Temperate elements were a little more than tropical elements, accounting for 50.84% and 49.16% in total (excluding cosmopolitans) respectively. Reserve had 3 Chinese endemic genera, reflecting certain ancient and relict. The purpose of the research is to provide background information and scientific basis for the protection, construction, management and rational utilization of plant resources in the reserve. 1 Preface temperature is 17℃, the annual average rainfall is 1663mm, the average sunshine hour is 2061 hours, and Flora refers to the sum of all plant species in a certain the frost free period is 270 days. The rain bearing area is region or country. It is the result of the development and 208500 hectares, and the annual average water level is evolution of the plant kingdom under certain natural and 17.81m. -

Curriculum Vitae
CURRICULUM VITAE ORCID ID: 0000-0003-0186-6546 Gar W. Rothwell Edwin and Ruth Kennedy Distinguished Professor Emeritus Department of Environmental and Plant Biology Porter Hall 401E T: 740 593 1129 Ohio University F: 740 593 1130 Athens, OH 45701 E: [email protected] also Courtesy Professor Department of Botany and PlantPathology Oregon State University T: 541 737- 5252 Corvallis, OR 97331 E: [email protected] Education Ph.D.,1973 University of Alberta (Botany) M.S., 1969 University of Illinois, Chicago (Biology) B.A., 1966 Central Washington University (Biology) Academic Awards and Honors 2018 International Organisation of Palaeobotany lifetime Honorary Membership 2014 Fellow of the Paleontological Society 2009 Distinguished Fellow of the Botanical Society of America 2004 Ohio University Distinguished Professor 2002 Michael A. Cichan Award, Botanical Society of America 1999-2004 Ohio University Presidential Research Scholar in Biomedical and Life Sciences 1993 Edgar T. Wherry Award, Botanical Society of America 1991-1992 Outstanding Graduate Faculty Award, Ohio University 1982-1983 Chairman, Paleobotanical Section, Botanical Society of America 1972-1973 University of Alberta Dissertation Fellow 1971 Paleobotanical (Isabel Cookson) Award, Botanical Society of America Positions Held 2011-present Courtesy Professor of Botany and Plant Pathology, Oregon State University 2008-2009 Visiting Senior Researcher, University of Alberta 2004-present Edwin and Ruth Kennedy Distinguished Professor of Environmental and Plant Biology, Ohio -

Diversity and Evolution of the Megaphyll in Euphyllophytes
G Model PALEVO-665; No. of Pages 16 ARTICLE IN PRESS C. R. Palevol xxx (2012) xxx–xxx Contents lists available at SciVerse ScienceDirect Comptes Rendus Palevol w ww.sciencedirect.com General palaeontology, systematics and evolution (Palaeobotany) Diversity and evolution of the megaphyll in Euphyllophytes: Phylogenetic hypotheses and the problem of foliar organ definition Diversité et évolution de la mégaphylle chez les Euphyllophytes : hypothèses phylogénétiques et le problème de la définition de l’organe foliaire ∗ Adèle Corvez , Véronique Barriel , Jean-Yves Dubuisson UMR 7207 CNRS-MNHN-UPMC, centre de recherches en paléobiodiversité et paléoenvironnements, 57, rue Cuvier, CP 48, 75005 Paris, France a r t i c l e i n f o a b s t r a c t Article history: Recent paleobotanical studies suggest that megaphylls evolved several times in land plant st Received 1 February 2012 evolution, implying that behind the single word “megaphyll” are hidden very differ- Accepted after revision 23 May 2012 ent notions and concepts. We therefore review current knowledge about diverse foliar Available online xxx organs and related characters observed in fossil and living plants, using one phylogenetic hypothesis to infer their origins and evolution. Four foliar organs and one lateral axis are Presented by Philippe Taquet described in detail and differ by the different combination of four main characters: lateral organ symmetry, abdaxity, planation and webbing. Phylogenetic analyses show that the Keywords: “true” megaphyll appeared at least twice in Euphyllophytes, and that the history of the Euphyllophytes Megaphyll four main characters is different in each case. The current definition of the megaphyll is questioned; we propose a clear and accurate terminology in order to remove ambiguities Bilateral symmetry Abdaxity of the current vocabulary. -

Late Devonian Spermatophyte Diversity and Paleoecology at Red Hill, North-Central Pennsylvania, U.S.A. Walter L
West Chester University Digital Commons @ West Chester University Geology & Astronomy Faculty Publications Geology & Astronomy 2010 Late Devonian spermatophyte diversity and paleoecology at Red Hill, north-central Pennsylvania, U.S.A. Walter L. Cressler III West Chester University, [email protected] Cyrille Prestianni Ben A. LePage Follow this and additional works at: http://digitalcommons.wcupa.edu/geol_facpub Part of the Geology Commons, Paleobiology Commons, and the Paleontology Commons Recommended Citation Cressler III, W.L., Prestianni, C., and LePage, B.A. 2010. Late Devonian spermatophyte diversity and paleoecology at Red Hill, north- central Pennsylvania, U.S.A. International Journal of Coal Geology 83, 91-102. This Article is brought to you for free and open access by the Geology & Astronomy at Digital Commons @ West Chester University. It has been accepted for inclusion in Geology & Astronomy Faculty Publications by an authorized administrator of Digital Commons @ West Chester University. For more information, please contact [email protected]. ARTICLE IN PRESS COGEL-01655; No of Pages 12 International Journal of Coal Geology xxx (2009) xxx–xxx Contents lists available at ScienceDirect International Journal of Coal Geology journal homepage: www.elsevier.com/locate/ijcoalgeo Late Devonian spermatophyte diversity and paleoecology at Red Hill, north-central Pennsylvania, USA Walter L. Cressler III a,⁎, Cyrille Prestianni b, Ben A. LePage c a Francis Harvey Green Library, 29 West Rosedale Avenue, West Chester University, West Chester, PA, 19383, USA b Université de Liège, Boulevard du Rectorat B18, Liège 4000 Belgium c The Academy of Natural Sciences, 1900 Benjamin Franklin Parkway, Philadelphia, PA, 19103 and PECO Energy Company, 2301 Market Avenue, S9-1, Philadelphia, PA 19103, USA article info abstract Article history: Early spermatophytes have been discovered at Red Hill, a Late Devonian (Famennian) fossil locality in north- Received 9 January 2009 central Pennsylvania, USA. -
Workshop on Alternation of Generations by Dana Krempels
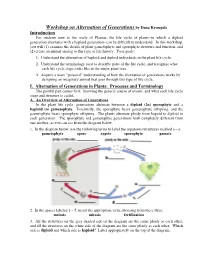
Workshop on Alternation of Generations by Dana Krempels Introduction For students new to the study of Plantae, the life cycle of plants--in which a diploid generation alternates with a haploid generation--can be difficult to understand. In this workshop, you will (1) examine the details of plant gametophyte and sporophyte structure and function, and (2) create an animal analog to this type of life history. Your goals: 1. Understand the alternation of haploid and diploid individuals in the plant life cycle. 2. Understand the terminology used to describe parts of the life cycle, and recognize what each life cycle stage looks like in the major plant taxa. 3. Acquire a more "personal" understanding of how the alternation of generations works by designing an imaginary animal that goes through this type of life cycle. I. Alternation of Generations in Plants: Processes and Terminology The painful part comes first: knowing the general course of events, and what each life cycle stage and structure is called. A. An Overview of Alternation of Generations In the plant life cycle, generations alternate between a diploid (2n) sporophyte and a haploid (n) gametophyte. Essentially, the sporophyte bears gametophyte offspring, and the gametophyte bears sporophyte offspring. The plants alternate ploidy from hapoid to diploid in each generation. The sporophyte and gametophye generations look completely different from one another, as you can see from the diagram below. 1. In the diagram below, use the following terms to label the organisms/structures marked a – e. gametophyte spore zygote sporophyte gamete 2. In the spaces labeled 1 – 5, insert the appropriate term, choosing from these three: meiosis mitosis fertilization 3. -

Protista and Fungi
Biology 2 – Practicum 2 1 Biology 2 Lab Packet For Practical 2 Biology 2 – Practicum 2 2 PLANT CLASSIFICATION: Domain: Eukarya (Supergroup: Archaeplastida) Kingdom: Plantae Nonvascular Seedless Plants Division: Hepatophyta - Liverworts Division: Bryophyta - Mosses Vascular Seedless Plants Division: Lycophyta - Club Mosses Division: Pterophyta (Psilophyta) - Whisk ferns (Sphenophyta) - Horsetails (Pterophyta) - Ferns Red Algae Archaeplastida Chlorophytes Charophyceans Liverworts Plants Mosses Hornworts Lycophytes Pterophytes Gymnosperms INTRODUCTION TO PLANTS Angiosperms The kingdom Plantae includes about twelve divisions. They are placed in the clade Archaeplastida along with the green algae and charophytes. They are all eukaryotic and multicellular with distinct cell walls. Photosynthetic pigments occur in organelles called plastids. Plants have adapted to the terrestrial environment with an increase in structural complexity. Many plants have developed organs for anchorage, conduction, support, and photosynthesis. Reproduction is primarily sexual, with an alternation of generation of haploid and diploid generations. The sporophyte generation becomes increasingly predominant as plants evolve. The nonvascular plants lack conductive tissue and are limited to a specific range of terrestrial habitats. These plants display two adaptations that first made the move onto land possible. They possess a waxy cuticle to reduce water loss and their gametes develop within gametangia for protection of the embryo. These plants are limited in range because they require water for reproduction. They lack vascular tissue, which means they must live in moist environments and lack woody structures for support therefore they grow low to the ground. Station 1 – Kingdom Planate 1. What supergroup do they belong to and what characteristic are responsible for this positioning? 2. What characteristics are specific to plants? 3.