Community of Actinomycetes in 42 Species of Animal Feces
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Tessaracoccus Arenae Sp. Nov., Isolated from Sea Sand
TAXONOMIC DESCRIPTION Thongphrom et al., Int J Syst Evol Microbiol 2017;67:2008–2013 DOI 10.1099/ijsem.0.001907 Tessaracoccus arenae sp. nov., isolated from sea sand Chutimon Thongphrom,1 Jong-Hwa Kim,1 Nagamani Bora2,* and Wonyong Kim1,* Abstract A Gram-stain positive, non-spore-forming, non-motile, facultatively anaerobic bacterial strain, designated CAU 1319T, was isolated from sea sand and the strain’s taxonomic position was investigated using a polyphasic approach. Strain CAU 1319T grew optimally at 30 C and at pH 7.5 in the presence of 2 % (w/v) NaCl. Phylogenetic analysis, based on the 16S rRNA gene sequence, revealed that strain CAU 1319T belongs to the genus Tessaracoccus, and is closely related to Tessaracoccus lapidicaptus IPBSL-7T (similarity 97.69 %), Tessaracoccus bendigoensis Ben 106T (similarity 95.64 %) and Tessaracoccus T T flavescens SST-39 (similarity 95.84 %). Strain CAU 1319 had LL-diaminopimelic acid as the diagnostic diamino acid in the cell-wall peptidoglycan, MK-9 (H4) as the predominant menaquinone, and anteiso-C15 : 0 as the major fatty acid. The polar lipids consisted of phosphatidylglycerol, phosphatidylinositol, two unidentified aminolipids, three unidentified phospholipids and one unidentified glycolipid. Predominant polyamines were spermine and spermidine. The DNA–DNA hybridization value between strain CAU 1319T and T. lapidicaptus IPBSL-7T was 24 %±0.2. The DNA G+C content of the novel strain was 69.5 mol %. On the basis of phenotypic and chemotaxonomic properties, as well as phylogenetic relatedness, strain CAU 1319Tshould be classified as a novel species of the genus Tessaracoccus, for which the name Tessaracoccus arenae sp. -

Corynebacterium Sp.|NML98-0116
1 Limnochorda_pilosa~GCF_001544015.1@NZ_AP014924=Bacteria-Firmicutes-Limnochordia-Limnochordales-Limnochordaceae-Limnochorda-Limnochorda_pilosa 0,9635 Ammonifex_degensii|KC4~GCF_000024605.1@NC_013385=Bacteria-Firmicutes-Clostridia-Thermoanaerobacterales-Thermoanaerobacteraceae-Ammonifex-Ammonifex_degensii 0,985 Symbiobacterium_thermophilum|IAM14863~GCF_000009905.1@NC_006177=Bacteria-Firmicutes-Clostridia-Clostridiales-Symbiobacteriaceae-Symbiobacterium-Symbiobacterium_thermophilum Varibaculum_timonense~GCF_900169515.1@NZ_LT827020=Bacteria-Actinobacteria-Actinobacteria-Actinomycetales-Actinomycetaceae-Varibaculum-Varibaculum_timonense 1 Rubrobacter_aplysinae~GCF_001029505.1@NZ_LEKH01000003=Bacteria-Actinobacteria-Rubrobacteria-Rubrobacterales-Rubrobacteraceae-Rubrobacter-Rubrobacter_aplysinae 0,975 Rubrobacter_xylanophilus|DSM9941~GCF_000014185.1@NC_008148=Bacteria-Actinobacteria-Rubrobacteria-Rubrobacterales-Rubrobacteraceae-Rubrobacter-Rubrobacter_xylanophilus 1 Rubrobacter_radiotolerans~GCF_000661895.1@NZ_CP007514=Bacteria-Actinobacteria-Rubrobacteria-Rubrobacterales-Rubrobacteraceae-Rubrobacter-Rubrobacter_radiotolerans Actinobacteria_bacterium_rbg_16_64_13~GCA_001768675.1@MELN01000053=Bacteria-Actinobacteria-unknown_class-unknown_order-unknown_family-unknown_genus-Actinobacteria_bacterium_rbg_16_64_13 1 Actinobacteria_bacterium_13_2_20cm_68_14~GCA_001914705.1@MNDB01000040=Bacteria-Actinobacteria-unknown_class-unknown_order-unknown_family-unknown_genus-Actinobacteria_bacterium_13_2_20cm_68_14 1 0,9803 Thermoleophilum_album~GCF_900108055.1@NZ_FNWJ01000001=Bacteria-Actinobacteria-Thermoleophilia-Thermoleophilales-Thermoleophilaceae-Thermoleophilum-Thermoleophilum_album -

Study of Dental Fluorosis in Subjects Related to a Phosphatic Fertilizer
Indian Journal of Geo Marine Sciences Vol. 46 (06), June 2017, pp. 1116-1127 Diversity and enzymatic profile of bacterial flora in the gut of an estuarine fish, Mugil jerdoni Ankita A. Datta, Amit K. Sharma, Rahul Kundu & Satya P. Singh* UGC-CAS Department of Biosciences, Saurashtra University, Rajkot 360 005, India *[E-mail: [email protected]] Received 12 August 2015 ; revised 07 December 2015 In order to examine the bacterial diversity and enzymatic potential, the isolates were screened for the amylolytic, cellulolytic, lipolytic and proteolytic activities using selective media. Significant proportion of the isolates (44%) exhibited lipase activity, while only few (11%) had protease activity. The 16S rRNA gene sequence analysis revealed that most of the isolates related to the genera Bacillus, Acinetobacter, Staphylococcus, Aeromonas, Psychrobacter, Dietzia and Isoptericola. Examined bacteria displayed significant tolerance against varying salt concentrations. Most of the isolates displayed antagonism against 10 selected target organisms. Bacteria were also assessed for their resistance and sensitivity against different antibiotics. Staphylococcus epidermis MJMG8.1 and Dietzia sp. MJMG8.2 grew significantly in the presence of different organic solvents. [Key Words: Microbial enzymes, antibiotic resistance, 16S rRNA sequencing, phylogeny, microbial diversity, fish-gut microflora] Introduction The culturable bacterial flora in the gut of Staphylococcus, unidentified anaerobes and yeast are freshwater and marine water fishes has been explored reported to produce exogenous enzymes. Amylases, in limited sense 1. Bacterial population in the fish gut proteases, lipases, cellulases, chitinases and few is governed by various environmental and intrinsic others have been produced by the gut flora 5, 7. factors such as species, developmental stage, feeding Studies have revealed that the gut microflora prevents strategy, structure of the digestive system and other the establishment of opportunistic pathogens in the physiological factors 2. -

Extensive Microbial Diversity Within the Chicken Gut Microbiome Revealed by Metagenomics and Culture
Extensive microbial diversity within the chicken gut microbiome revealed by metagenomics and culture Rachel Gilroy1, Anuradha Ravi1, Maria Getino2, Isabella Pursley2, Daniel L. Horton2, Nabil-Fareed Alikhan1, Dave Baker1, Karim Gharbi3, Neil Hall3,4, Mick Watson5, Evelien M. Adriaenssens1, Ebenezer Foster-Nyarko1, Sheikh Jarju6, Arss Secka7, Martin Antonio6, Aharon Oren8, Roy R. Chaudhuri9, Roberto La Ragione2, Falk Hildebrand1,3 and Mark J. Pallen1,2,4 1 Quadram Institute Bioscience, Norwich, UK 2 School of Veterinary Medicine, University of Surrey, Guildford, UK 3 Earlham Institute, Norwich Research Park, Norwich, UK 4 University of East Anglia, Norwich, UK 5 Roslin Institute, University of Edinburgh, Edinburgh, UK 6 Medical Research Council Unit The Gambia at the London School of Hygiene and Tropical Medicine, Atlantic Boulevard, Banjul, The Gambia 7 West Africa Livestock Innovation Centre, Banjul, The Gambia 8 Department of Plant and Environmental Sciences, The Alexander Silberman Institute of Life Sciences, Edmond J. Safra Campus, Hebrew University of Jerusalem, Jerusalem, Israel 9 Department of Molecular Biology and Biotechnology, University of Sheffield, Sheffield, UK ABSTRACT Background: The chicken is the most abundant food animal in the world. However, despite its importance, the chicken gut microbiome remains largely undefined. Here, we exploit culture-independent and culture-dependent approaches to reveal extensive taxonomic diversity within this complex microbial community. Results: We performed metagenomic sequencing of fifty chicken faecal samples from Submitted 4 December 2020 two breeds and analysed these, alongside all (n = 582) relevant publicly available Accepted 22 January 2021 chicken metagenomes, to cluster over 20 million non-redundant genes and to Published 6 April 2021 construct over 5,500 metagenome-assembled bacterial genomes. -

Halophilic Microorganisms Are Responsible for the Rosy Discolouration of Saline Environments in Three Historical Buildings with Mural Paintings
Halophilic Microorganisms Are Responsible for the Rosy Discolouration of Saline Environments in Three Historical Buildings with Mural Paintings Jo¨ rg D. Ettenauer1*, Valme Jurado2, Guadalupe Pin˜ ar1, Ana Z. Miller2,3, Markus Santner4, Cesareo Saiz-Jimenez2, Katja Sterflinger1 1 VIBT-BOKU, University of Natural Resources and Life Sciences, Department of Biotechnology, Vienna, Austria, 2 Instituto de Recursos Naturales y Agrobiologia, IRNAS- CSIC, Sevilla, Spain, 3 CEPGIST/CERENA, Instituto Superior Te´cnico, Universidade de Lisboa, Lisboa, Portugal, 4 Bundesdenkmalamt, Abteilung fu¨r Konservierung und Restaurierung, Vienna, Austria Abstract A number of mural paintings and building materials from monuments located in central and south Europe are characterized by the presence of an intriguing rosy discolouration phenomenon. Although some similarities were observed among the bacterial and archaeal microbiota detected in these monuments, their origin and nature is still unknown. In order to get a complete overview of this biodeterioration process, we investigated the microbial communities in saline environments causing the rosy discolouration of mural paintings in three Austrian historical buildings using a combination of culture- dependent and -independent techniques as well as microscopic techniques. The bacterial communities were dominated by halophilic members of Actinobacteria, mainly of the genus Rubrobacter. Representatives of the Archaea were also detected with the predominating genera Halobacterium, Halococcus and Halalkalicoccus. Furthermore, halophilic bacterial strains, mainly of the phylum Firmicutes, could be retrieved from two monuments using special culture media. Inoculation of building materials (limestone and gypsum plaster) with selected isolates reproduced the unaesthetic rosy effect and biodeterioration in the laboratory. Citation: Ettenauer JD, Jurado V, Pin˜ar G, Miller AZ, Santner M, et al. -

Table S5. the Information of the Bacteria Annotated in the Soil Community at Species Level
Table S5. The information of the bacteria annotated in the soil community at species level No. Phylum Class Order Family Genus Species The number of contigs Abundance(%) 1 Firmicutes Bacilli Bacillales Bacillaceae Bacillus Bacillus cereus 1749 5.145782459 2 Bacteroidetes Cytophagia Cytophagales Hymenobacteraceae Hymenobacter Hymenobacter sedentarius 1538 4.52499338 3 Gemmatimonadetes Gemmatimonadetes Gemmatimonadales Gemmatimonadaceae Gemmatirosa Gemmatirosa kalamazoonesis 1020 3.000970902 4 Proteobacteria Alphaproteobacteria Sphingomonadales Sphingomonadaceae Sphingomonas Sphingomonas indica 797 2.344876284 5 Firmicutes Bacilli Lactobacillales Streptococcaceae Lactococcus Lactococcus piscium 542 1.594633558 6 Actinobacteria Thermoleophilia Solirubrobacterales Conexibacteraceae Conexibacter Conexibacter woesei 471 1.385742446 7 Proteobacteria Alphaproteobacteria Sphingomonadales Sphingomonadaceae Sphingomonas Sphingomonas taxi 430 1.265115184 8 Proteobacteria Alphaproteobacteria Sphingomonadales Sphingomonadaceae Sphingomonas Sphingomonas wittichii 388 1.141545794 9 Proteobacteria Alphaproteobacteria Sphingomonadales Sphingomonadaceae Sphingomonas Sphingomonas sp. FARSPH 298 0.876754244 10 Proteobacteria Alphaproteobacteria Sphingomonadales Sphingomonadaceae Sphingomonas Sorangium cellulosum 260 0.764953367 11 Proteobacteria Deltaproteobacteria Myxococcales Polyangiaceae Sorangium Sphingomonas sp. Cra20 260 0.764953367 12 Proteobacteria Alphaproteobacteria Sphingomonadales Sphingomonadaceae Sphingomonas Sphingomonas panacis 252 0.741416341 -

Raineyella Antarctica Gen. Nov., Sp. Nov., a Psychrotolerant, D-Amino
International Journal of Systematic and Evolutionary Microbiology (2016), 66, 5529–5536 DOI 10.1099/ijsem.0.001552 Raineyella antarctica gen. nov., sp. nov., a psychrotolerant, D-amino-acid-utilizing anaerobe isolated from two geographic locations of the Southern Hemisphere Elena Vladimirovna Pikuta,1 Rodolfo Javier Menes,2 Alisa Michelle Bruce,3† Zhe Lyu,4 Nisha B. Patel,5 Yuchen Liu,6 Richard Brice Hoover,1 Hans-Jürgen Busse,7 Paul Alexander Lawson5 and William Barney Whitman4 Correspondence 1Department of Mathematical, Computer and Natural Sciences, Athens State University, Athens, Elena Vladimirovna Pikuta AL 35611, USA [email protected] 2Catedra de Microbiología, Facultad de Química y Facultad de Ciencias, UDELAR, 11800 or Montevideo, Uruguay [email protected] 3Biology Department, University of Alabama in Huntsville, Huntsville, AL 35899, USA 4Microbiology Department, University of Georgia in Athens, Athens, GA 30602, USA 5Department of Microbiology and Plant Biology, University of Oklahoma, Norman, OK 73019, USA 6Department of Biological Sciences, Louisiana State University, Baton Rouge, LA 70803, USA 7Institut für Mikrobiologie - Veterinarmedizinische€ Universitat€ Wien, A-1210 Wien, Austria A Gram-stain-positive bacterium, strain LZ-22T, was isolated from a rhizosphere of moss Leptobryum sp. collected at the shore of Lake Zub in Antarctica. Cells were motile, straight or pleomorphic rods with sizes of 0.6–1.0Â3.5–10 µm. The novel isolate was a facultatively anaerobic, catalase-positive, psychrotolerant mesophile. Growth was observed at 3–41 C (optimum 24–28 C), with 0–7 % (w/v) NaCl (optimum 0.25 %) and at pH 4.0–9.0 (optimum pH 7.8). The quinone system of strain LZ-22T possessed predominately menaquinone MK-9(H4). -

Aestuariimicrobium Ganziense Sp. Nov., a New Gram-Positive Bacterium Isolated from Soil in the Ganzi Tibetan Autonomous Prefecture, China
Aestuariimicrobium ganziense sp. nov., a new Gram-positive bacterium isolated from soil in the Ganzi Tibetan Autonomous Prefecture, China Yu Geng Yunnan University Jiang-Yuan Zhao Yunnan University Hui-Ren Yuan Yunnan University Le-Le Li Yunnan University Meng-Liang Wen yunnan university Ming-Gang Li yunnan university Shu-Kun Tang ( [email protected] ) Yunnan Institute of Microbiology, Yunnan University https://orcid.org/0000-0001-9141-6244 Research Article Keywords: Aestuariimicrobium ganziense sp. nov., Chemotaxonomy, 16S rRNA sequence analysis Posted Date: February 11th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-215613/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Version of Record: A version of this preprint was published at Archives of Microbiology on March 12th, 2021. See the published version at https://doi.org/10.1007/s00203-021-02261-2. Page 1/11 Abstract A novel Gram-stain positive, oval shaped and non-agellated bacterium, designated YIM S02566T, was isolated from alpine soil in Shadui Towns, Ganzi County, Ganzi Tibetan Autonomous Prefecture, Sichuan Province, PR China. Growth occurred at 23–35°C (optimum, 30°C) in the presence of 0.5-4 % (w/v) NaCl (optimum, 1%) and at pH 7.0–8.0 (optimum, pH 7.0). The phylogenetic analysis based on 16S rRNA gene sequence revealed that strain YIM S02566T was most closely related to the genus Aestuariimicrobium, with Aestuariimicrobium kwangyangense R27T and Aestuariimicrobium soli D6T as its closest relative (sequence similarities were 96.3% and 95.4%, respectively). YIM S02566T contained LL-diaminopimelic acid in the cell wall. -

Marine Rare Actinomycetes: a Promising Source of Structurally Diverse and Unique Novel Natural Products
Review Marine Rare Actinomycetes: A Promising Source of Structurally Diverse and Unique Novel Natural Products Ramesh Subramani 1 and Detmer Sipkema 2,* 1 School of Biological and Chemical Sciences, Faculty of Science, Technology & Environment, The University of the South Pacific, Laucala Campus, Private Mail Bag, Suva, Republic of Fiji; [email protected] 2 Laboratory of Microbiology, Wageningen University & Research, Stippeneng 4, 6708 WE Wageningen, The Netherlands * Correspondence: [email protected]; Tel.: +31-317-483113 Received: 7 March 2019; Accepted: 23 April 2019; Published: 26 April 2019 Abstract: Rare actinomycetes are prolific in the marine environment; however, knowledge about their diversity, distribution and biochemistry is limited. Marine rare actinomycetes represent a rather untapped source of chemically diverse secondary metabolites and novel bioactive compounds. In this review, we aim to summarize the present knowledge on the isolation, diversity, distribution and natural product discovery of marine rare actinomycetes reported from mid-2013 to 2017. A total of 97 new species, representing 9 novel genera and belonging to 27 families of marine rare actinomycetes have been reported, with the highest numbers of novel isolates from the families Pseudonocardiaceae, Demequinaceae, Micromonosporaceae and Nocardioidaceae. Additionally, this study reviewed 167 new bioactive compounds produced by 58 different rare actinomycete species representing 24 genera. Most of the compounds produced by the marine rare actinomycetes present antibacterial, antifungal, antiparasitic, anticancer or antimalarial activities. The highest numbers of natural products were derived from the genera Nocardiopsis, Micromonospora, Salinispora and Pseudonocardia. Members of the genus Micromonospora were revealed to be the richest source of chemically diverse and unique bioactive natural products. -

Emergence of Carbapenem-Resistant Pseudomonas Aeruginosa and Acinetobacter Baumannii Clinical Isolates Collected from Some Libyan Hospitals
UNIVERSITE DE TUNIS EL MANAR Ecole Doctorale des Sciences Biologiques et AIX-MARSEILLE UNIVERSITE Ecole Doctorale des Sciences de la Vie et de la Santé THESE DE DOCTORAT EN COTUTELLE En Vue de l’obtention de grade de Docteur de l’Université de Tunis et d’Aix-Marseille Université Spécialités: Microbiologie / Pathologie humaine et maladies infectieuses Déterminisme du support moléculaire et de l’épidémiologie de la résistance aux ȕ-lactamines chez des bacilles à Gram négatif isolés dans des hôpitaux tunisiens et libyens Présentée par: Najla MATHLOUTHI Composition du Jury: Pr. Imane ZOUARI Université Tunis el Manar Président de Jury Dr. Marie KEMPF Université d’Angers Rapporteur Dr. Taoufik GHRAIRI Université Tunis el Manar Rapporteur Pr. Philippe COLSON Université d’Aix-Marseille Examinateur Pr. Mr. Jean-Marc ROLAIN Université d’Aix-Marseille Directeur de Thèse Dr. Chedly CHOUCHANI Université de Carthage Directeur de Thèse (8 Avril 2017) SOMMAIRE AVANT PROPOS ...................................................................... 3 RESUME .................................................................................... 4 SUMMARY ................................................................................ 5 INTRODUCTION ..................................................................... 7 CHAPITRE I: Revue: L’émergence et la dissémination des carbapénèmases produites par les bacilles à Gram négatifs dans les pays du bassin méditerranéen ............................................... 15 Article 1: Prevalence and emergence of -

THE LAKE VOSTOK Ferran Romero Blanch
MICROBIOLOGY OF A SUBGLACIAL LAKE: THE LAKE VOSTOK Ferran Romero Blanch OBJECTIVES INTRODUCTION: Lake Vostok is the largest and deepest The aim of the present review is to describe subglacial body of water in Antarctica. It has the communities present in a subglacial lake an area of 14000Km2 and a volume of (the Lake Vostok). 5600Km3. Nowadays, it is accepted that it has been buried under glacial ice for 14-15 millions It is also objective of this review to describe of years (Figure 1). Discovered between 1950 the use of ice-binding proteins as a and 1960, it has been recently reported that it method to survive in extremely cold forms a complex ecosystem with both environments. prokaryotic and eukaryotic microorganisms1,2. The living forms here have evolved to survive in an extremely oligotrophic, Main characteristics of Lake Vostok hyperbaric, cold and dark Temperature: -2°C environment. Pressure: 350atm (as it is covered by a 4- kilometer-thick layer of glacial ice) Nutrient availability: very slow Figure 1. Location of lake Vostok under Antarctic glacial ice1. Light: absent The ice above Lake Vostok freezes forming a 220m layer of accreted ice. It is generally accepted that this 220m layer of accreted ice reflects the contents into the lake and, because of this, it has been under study during the last 30 years. Ice accreting near the embayment has higher concentrations of ions, biomass and solid inclusions and it is known as type I accretion ice. The ice that forms over open water contains lower concentrations of biomass and ions and has been termed type II accretion ice3 (Figure 2). -
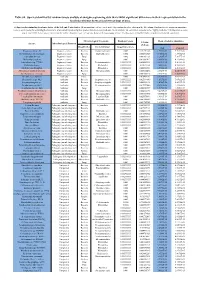
Table S8. Species Identified by Random Forests Analysis of Shotgun Sequencing Data That Exhibit Significant Differences In
Table S8. Species identified by random forests analysis of shotgun sequencing data that exhibit significant differences in their representation in the fecal microbiomes between each two groups of mice. (a) Species discriminating fecal microbiota of the Soil and Control mice. Mean importance of species identified by random forest are shown in the 5th column. Random forests assigns an importance score to each species by estimating the increase in error caused by removing that species from the set of predictors. In our analysis, we considered a species to be “highly predictive” if its importance score was at least 0.001. T-test was performed for the relative abundances of each species between the two groups of mice. P-values were at least 0.05 to be considered statistically significant. Microbiological Taxonomy Random Forests Mean of relative abundance P-Value Species Microbiological Function (T-Test) Classification Bacterial Order Importance Score Soil Control Rhodococcus sp. 2G Engineered strain Bacteria Corynebacteriales 0.002 5.73791E-05 1.9325E-05 9.3737E-06 Herminiimonas arsenitoxidans Engineered strain Bacteria Burkholderiales 0.002 0.005112829 7.1580E-05 1.3995E-05 Aspergillus ibericus Engineered strain Fungi 0.002 0.001061181 9.2368E-05 7.3057E-05 Dichomitus squalens Engineered strain Fungi 0.002 0.018887472 8.0887E-05 4.1254E-05 Acinetobacter sp. TTH0-4 Engineered strain Bacteria Pseudomonadales 0.001333333 0.025523638 2.2311E-05 8.2612E-06 Rhizobium tropici Engineered strain Bacteria Rhizobiales 0.001333333 0.02079554 7.0081E-05 4.2000E-05 Methylocystis bryophila Engineered strain Bacteria Rhizobiales 0.001333333 0.006513543 3.5401E-05 2.2044E-05 Alteromonas naphthalenivorans Engineered strain Bacteria Alteromonadales 0.001 0.000660472 2.0747E-05 4.6463E-05 Saccharomyces cerevisiae Engineered strain Fungi 0.001 0.002980726 3.9901E-05 7.3043E-05 Bacillus phage Belinda Antibiotic Phage 0.002 0.016409765 6.8789E-07 6.0681E-08 Streptomyces sp.