QKI Is a Critical Pre-Mrna Alternative Splicing Regulator of Cardiac Myofibrillogenesis and Contractile Function
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
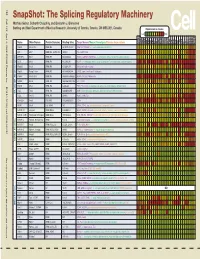
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

The Long Non-Coding RNA Gomafu Is Acutely Regulated in Response to Neuronal Activation and Involved in Schizophrenia-Associated Alternative Splicing
Molecular Psychiatry (2014) 19, 486–494 & 2014 Macmillan Publishers Limited All rights reserved 1359-4184/14 www.nature.com/mp ORIGINAL ARTICLE The long non-coding RNA Gomafu is acutely regulated in response to neuronal activation and involved in schizophrenia-associated alternative splicing G Barry1, JA Briggs2, DP Vanichkina1, EM Poth3, NJ Beveridge4,5, VS Ratnu6, SP Nayler2, K Nones7,JHu8, TW Bredy6, S Nakagawa9, F Rigo10, RJ Taft1, MJ Cairns4,5, S Blackshaw3, EJ Wolvetang2 and JS Mattick1,11,12 Schizophrenia (SZ) is a complex disease characterized by impaired neuronal functioning. Although defective alternative splicing has been linked to SZ, the molecular mechanisms responsible are unknown. Additionally, there is limited understanding of the early transcriptomic responses to neuronal activation. Here, we profile these transcriptomic responses and show that long non-coding RNAs (lncRNAs) are dynamically regulated by neuronal activation, including acute downregulation of the lncRNA Gomafu, previously implicated in brain and retinal development. Moreover, we demonstrate that Gomafu binds directly to the splicing factors QKI and SRSF1 (serine/arginine-rich splicing factor 1) and dysregulation of Gomafu leads to alternative splicing patterns that resemble those observed in SZ for the archetypal SZ-associated genes DISC1 and ERBB4. Finally, we show that Gomafu is downregulated in post-mortem cortical gray matter from the superior temporal gyrus in SZ. These results functionally link activity-regulated lncRNAs and alternative splicing in neuronal function and suggest that their dysregulation may contribute to neurological disorders. Molecular Psychiatry (2014) 19, 486–494; doi:10.1038/mp.2013.45; published online 30 April 2013 Keywords: alternative splicing; Gomafu; neuronal activation; quaking homolog; schizophrenia INTRODUCTION LncRNAs are arbitrarily defined as transcripts longer than 200 bp 10 Schizophrenia (SZ) disorder, a debilitating mental disorder that possess no protein-coding capacity. -

The Emerging Roles of the RNA Binding Protein QKI in Cardiovascular Development and Function
fcell-09-668659 June 14, 2021 Time: 13:17 # 1 MINI REVIEW published: 16 June 2021 doi: 10.3389/fcell.2021.668659 The Emerging Roles of the RNA Binding Protein QKI in Cardiovascular Development and Function Xinyun Chen1,2†, Jianwen Yin3†, Dayan Cao1, Deyong Xiao1, Zhongjun Zhou4, Ying Liu1* and Weinian Shou1* 1 Department of Pediatrics, Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN, United States, 2 Guangdong Key Laboratory for Genome Stability and Human Disease Prevention, Department of Biochemistry and Molecular Biology, School of Basic Medical Sciences, Shenzhen University, Shenzhen, China, 3 Department of Foot, Ankle and Hand Surgery, Shenzhen Second People’s Hospital, First Affiliated Hospital of Shenzhen University, Shenzhen, China, 4 Faculty of Medicine, School of Biomedical Sciences, The University of Hong Kong, Hong Kong Edited by: Janine M. Van Gils, Leiden University Medical Center, RNA binding proteins (RBPs) have a broad biological and physiological function and Netherlands are critical in regulating pre-mRNA posttranscriptional processing, intracellular migration, Reviewed by: Jianbo Wang, and mRNA stability. QKI, also known as Quaking, is a member of the signal transduction University of Alabama at Birmingham, and activation of RNA (STAR) family, which also belongs to the heterogeneous nuclear United States ribonucleoprotein K- (hnRNP K-) homology domain protein family. There are three Wuqiang Zhu, Mayo Clinic Arizona, United States major alternatively spliced isoforms, QKI-5, QKI-6, and QKI-7, differing in carboxy- *Correspondence: terminal domains. They share a common RNA binding property, but each isoform can Ying Liu regulate pre-mRNA splicing, transportation or stability differently in a unique cell type- [email protected] Weinian Shou specific manner. -

A Fast, Non-Radioactive Method to Identify in Vivo Targets of RNA-Binding Proteins
bioRxiv preprint doi: https://doi.org/10.1101/158410; this version posted July 1, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. uvCLAP: a fast, non-radioactive method to identify in vivo targets of RNA-binding proteins Daniel Maticzka1,4, Ibrahim Avsar Ilik2,4, Tugce Aktas2,4, Rolf Backofen1,3,# and Asifa Akhtar2,#* 1 Bioinformatics Group, Department of Computer Science, University of Freiburg, Freiburg, Germany 2 Max Planck Institute of Immunobiology and Epigenetics, 79108 Freiburg im Breisgau, Germany 3 Centre for Biological Signalling Studies (BIOSS), University of Freiburg, Freiburg, Germany 4 These authors contributed equally to this work # Corresponding author [email protected] Phone: +49 761 203 7461 Fax: +49 761 203 7462 # Corresponding author, * Lead contact [email protected] Phone: +49 (0)7615108565 Fax: +49 (0)7615108566 1 bioRxiv preprint doi: https://doi.org/10.1101/158410; this version posted July 1, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract RNA-binding proteins (RBPs) play important and essential roles in eukaryotic gene expression regulating splicing, localization, translation and stability of mRNAs. Understanding the exact contribution of RBPs to gene regulation is crucial as many RBPs are frequently mis-regulated in several neurological diseases and certain cancers. While recently developed techniques provide binding sites of RBPs, they are labor-intensive and generally rely on radioactive labeling of RNA. With more than 1,000 RBPs in a human cell, it is imperative to develop easy, robust, reproducible and high-throughput methods to determine in vivo targets of RBPs. -

Alternatively Spliced Genes
1 Alternatively Spliced Genes Jane Y. Wu1,2,LiyaYuan1 and Necat Havlioglu1 1Washington University School of Medicine, St. Louis, MO, USA 2John F. Kennedy Center for Research on Human Development, Vanderbilt University Medical Center, Nashville, TN, USA 1 Pre-mRNA Splicing and Splicing Machinery 3 1.1 Splicing Machinery: Spliceosome 3 1.2 Splicing Signals 4 1.3 Spliceosomal UsnRNP Biogenesis 12 1.4 Spliceosome Assembly 13 1.5 Biochemical Mechanisms of pre-mRNA Splicing 17 2 Alternative pre-mRNA Splicing 17 2.1 Alternative Splicing and its Role in Regulating Gene Activities and Generating Genetic Diversity 17 2.1.1 Different Patterns of Alternative Splicing 17 2.1.2 Alternative Splicing and Genetic Diversity 18 2.2 Mechanisms Underlying Alternative Splicing Regulation 19 2.2.1 Splicing Signals and Splicing Regulatory Elements 20 2.2.2 Trans-acting Splicing Regulators 23 2.3 Tissue-specific and Developmentally Regulated Alternative Splicing 26 2.4 Regulation of Alternative Splicing in Response to Extracellular Stimuli 27 3 Pre-mRNA Splicing and Human Diseases 28 3.1 Splicing Defects in Human Diseases 28 3.2 Molecular Mechanisms Underlying Splicing Defects Associated with Disease 33 4 Perspectives on Diagnosis and Treatment of Diseases Caused by pre-mRNA Splicing Defects 36 4.1 Diagnosis of Human Diseases Caused by Splicing Defects 36 2 Alternatively Spliced Genes 4.2 Potential Therapeutic Approaches 37 4.2.1 Oligonucleotide-based Approaches: Antisense, RNAi, and Chimeric Molecules 37 4.2.2 Ribozymes 37 4.2.3 SMaRT 38 4.2.4 Chemical Compounds 38 5 Concluding Remarks 38 Acknowledgment 39 Bibliography 39 Books and Reviews 39 Keywords Pre-mRNA Nascent transcripts that are precursors of mature messenger RNAs. -

STAR RNA-Binding Protein Quaking Suppresses Cancer Via Stabilization of Specific Mirna
Downloaded from genesdev.cshlp.org on September 30, 2021 - Published by Cold Spring Harbor Laboratory Press STAR RNA-binding protein Quaking suppresses cancer via stabilization of specific miRNA An-Jou Chen,1,2,3 Ji-Hye Paik,1,2,3 Hailei Zhang,1,2 Sachet A. Shukla,1,2 Richard Mortensen,4 Jian Hu,1,2,3,5 Haoqiang Ying,1,2,3,5 Baoli Hu,1,2,5 Jessica Hurt,6 Natalie Farny,6 Caroline Dong,1,2 Yonghong Xiao,1,2 Y. Alan Wang,1,2,5 Pamela A. Silver,6 Lynda Chin,1,2,5,7 Shobha Vasudevan,4 and Ronald A. DePinho1,2,3,8,9 1Belfer Institute for Applied Cancer Science, 2Department of Medical Oncology, Dana-Farber Cancer Institute, Harvard Medical School, Boston, Massachusetts 02115, USA; 3Department of Medicine and Genetics, Harvard Medical School, Boston, Massachusetts 02115, USA; 4Cancer Center, Massachusetts General Hospital, Harvard Medical School, Boston, Massachusetts 02129, USA; 5Department of Genomic Medicine, University of Texas MD Anderson Cancer Center, Houston, Texas 77030, USA; 6Department of Systems Biology, Harvard Medical School, Boston, Massachusetts 02115, USA; 7Department of Dermatology, Brigham and Women’s Hospital, Harvard Medical School, Boston, Massachusetts 02115, USA; 8Department of Cancer Biology, University of Texas MD Anderson Cancer Center, Houston, Texas 77030, USA Multidimensional cancer genome analysis and validation has defined Quaking (QKI), a member of the signal transduction and activation of RNA (STAR) family of RNA-binding proteins, as a novel glioblastoma multiforme (GBM) tumor suppressor. Here, we establish that p53 directly regulates QKI gene expression, and QKI protein associates with and leads to the stabilization of miR-20a; miR-20a, in turn, regulates TGFbR2 and the TGFb signaling network. -

CRISPR-Trapseq Identifies the QKI RNA Binding Protein As Important for Astrocytic Maturation and Control of Thalamocortical Synapses
bioRxiv preprint doi: https://doi.org/10.1101/2020.03.13.991224; this version posted March 14, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. CRISPR-TRAPSeq identifies the QKI RNA binding protein as important for astrocytic maturation and control of thalamocortical synapses. Kristina Sakers1,2, Yating Liu1,2,, Lorida Llaci3, Michael J. Vasek1,2, Michael A. Rieger1,2,, Sean Brophy1,2, Eric Tycksen4, Renate Lewis,5 Susan E. Maloney2,, Joseph D. Dougherty*1,2 Affiliations: 1Department of Genetics, Washington University School of Medicine, 660 S. Euclid Ave, Saint Louis MO, 63108, USA. 2Department of Psychiatry, Washington University School of Medicine. 3Division of Biomedical and Biological Sciences, Washington University School of Medicine, 4Genome Technology Access Center, McDonnell Genome Institute, Washington University School of Medicine. 5Hope Center, Washington University School of Medicine and Saint Louis University Contact Information: Dr. Joseph Dougherty Department of Genetics Campus Box 8232 4566 Scott Ave. St. Louis, MO. 63110-1093 P: 314-286-0752 F: 314-362-7855 E:[email protected] Keywords: Quaking, QK, CLIP, SPARC, CRISPR Abstract Quaking RNA binding protein(QKI) is essential for oligodendrocyte development as myelination requires MBP mRNA regulation and localization by the cytoplasmic isoforms(e.g. QKI-6). QKI-6 is also highly expressed in astrocytes, which were recently demonstrated to have regulated mRNA localization. Here, we show via CLIPseq that QKI-6 binds 3’UTRs of a subset of astrocytic mRNAs, including many enriched in peripheral processes. -

Influence of Chromosome 6 Deletion on the Expressional Characteristics in Humans
MOJ Biology and Medicine Mini Review Open Access Influence of chromosome 6 deletion on the expressional characteristics in humans Abstract Volume 6 Issue 1 - 2021 Deletion of chromosomal fragments bring about the devastating disease onset which Taskina Murshed, Rashed Noor is apparently almost incurable. Such deletions usually arise denovo as well as from Department of Life Sciences (DLS), School of Environment and the unstable chromosomal translocations, which in turn affect the terminal part of a Life Sciences (SELS), Independent University, Bangladesh chromosome or an interstitial region, making a large number of genes malfunctioning with the onset of various complications. Chromosome 6q deletion happens due to a missing Correspondence: Rashed Noor, Associate Professor, copy of the genetic material existing within the long arm (q) of the chromosome, which Department of Life Sciences (DLS), School of Environment and causes the loss of multiple necessary genes ending up with an array of physical and mental Life Sciences (SELS), Independent University, Bangladesh (IUB), abnormalities. current review discussed the background of such deleterious effects along Plot 16, Block B, Bashundhara, Dhaka 1229, Bangladesh, with some examples, and depicted on the possible mechanisms; i.e., the genetic insights of Tel +8801749401451, Email chromosomal breakpoint based on the information gathered from the previous literature which could be useful in perspective of the surveillance of genetic disease as well as for the Received: December 11, 2020 | Published: -

Principles of RNA Processing from Analysis of Enhanced CLIP Maps for 150 RNA Binding Proteins Eric L
Van Nostrand et al. Genome Biology (2020) 21:90 https://doi.org/10.1186/s13059-020-01982-9 RESEARCH Open Access Principles of RNA processing from analysis of enhanced CLIP maps for 150 RNA binding proteins Eric L. Van Nostrand1,2, Gabriel A. Pratt1,2, Brian A. Yee1,2, Emily C. Wheeler1,2, Steven M. Blue1,2, Jasmine Mueller1,2, Samuel S. Park1,2, Keri E. Garcia1,2, Chelsea Gelboin-Burkhart1,2, Thai B. Nguyen1,2, Ines Rabano1,2, Rebecca Stanton1,2, Balaji Sundararaman1,2, Ruth Wang1,2, Xiang-Dong Fu1,2, Brenton R. Graveley3* and Gene W. Yeo1,2* Abstract Background: A critical step in uncovering rules of RNA processing is to study the in vivo regulatory networks of RNA binding proteins (RBPs). Crosslinking and immunoprecipitation (CLIP) methods enable mapping RBP targets transcriptome-wide, but methodological differences present challenges to large-scale analysis across datasets. The development of enhanced CLIP (eCLIP) enabled the mapping of targets for 150 RBPs in K562 and HepG2, creating a unique resource of RBP interactomes profiled with a standardized methodology in the same cell types. Results: Our analysis of 223 eCLIP datasets reveals a range of binding modalities, including highly resolved positioning around splicing signals and mRNA untranslated regions that associate with distinct RBP functions. Quantification of enrichment for repetitive and abundant multicopy elements reveals 70% of RBPs have enrichment for non-mRNA element classes, enables identification of novel ribosomal RNA processing factors and sites, and suggests that association with retrotransposable elements reflects multiple RBP mechanisms of action. Analysis of spliceosomal RBPs indicates that eCLIP resolves AQR association after intronic lariat formation, enabling identification of branch points with single-nucleotide resolution, and provides genome-wide validation for a branch point-based scanning model for 3′ splice site recognition. -

Epigenetic Regulation of Alternative Splicing: How Lncrnas Tailor the Message
non-coding RNA Review Epigenetic Regulation of Alternative Splicing: How LncRNAs Tailor the Message Giuseppina Pisignano 1,* and Michael Ladomery 2,* 1 Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, UK 2 Faculty of Health and Applied Sciences, University of the West of England, Coldharbour Lane, Frenchay, Bristol BS16 1QY, UK * Correspondence: [email protected] (G.P.); [email protected] (M.L.) Abstract: Alternative splicing is a highly fine-tuned regulated process and one of the main drivers of proteomic diversity across eukaryotes. The vast majority of human multi-exon genes is alternatively spliced in a cell type- and tissue-specific manner, and defects in alternative splicing can dramatically alter RNA and protein functions and lead to disease. The eukaryotic genome is also intensively transcribed into long and short non-coding RNAs which account for up to 90% of the entire tran- scriptome. Over the years, lncRNAs have received considerable attention as important players in the regulation of cellular processes including alternative splicing. In this review, we focus on recent discoveries that show how lncRNAs contribute significantly to the regulation of alternative splicing and explore how they are able to shape the expression of a diverse set of splice isoforms through several mechanisms. With the increasing number of lncRNAs being discovered and characterized, the contribution of lncRNAs to the regulation of alternative splicing is likely to grow significantly. Keywords: long non-coding RNAs; alternative splicing; splicing factors; post-transcriptional regulation Citation: Pisignano, G.; Ladomery, 1. Introduction M. Epigenetic Regulation of In the late 1970s, researchers were interested in gaining a better understanding of Alternative Splicing: How LncRNAs the mechanisms of adenoviral gene expression when they noticed something unusual, Tailor the Message. -

MYB-QKI Rearrangements in Angiocentric Glioma Drive Tumorigenicity Through a Tripartite Mechanism
MYB-QKI rearrangements in Angiocentric Glioma drive tumorigenicity through a tripartite mechanism The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Bandopadhayay, P., L. A. Ramkissoon, P. Jain, G. Bergthold, J. Wala, R. Zeid, S. E. Schumacher, et al. 2016. “MYB-QKI rearrangements in Angiocentric Glioma drive tumorigenicity through a tripartite mechanism.” Nature genetics 48 (3): 273-282. doi:10.1038/ng.3500. http://dx.doi.org/10.1038/ng.3500. Published Version doi:10.1038/ng.3500 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:29002633 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Nat Genet Manuscript Author . Author manuscript; Manuscript Author available in PMC 2016 August 01. Published in final edited form as: Nat Genet. 2016 February 24; 48(3): 273–282. doi:10.1038/ng.3500. MYB-QKI rearrangements in Angiocentric Glioma drive tumorigenicity through a tripartite mechanism A full list of authors and affiliations appears at the end of the article. Abstract Angiocentric gliomas are pediatric low-grade gliomas (PLGGs) without known recurrent genetic drivers. We performed genomic analysis of new and published data from 249 PLGGs including 19 Angiocentric Gliomas. We identified MYB-QKI fusions as a specific and single candidate driver event in Angiocentric Gliomas. -

Clinical Significance of Methylation and Reduced Expression of The
ANTICANCER RESEARCH 37 : 489-498 (2017) doi:10.21873/anticanres.11341 Clinical Significance of Methylation and Reduced Expression of the Quaking Gene in Colorectal Cancer NORIKO IWATA 1, TOSHIAKI ISHIKAWA 2, SATOSHI OKAZAKI 1, KAORU MOGUSHI 3, HIRONOBU BABA 1, MEGUMI ISHIGURO 4, HIROTOSHI KOBAYASHI 1, HIROSHI TANAKA 3, TATSUYUKI KAWANO 1, KENICHI SUGIHARA 2 and HIROYUKI UETAKE 2 1Division of Surgical Gastroenterology, 2Division of Surgical Specialties, and 4Department of Translational Oncology, Graduate School of Medical and Dental Sciences, and 3Department of Systems Biology, Graduate School of Biochemical Science, Tokyo Medical and Dental University, Tokyo, Japan Abstract. Background: This study investigated abnormal Colorectal cancer (CRC) is one of the best understood methylation in colorectal cancer (CRC) and the potential neoplasms from a genetic perspective, yet it remains the role of the Quaking RNA-binding protein (QKI) gene in second most common cause of cancer-related death. tumorigenesis. Materials and Methods: Oligonucleotide Although the diagnosis and treatment of CRC has microarray expression profiling was carried out on a panel significantly improved during recent decades, leading to a of primary CRC specimens (n=17) and CRC cell lines (n=5), substantial reduction in cancer-related mortality (1), CRC followed by methylation analysis using methylation-specific continues to be a major public health problem worldwide. polymerase chain reaction. QKI expression levels were It is now well established that widespread epigenetic assessed in 156 primary CRCs by qRT-PCR and changes, including alterations in DNA methylation profiles immunohistochemistry. Results: Low QKI expression was relative to non-neoplastic tissue, are a characteristic of many observed in 47.7% in CRCs.