Of Ciliates (Ciliophora) and Some Further Protists at the Upper Austrian Museum in Linz, Including a Guideline for “Typification” of Species*
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Denis BAURAIN Département Des Sciences De La Vie Université De Liège Société Royale Des Sciences De Liège 20 Septembre 2012 Plan De L’Exposé
L’évolution des Eucaryotes Denis BAURAIN Département des Sciences de la Vie Université de Liège Société Royale des Sciences de Liège 20 septembre 2012 Plan de l’exposé 1. Qu’est-ce qu’un Eucaryote ? 2. Quelle est la diversité des Eucaryotes ? 3. Quelles sont les relations de parenté entre les grands groupes d’Eucaryotes ? 4. D’où viennent les Eucaryotes ? Qu’est-ce1 qu’un Eucaryote ? Eukaryotic Cells définition ultrastructurale : organelles spécifiques • noyau (1) • nucléole (2) • RE (5, 8) • Golgi (6) • centriole(s) (13) • mitochondrie(s) (9) • chloroplaste(s) • ... http://en.wikipedia.org/ A eukaryotic gene is arranged in a patchwork of coding (exons) and non-coding sequences (introns). Introns are eliminated while exons are spliced together to yield the mature mRNA used for protein synthesis. http://reflexions.ulg.ac.be/ Gene DNA Transcription Exon1 Exon2 Exon3 Exon4 Exon5 Exon6 pre-mRNA Alternatif splicing mature mRNA Translation Protein In many Eukaryotes, almost all genes can lead to different proteins through a process termed alternative splicing. http://reflexions.ulg.ac.be/ REVIEWS Box 2 | Endosymbiotic evolution and the tree of genomes Intracellular endosymbionts that originally descended from free-living prokaryotes have been important in the evolution of eukaryotes by giving rise to two cytoplasmic organelles. Mitochondria arose from α-proteobacteria and chloroplasts arose from cyanobacteria. Both organelles have made substantial contributions to the complement of genes that are found in eukaryotic nuclei today. The figure shows a schematic diagram of the evolution of eukaryotes, highlighting the incorporation of mitochondria and chloroplasts into the eukaryotic lineage through endosymbiosis and the subsequent co-evolution of the nuclear and organelle genomes. -

Protists and the Wild, Wild West of Gene Expression
MI70CH09-Keeling ARI 3 August 2016 18:22 ANNUAL REVIEWS Further Click here to view this article's online features: • Download figures as PPT slides • Navigate linked references • Download citations Protists and the Wild, Wild • Explore related articles • Search keywords West of Gene Expression: New Frontiers, Lawlessness, and Misfits David Roy Smith1 and Patrick J. Keeling2 1Department of Biology, University of Western Ontario, London, Ontario, Canada N6A 5B7; email: [email protected] 2Canadian Institute for Advanced Research, Department of Botany, University of British Columbia, Vancouver, British Columbia, Canada V6T 1Z4; email: [email protected] Annu. Rev. Microbiol. 2016. 70:161–78 Keywords First published online as a Review in Advance on constructive neutral evolution, mitochondrial transcription, plastid June 17, 2016 transcription, posttranscriptional processing, RNA editing, trans-splicing The Annual Review of Microbiology is online at micro.annualreviews.org Abstract This article’s doi: The DNA double helix has been called one of life’s most elegant structures, 10.1146/annurev-micro-102215-095448 largely because of its universality, simplicity, and symmetry. The expression Annu. Rev. Microbiol. 2016.70:161-178. Downloaded from www.annualreviews.org Copyright c 2016 by Annual Reviews. Access provided by University of British Columbia on 09/24/17. For personal use only. of information encoded within DNA, however, can be far from simple or All rights reserved symmetric and is sometimes surprisingly variable, convoluted, and wantonly inefficient. Although exceptions to the rules exist in certain model systems, the true extent to which life has stretched the limits of gene expression is made clear by nonmodel systems, particularly protists (microbial eukary- otes). -

The Macronuclear Genome of Stentor Coeruleus Reveals Tiny Introns in a Giant Cell
University of Pennsylvania ScholarlyCommons Departmental Papers (Biology) Department of Biology 2-20-2017 The Macronuclear Genome of Stentor coeruleus Reveals Tiny Introns in a Giant Cell Mark M. Slabodnick University of California, San Francisco J. G. Ruby University of California, San Francisco Sarah B. Reiff University of California, San Francisco Estienne C. Swart University of Bern Sager J. Gosai University of Pennsylvania See next page for additional authors Follow this and additional works at: https://repository.upenn.edu/biology_papers Recommended Citation Slabodnick, M. M., Ruby, J. G., Reiff, S. B., Swart, E. C., Gosai, S. J., Prabakaran, S., Witkowska, E., Larue, G. E., Gregory, B. D., Nowacki, M., Derisi, J., Roy, S. W., Marshall, W. F., & Sood, P. (2017). The Macronuclear Genome of Stentor coeruleus Reveals Tiny Introns in a Giant Cell. Current Biology, 27 (4), 569-575. http://dx.doi.org/10.1016/j.cub.2016.12.057 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/biology_papers/49 For more information, please contact [email protected]. The Macronuclear Genome of Stentor coeruleus Reveals Tiny Introns in a Giant Cell Abstract The giant, single-celled organism Stentor coeruleus has a long history as a model system for studying pattern formation and regeneration in single cells. Stentor [1, 2] is a heterotrichous ciliate distantly related to familiar ciliate models, such as Tetrahymena or Paramecium. The primary distinguishing feature of Stentor is its incredible size: a single cell is 1 mm long. Early developmental biologists, including T.H. Morgan [3], were attracted to the system because of its regenerative abilities—if large portions of a cell are surgically removed, the remnant reorganizes into a normal-looking but smaller cell with correct proportionality [2, 3]. -

The Planktonic Protist Interactome: Where Do We Stand After a Century of Research?
bioRxiv preprint doi: https://doi.org/10.1101/587352; this version posted May 2, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Bjorbækmo et al., 23.03.2019 – preprint copy - BioRxiv The planktonic protist interactome: where do we stand after a century of research? Marit F. Markussen Bjorbækmo1*, Andreas Evenstad1* and Line Lieblein Røsæg1*, Anders K. Krabberød1**, and Ramiro Logares2,1** 1 University of Oslo, Department of Biosciences, Section for Genetics and Evolutionary Biology (Evogene), Blindernv. 31, N- 0316 Oslo, Norway 2 Institut de Ciències del Mar (CSIC), Passeig Marítim de la Barceloneta, 37-49, ES-08003, Barcelona, Catalonia, Spain * The three authors contributed equally ** Corresponding authors: Ramiro Logares: Institute of Marine Sciences (ICM-CSIC), Passeig Marítim de la Barceloneta 37-49, 08003, Barcelona, Catalonia, Spain. Phone: 34-93-2309500; Fax: 34-93-2309555. [email protected] Anders K. Krabberød: University of Oslo, Department of Biosciences, Section for Genetics and Evolutionary Biology (Evogene), Blindernv. 31, N-0316 Oslo, Norway. Phone +47 22845986, Fax: +47 22854726. [email protected] Abstract Microbial interactions are crucial for Earth ecosystem function, yet our knowledge about them is limited and has so far mainly existed as scattered records. Here, we have surveyed the literature involving planktonic protist interactions and gathered the information in a manually curated Protist Interaction DAtabase (PIDA). In total, we have registered ~2,500 ecological interactions from ~500 publications, spanning the last 150 years. -

Role of Metopus Es in the Anaerobic Degradation of Organic Matter and Biomethanation Process
Role of Metopus es in the anaerobic degradation of organic matter and biomethanation process Thesis submitted to the University of Kerala for award of the degree of Doctor of Philosophy in Microbiology By Nimi Narayanan Under the Supervision of Dr. V. B. Manilal Process Engineering and Environmental Technology Division, National Institute for Interdisciplinary Science and Technology (NIIST), CSIR, Thiruvananthapuram, Kerala, INDIA - 695019 2011 To my Family DECLARATION I hereby declare that the work presented in this thesis is based on the original work done by me under the guidance of Dr. V. B. Manilal, Principal Scientist, Process Engineering and Environmental Technology Division, National Institute for Interdisciplinary Science and Technology, and that no part of this has been included in any other thesis submitted previously for the award of any degree. Nimi Narayanan Acknowledgement It is a great pleasure to express my sincere gratitude and sense of appreciation to my research guide, Dr. V.B. Manilal, Principal Scientist, Environmental Technology, NIIST, Trivandrum, for his constant encouragement, enthusiastic support and valuable guidance throughout the period of study. I am indebted to him for giving me the ample freedom to do the work and express my ideas during this period. I am grateful to Dr. Ajit Haridas, Scientist-in-charge, Environmental Technology, NIIST for his valuable suggestions and constructive criticism during my tenure. It is an honor for me to thank the present Director, Dr. Suresh Das and the former directors of NIIST, Trivandrum for providing the necessary infrastructural facilities for the successful completion of work. I would like to thank Council of Scientific and Industrial Research, New Delhi, for the research fellowship. -

University of Oklahoma
UNIVERSITY OF OKLAHOMA GRADUATE COLLEGE MACRONUTRIENTS SHAPE MICROBIAL COMMUNITIES, GENE EXPRESSION AND PROTEIN EVOLUTION A DISSERTATION SUBMITTED TO THE GRADUATE FACULTY in partial fulfillment of the requirements for the Degree of DOCTOR OF PHILOSOPHY By JOSHUA THOMAS COOPER Norman, Oklahoma 2017 MACRONUTRIENTS SHAPE MICROBIAL COMMUNITIES, GENE EXPRESSION AND PROTEIN EVOLUTION A DISSERTATION APPROVED FOR THE DEPARTMENT OF MICROBIOLOGY AND PLANT BIOLOGY BY ______________________________ Dr. Boris Wawrik, Chair ______________________________ Dr. J. Phil Gibson ______________________________ Dr. Anne K. Dunn ______________________________ Dr. John Paul Masly ______________________________ Dr. K. David Hambright ii © Copyright by JOSHUA THOMAS COOPER 2017 All Rights Reserved. iii Acknowledgments I would like to thank my two advisors Dr. Boris Wawrik and Dr. J. Phil Gibson for helping me become a better scientist and better educator. I would also like to thank my committee members Dr. Anne K. Dunn, Dr. K. David Hambright, and Dr. J.P. Masly for providing valuable inputs that lead me to carefully consider my research questions. I would also like to thank Dr. J.P. Masly for the opportunity to coauthor a book chapter on the speciation of diatoms. It is still such a privilege that you believed in me and my crazy diatom ideas to form a concise chapter in addition to learn your style of writing has been a benefit to my professional development. I’m also thankful for my first undergraduate research mentor, Dr. Miriam Steinitz-Kannan, now retired from Northern Kentucky University, who was the first to show the amazing wonders of pond scum. Who knew that studying diatoms and algae as an undergraduate would lead me all the way to a Ph.D. -

Morphology and Phylogeny of the Soil Ciliate Metopus Yantaiensis N. Sp
Journal of Eukaryotic Microbiology ISSN 1066-5234 ORIGINAL ARTICLE Morphology and Phylogeny of the Soil Ciliate Metopus yantaiensis n. sp. (Ciliophora, Metopida), with Identification of the Intracellular Bacteria Atef Omara,b, Qianqian Zhanga, Songbao Zoua,c & Jun Gonga,c a Laboratory of Microbial Ecology and Matter Cycles, Yantai Institute of Coastal Zone Research, Chinese Academy of Sciences, Yantai 264003, China b Department of Zoology, Al-Azhar University, Assiut 71524, Egypt c University of Chinese Academy of Sciences, Beijing 100049, China Keywords ABSTRACT Anaerobic ciliates; Armophorea; digestion- resistant bacteria; Metopus contortus; The morphology and infraciliature of a new ciliate, Metopus yantaiensis n. sp., Metopus hasei. discovered in coastal soil of northern China, were investigated. It is distin- guished from its congeners by a combination of the following features: nuclear Correspondence apparatus situated in the preoral dome; 18–21 somatic ciliary rows, of which J. Gong, Yantai Institute of Coastal Zone three extend onto the preoral dome (dome kineties); three to five distinctly Research, Chinese Academy of Sciences, elongated caudal cilia, and 21–29 adoral polykinetids. The 18S rRNA genes of Yantai 264003, China this new species and two congeners, Metopus contortus and Metopus hasei, Telephone number: +86-535-2109123; were sequenced and phylogenetically analyzed. The new species is more clo- FAX number: +86-535-2109000; sely related to M. hasei and the clevelandellids than to other congeners; both e-mail: [email protected] the genus Metopus and the order Metopida are not monophyletic. In addition, the digestion-resistant bacteria in the cytoplasm of M. yantaiensis were identi- Received: 13 January 2017; revised 2 March fied, using a 16S rRNA gene clone library, sequencing, and fluorescence 2017; accepted March 8, 2017. -

Old Woman Creek National Estuarine Research Reserve Management Plan 2011-2016
Old Woman Creek National Estuarine Research Reserve Management Plan 2011-2016 April 1981 Revised, May 1982 2nd revision, April 1983 3rd revision, December 1999 4th revision, May 2011 Prepared for U.S. Department of Commerce Ohio Department of Natural Resources National Oceanic and Atmospheric Administration Division of Wildlife Office of Ocean and Coastal Resource Management 2045 Morse Road, Bldg. G Estuarine Reserves Division Columbus, Ohio 1305 East West Highway 43229-6693 Silver Spring, MD 20910 This management plan has been developed in accordance with NOAA regulations, including all provisions for public involvement. It is consistent with the congressional intent of Section 315 of the Coastal Zone Management Act of 1972, as amended, and the provisions of the Ohio Coastal Management Program. OWC NERR Management Plan, 2011 - 2016 Acknowledgements This management plan was prepared by the staff and Advisory Council of the Old Woman Creek National Estuarine Research Reserve (OWC NERR), in collaboration with the Ohio Department of Natural Resources-Division of Wildlife. Participants in the planning process included: Manager, Frank Lopez; Research Coordinator, Dr. David Klarer; Coastal Training Program Coordinator, Heather Elmer; Education Coordinator, Ann Keefe; Education Specialist Phoebe Van Zoest; and Office Assistant, Gloria Pasterak. Other Reserve staff including Dick Boyer and Marje Bernhardt contributed their expertise to numerous planning meetings. The Reserve is grateful for the input and recommendations provided by members of the Old Woman Creek NERR Advisory Council. The Reserve is appreciative of the review, guidance, and council of Division of Wildlife Executive Administrator Dave Scott and the mapping expertise of Keith Lott and the late Steve Barry. -

A Fossilized Microcenosis in Triassic Amber
J Eukaqlnr. Micmbrol., 46(6), 1999 pp. 571-584 0 1999 by the Society of Protozoologists A Fossilized Microcenosis in Triassic Amber WILFRIED SCHONBORN,a HEINRICH DORFELT? WILHELM FOISSNER,’ LOTHAR KRIENITZd and URSULA SCHAFER” “Friedrich-Schiller-UniversitatJena, Institut fur Okologie, Arbeitsgruppe Limnologie, Winzerlaer StraJe 10, 0-0774.5 Jena, Germany, and bFriedrich-Schiller-Univer.sitatJenn,lnstitut fiir Ernahrung und Umwelt, Lehrgebiet Lnndschaftsokologie und Naturschutz, DornburgerslraJe 159, 0-07743 Jena, Germany, and ‘Universitut Salzburg, Institut fur Zoologie, HellbrunnerstraJe 34, A-5020 Salzburg, Austria, and dInstitut fur Gewasseriikologie und Binnenjscherei, Alte Fischerhiitte 2, 0-16775 Neuglobsow, Germany ABSTRACT. Detailed data on bacterial and protistan microfossils are presented from a 0.003 mm3 piece of Triassic amber (Schlier- seerit, Upper Triassic period, 220-230 million years old). This microcenosis, which actually existed as such within a very small, probably semiaquatic habitat, included the remains of about two bacteria species, four fungi (Palaeodikaryomyces baueri, Pithomyces-like conidia, capillitium-like hyphae, yeast cells) two euglenoids, two chlamydomonas (Chlamydomonas sp., Chloromonas sp.), two coccal green microalgae (Chlorellu sp., Chorzcystis-like cells), one zooflagellate, three testate amoebae (Centropyxis aculeata var. oblonga-like, Cyclopyxis eurystoma-like, Hyalosphenia baueri n. sp.), seven ciliates (Pseudoplatyophrya nana-like, Mykophagophrys rerricola-like, Cvrtolophosis mucicola-like, Paracondylostoma -
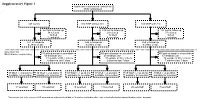
Supplementary Figure 1 Multicenter Randomised Control Trial 2746 Randomised
Supplementary Figure 1 Multicenter randomised control trial 2746 randomised 947 control 910 MNP without zinc 889 MNP with zinc 223 lost to follow up 219 lost to follow up 183 lost to follow up 34 refused 29 refused 37 refused 16 died 12 died 9 died 3 excluded 4 excluded 1 excluded 671 in follow-up 646 in follow-up 659 in follow-up at 24mo of age at 24mo of age at 24mo of age Selection for Microbiome sequencing 516 paired samples unavailable 469 paired samples unavailable 497 paired samples unavailable 69 antibiotic use 63 antibiotic use 67 antibiotic use 31 outside of WLZ criteria 37 outside of WLZ criteria 34 outside of WLZ criteria 6 diarrhea last 7 days 2 diarrhea last 7 days 7 diarrhea last 7 days 39 WLZ > -1 at 24 mo 10 WLZ < -2 at 24mo 58 WLZ > -1 at 24 mo 17 WLZ < -2 at 24mo 48 WLZ > -1 at 24 mo 8 WLZ < -2 at 24mo available for selection available for selection available for selection available for selection available for selection1 available for selection1 14 selected 10 selected 15 selected 14 selected 20 selected1 7 selected1 1 Two subjects (one in the reference WLZ group and one undernourished) had, at 12 months, no diarrhea within 1 day of stool collection but reported diarrhea within 7 days prior. Length, cm kg Weight, Supplementary Figure 2. Length (left) and weight (right) z-scores of children recruited into clinical trial NCT00705445 during the first 24 months of life. Median and quantile values are shown, with medians for participants profiled in current study indicated by red (undernourished) and black (reference WLZ) lines. -

Protistology an International Journal Vol
Protistology An International Journal Vol. 10, Number 2, 2016 ___________________________________________________________________________________ CONTENTS INTERNATIONAL SCIENTIFIC FORUM «PROTIST–2016» Yuri Mazei (Vice-Chairman) Welcome Address 2 Organizing Committee 3 Organizers and Sponsors 4 Abstracts 5 Author Index 94 Forum “PROTIST-2016” June 6–10, 2016 Moscow, Russia Website: http://onlinereg.ru/protist-2016 WELCOME ADDRESS Dear colleagues! Republic) entitled “Diplonemids – new kids on the block”. The third lecture will be given by Alexey The Forum “PROTIST–2016” aims at gathering Smirnov (Saint Petersburg State University, Russia): the researchers in all protistological fields, from “Phylogeny, diversity, and evolution of Amoebozoa: molecular biology to ecology, to stimulate cross- new findings and new problems”. Then Sandra disciplinary interactions and establish long-term Baldauf (Uppsala University, Sweden) will make a international scientific cooperation. The conference plenary presentation “The search for the eukaryote will cover a wide range of fundamental and applied root, now you see it now you don’t”, and the fifth topics in Protistology, with the major focus on plenary lecture “Protist-based methods for assessing evolution and phylogeny, taxonomy, systematics and marine water quality” will be made by Alan Warren DNA barcoding, genomics and molecular biology, (Natural History Museum, United Kingdom). cell biology, organismal biology, parasitology, diversity and biogeography, ecology of soil and There will be two symposia sponsored by ISoP: aquatic protists, bioindicators and palaeoecology. “Integrative co-evolution between mitochondria and their hosts” organized by Sergio A. Muñoz- The Forum is organized jointly by the International Gómez, Claudio H. Slamovits, and Andrew J. Society of Protistologists (ISoP), International Roger, and “Protists of Marine Sediments” orga- Society for Evolutionary Protistology (ISEP), nized by Jun Gong and Virginia Edgcomb. -

Plenary Lecture & Symposium
PLENARY LECTURE & SYMPOSIUM SYMPOSIuM From genomics to flagellar and ciliary struc - MONDAY 29 JulY tures and cytoskeleton dynamics (by FEPS) PlENARY lECTuRE (ISoP Honorary Member lECTuRE) Chairs (by ISoP) Cristina Miceli , University of Camerino, Camerino, Italy Helena Soares , University of Lisbon and Gulbenkian Foun - Introduction - John Dolan , CNRS-Sorbonne University, Ville - dation, Lisbon, Portugal franche-sur-Mer, France. Jack Sunter - Oxford Brookes University, Oxford, UK- Genome Tom Fenchel University of Copenhagen, Copenhagen, Den - wide tagging in trypanosomes uncovers flagellum asymmetries mark Dorota Wloga - Nencki Institute of Experimental Biology, War - ISoP Honorary Member saw, Poland - Deciphering the molecular mechanisms that coor - dinate ciliary outer doublet complexes – search for “missing Size, Shape and Function among Protozoa links” Helena Soares - University of Lisbon and Polytechnic Institute of Lisbon, Lisbon, Portugal - From centrosomal microtubule an - SYMPOSIuM on ciliate biology and taxonomy in memory choring and organization to basal body positioning: TBCCD1 an of Denis lynn (by FEPS/ISoP) elusive protein Chairs Pierangelo luporini , University of Camerino, Camerino, Italy Roberto Docampo , University of Georgia, Athens, Georgia TuESDAY 30 JulY Alan Warren - Natural History Museum, London, UK. The bio - logy and systematics of peritrich ciliates: old concepts and new PlENARY lECTuRE (PAST-PRESIDENT LECTURE, by ISoP) findings Rebecca Zufall - University of Houston, Houston, USA. Amitosis Introduction - Avelina Espinosa , Roger Williams University, and the Evolution of Asexuality in Tetrahymena Ciliates Bristol, USA Sabine Agatha - University of Salzburg, Salzburg, Austria. The biology and systematics of oligotrichean ciliates: new findings David Bass and old concepts Natural History Museum London, London & Cefas, Weymouth, laura utz - School of Sciences, PUCRS, Porto Alegre, Brazil.