The Transcription Factor NF-Atc1 Regulates Lymphocyte Proliferation and Th2 Cytokine Production
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Transcriptional Regulation by Extracellular Signals 209
Cell, Vol. 80, 199-211, January 27, 1995, Copyright © 1995 by Cell Press Transcriptional Regulation Review by Extracellular Signals: Mechanisms and Specificity Caroline S. Hill and Richard Treisman Nuclear Translocation Transcription Laboratory In principle, regulated nuclear localization of transcription Imperial Cancer Research Fund factors can involve regulated activity of either nuclear lo- Lincoln's Inn Fields calization signals (NLSs) or cytoplasmic retention signals, London WC2A 3PX although no well-characterized case of the latter has yet England been reported. N LS activity, which is generally dependent on short regions of basic amino acids, can be regulated either by masking mechanisms or by phosphorylations Changes in cell behavior induced by extracellular signal- within the NLS itself (Hunter and Karin, 1992). For exam- ing molecules such as growth factors and cytokines re- ple, association with an inhibitory subunit masks the NLS quire execution of a complex program of transcriptional of NF-KB and its relatives (Figure 1; for review see Beg events. While the route followed by the intracellular signal and Baldwin, 1993), while an intramolecular mechanism from the cell membrane to its transcription factor targets may mask NLS activity in the heat shock regulatory factor can be traced in an increasing number of cases, how the HSF2 (Sheldon and Kingston, 1993). When transcription specificity of the transcriptional response of the cell to factor localization is dependent on regulated NLS activity, different stimuli is determined is much less clear. How- linkage to a constitutively acting NLS may be sufficient to ever, it is possible to understand at least in principle how render nuclear localization independent of signaling (Beg different stimuli can activate the same signal pathway yet et al., 1992). -

NF-Κb Modifies the Mammalian Circadian Clock Through Interaction with the Core Clock Protein BMAL1
bioRxiv preprint doi: https://doi.org/10.1101/2020.09.06.285254; this version posted September 6, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. NF-κB modifies the mammalian circadian clock through interaction with the core clock protein BMAL1 Yang Shen1, Wei Wang2, Mehari Endale1, Lauren J. Francey3, Rachel L. Harold4, David W. Hammers5, Zhiguang Huo6, Carrie L. Partch4, John B. Hogenesch3, Zhao-Hui Wu2, and Andrew C. Liu1* 1Department of Physiology and Functional Genomics, University of Florida College of Medicine; 2Department of Radiation Oncology and Center for Cancer Research, University of Tennessee Health Science Center; 3Divisions of Human Genetics and Immunobiology, Cincinnati Children's Hospital Medical Center; 4Department of Chemistry and Biochemistry, University of California Santa Cruz; 5Department of Pharmacology and Therapeutics, University of Florida College of Medicine; 6Department of Biostatistics, College of Public Health & Health Professions, University of Florida College of Medicine Running title: NF-κB modifies circadian clock * Correspondence: Andrew C. Liu, [email protected] Keywords: Circadian clock, inflammation, NF-κB, suprachiasmatic nucleus (SCN), BMAL1 Abbreviations: NF-κB, nuclear factor kappa B; SCN, suprachiasmatic nucleus; IKK, IκB kinase; IκBα, inhibitor of NF-κB; Luc, luciferase; RHD, Rel homology domain; CRD, C-terminal oscillation regulatory domain, TAD, transactivation domain 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.09.06.285254; this version posted September 6, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. -
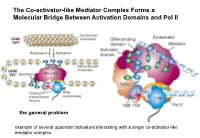
The Co-Activator-Like Mediator Complex Forms a Molecular Bridge Between Activation Domains and Pol II
The Co-activator-like Mediator Complex Forms a Molecular Bridge Between Activation Domains and Pol II Co-activator the general problem example of several upstream activators interacting with a single co-activator-like mediator complex Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P. Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00 REGULATORY MECHANISMS OF TRANSCRIPTION FACTOR FUNCTION •Protein synthesized •Protein phosphorylated •Ligand binding •Release inhibitor •Change partner, etc RG Clerc April 6. 2016 Transcription factors as final effectors of the cellular signaling cascade Regulatory Mechanisms of Transcription Factor Function Genes X. Lewin B. editor Regulatory Mechanisms of Transcription Factor Function CREB FOXO NFAT Genes X. Lewin B. editor Body plan is constructed through interactions of the developmentally regulated homeotic gene expression anterior early expression posterior late expression high RA response low RA response hindbrain trunk Alexander T and Krumlauf R. Ann.Rev.Cell.Dev.Biol: 25_431 (2009) Hoxb transcription factors mRNA distribution along the AP axis A staggered expression of the anterior border within somites is a property of the physical ordering along the chromosome: the colinearity Alexander T and Krumlauf R. Ann.Rev.Cell.Dev.Biol: 25_431 (2009) Transcription control of the Hox genes: insight into colinear activation P A p p p p a p a p a a a a Tarchini B and Duboule D. Dev.Cell 10_93 (2006) correlation between linear arrangement along the chromosome and spatial -

The C-Rel Transcription Factor and B-Cell Proliferation: a Deal with the Devil
Oncogene (2004) 23, 2275–2286 & 2004 Nature Publishing Group All rights reserved 0950-9232/04 $25.00 www.nature.com/onc REVIEW The c-Rel transcription factor and B-cell proliferation: a deal with the devil Thomas D Gilmore*,1, Demetrios Kalaitzidis1, Mei-Chih Liang1 and Daniel T Starczynowski1 1Department of Biology, Boston University, 5 Cummington Street, Boston, MA 02215, USA Activation of the Rel/NF-jB signal transduction pathway called the Rel homology domain, which has sequences has been associated with a varietyof animal and human required for DNA binding, dimerization, nuclear malignancies. However, among the Rel/NF-jB family localization, and inhibitor (IkB) binding. Sequences members, onlyc-Rel has been consistentlyshown to be C-terminal to the Rel homology domain in c-Rel, RelA, able to malignantlytransform cells in culture. In addition, and RelB contain transcriptional activation domains, c-rel has been activated bya retroviral promoter insertion and therefore dimers that contain these Rel/NF-kB in an avian B-cell lymphoma, and amplifications of REL members usually increase transcription of target genes. (human c-rel) are frequentlyseen in Hodgkin’s lympho- Although the Rel homology domain sequences of c-Rel mas and diffuse large B-cell lymphomas, and in some proteins across species are highly conserved, their follicular and mediastinal B-cell lymphomas. Phenotypic C-terminal transactivation domains are not. For exam- analysis of c-rel knockout mice demonstrates that c-Rel ple, the Rel homology domains of chicken and human has a normal role in B-cell proliferation and survival; c-Rel are approximately 85% identical, whereas their moreover, c-Rel nuclear activityis required for B-cell C-terminal transactivation domains are only about 10% development. -

The Role of Ubiquitination in NF-Κb Signaling During Virus Infection
viruses Review The Role of Ubiquitination in NF-κB Signaling during Virus Infection Kun Song and Shitao Li * Department of Microbiology and Immunology, Tulane University, New Orleans, LA 70112, USA; [email protected] * Correspondence: [email protected] Abstract: The nuclear factor κB (NF-κB) family are the master transcription factors that control cell proliferation, apoptosis, the expression of interferons and proinflammatory factors, and viral infection. During viral infection, host innate immune system senses viral products, such as viral nucleic acids, to activate innate defense pathways, including the NF-κB signaling axis, thereby inhibiting viral infection. In these NF-κB signaling pathways, diverse types of ubiquitination have been shown to participate in different steps of the signal cascades. Recent advances find that viruses also modulate the ubiquitination in NF-κB signaling pathways to activate viral gene expression or inhibit host NF-κB activation and inflammation, thereby facilitating viral infection. Understanding the role of ubiquitination in NF-κB signaling during viral infection will advance our knowledge of regulatory mechanisms of NF-κB signaling and pave the avenue for potential antiviral therapeutics. Thus, here we systematically review the ubiquitination in NF-κB signaling, delineate how viruses modulate the NF-κB signaling via ubiquitination and discuss the potential future directions. Keywords: NF-κB; polyubiquitination; linear ubiquitination; inflammation; host defense; viral infection Citation: Song, K.; Li, S. The Role of 1. Introduction Ubiquitination in NF-κB Signaling The nuclear factor κB (NF-κB) is a small family of five transcription factors, including during Virus Infection. Viruses 2021, RelA (also known as p65), RelB, c-Rel, p50 and p52 [1]. -

And C-Terminal Domains of Relb Are Required for Full Transactivation
MOLECULAR AND CELLULAR BIOLOGY, Mar. 1993, p. 1572-1582 Vol. 13, No. 3 0270-7306/93/031572-11$02.00/0 Copyright © 1993, American Society for Microbiology Both N- and C-Terminal Domains of RelB Are Required for Full Transactivation: Role of the N-Terminal Leucine Zipper-Like Motif PAWEL DOBRZANSKI, ROLF-PETER RYSECK, AND RODRIGO BRAVO* Department ofMolecular Biology, Bristol-Myers Squibb Pharmaceutical Research Institute, P.O. Box 4000, Princeton, New Jersey 08543-4000 Received 18 September 1992/Returned for modification 11 November 1992/Accepted 12 December 1992 RelB, a member of the Rel family of transcription factors, can stimulate promoter activity in the presence of p50-NF-KB or p5OB/p49-NF-KB in mammalian cells. Transcriptional activation analysis reveals that the N and C termini of RelB are required for full transactivation in the presence of p50-NF-KB. RelB/p50-NF-KB hybrid molecules containing the Rel homology domain of p50-NF-KB and the N and C termini of RelB have high transcriptional activity compared with wild-type p50-NF-KB. The N and C termini of RelB cooperate in transactivation in cis or trans configuration. Alterations in the structure of the leucine zipper-like motif present in the N terminus of RelB significantly decrease the transcriptional capacity of RelB and of different RelB/p50-NF-KB hybrid molecules. Serum stimulation of quiescent fibroblasts induces the regulated by the inhibitory factor cactus (28). The different expression of more than 100 genes, including several proto- homodimers and heterodimers formed between the members oncogenes encoding for transcriptional factors belonging to of the Rel family regulate gene expression in vivo (20, 22, 42) the Jun, Fos, and Rel families (2, 11, 13, 24, 34-36, 38). -

Direct Interaction Between Estrogen Receptor and NF-B in the Nucleus Of
Direct interaction between estrogen receptor α and NF-κB in the nucleus of living cells Monique E. Quaedackers, Christina E. van den Brink, Paul T. van der Saag, Leon G.J. Tertoolen To cite this version: Monique E. Quaedackers, Christina E. van den Brink, Paul T. van der Saag, Leon G.J. Tertoolen. Direct interaction between estrogen receptor α and NF-κB in the nucleus of living cells. Molecular and Cellular Endocrinology, Elsevier, 2007, 273 (1-2), pp.42. 10.1016/j.mce.2007.05.002. hal-00531925 HAL Id: hal-00531925 https://hal.archives-ouvertes.fr/hal-00531925 Submitted on 4 Nov 2010 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Accepted Manuscript Title: Direct interaction between estrogen receptor ␣ and NF-B in the nucleus of living cells Authors: Monique E. Quaedackers, Christina E. van den Brink, Paul T. van der Saag, Leon G.J. Tertoolen PII: S0303-7207(07)00185-2 DOI: doi:10.1016/j.mce.2007.05.002 Reference: MCE 6650 To appear in: Molecular and Cellular Endocrinology Received date: 27-3-2007 Revised date: 7-5-2007 Accepted date: 8-5-2007 Please cite this article as: Quaedackers, M.E., van den Brink, C.E., van der Saag, P.T., Tertoolen, L.G.J., Direct interaction between estrogen receptor ␣ and NF- B in the nucleus of living cells, Molecular and Cellular Endocrinology (2007), doi:10.1016/j.mce.2007.05.002 This is a PDF file of an unedited manuscript that has been accepted for publication. -

Review Recognition of Specific DNA Sequences
Molecular Cell, Vol. 8, 937–946, November, 2001, Copyright 2001 by Cell Press Recognition of Specific DNA Sequences Review Colin W. Garvie and Cynthia Wolberger1 teins that mediate recombination, DNA cleavage, meth- Department of Biophysics and Biophysical Chemistry ylation, and other processes. The broad principles of and the Howard Hughes Medical Institute DNA binding outlined here pertain to these proteins as Johns Hopkins University School of Medicine well, although the structural details differ. Baltimore, Maryland 21205 Basic Requirements for DNA Binding Double-stranded DNA is a polymer of relatively uniform Proteins that recognize specific DNA sequences play structure, with a highly negatively charged sugar-phos- a central role in the regulation of transcription. The phate backbone and a core of stacked base pairs whose tremendous increase in structural information on pro- edges are exposed in the major and minor grooves. tein-DNA complexes has uncovered a remarkable Since each base pair has a characteristic set of func- structural diversity in DNA binding folds, while at the tional groups, each DNA sequence has a chemical “sig- same time revealing common themes in binding to nature” characterized by the pattern of these groups target sites in the genome. exposed in the DNA grooves. It is this chemical surface, along with sequence-dependent variations in DNA Introduction structure and flexibility, that is recognized by proteins. Proteins recognize a particular sequence by having a The ability of a protein to bind selectively to a particular surface that is chemically complementary to that of the DNA site in the genome is the foundation upon which DNA, forming a series of favorable electrostatic and van transcriptional regulatory pathways are built. -

Insect Transcription Factors: a Landscape of Their Structures and Biological Functions in Drosophila and Beyond
International Journal of Molecular Sciences Review Insect Transcription Factors: A Landscape of Their Structures and Biological Functions in Drosophila and beyond Zhaojiang Guo 1,† , Jianying Qin 1,2,†, Xiaomao Zhou 2 and Youjun Zhang 1,* 1 Department of Plant Protection, Institute of Vegetables and Flowers, Chinese Academy of Agricultural Sciences, Beijing 100081, China; [email protected] (Z.G.); [email protected] (J.Q.) 2 Longping Branch, Graduate School of Hunan University, Changsha 410125, China; [email protected] * Correspondence: [email protected]; Tel.: +86-10-82109518 † These authors contributed equally to this work. Received: 23 October 2018; Accepted: 16 November 2018; Published: 21 November 2018 Abstract: Transcription factors (TFs) play essential roles in the transcriptional regulation of functional genes, and are involved in diverse physiological processes in living organisms. The fruit fly Drosophila melanogaster, a simple and easily manipulated organismal model, has been extensively applied to study the biological functions of TFs and their related transcriptional regulation mechanisms. It is noteworthy that with the development of genetic tools such as CRISPR/Cas9 and the next-generation genome sequencing techniques in recent years, identification and dissection the complex genetic regulatory networks of TFs have also made great progress in other insects beyond Drosophila. However, unfortunately, there is no comprehensive review that systematically summarizes the structures and biological functions of TFs in both model and non-model insects. Here, we spend extensive effort in collecting vast related studies, and attempt to provide an impartial overview of the progress of the structure and biological functions of current documented TFs in insects, as well as the classical and emerging research methods for studying their regulatory functions. -

Requirement for Transcription Factor NFAT in Interleukin-2 Expression
University of Massachusetts Medical School eScholarship@UMMS Open Access Articles Open Access Publications by UMMS Authors 1999-02-18 Requirement for transcription factor NFAT in interleukin-2 expression Chi-Wing Chow University of Massachusetts Medical School Et al. Let us know how access to this document benefits ou.y Follow this and additional works at: https://escholarship.umassmed.edu/oapubs Part of the Life Sciences Commons, and the Medicine and Health Sciences Commons Repository Citation Chow C, Rincon M, Davis RJ. (1999). Requirement for transcription factor NFAT in interleukin-2 expression. Open Access Articles. Retrieved from https://escholarship.umassmed.edu/oapubs/1444 This material is brought to you by eScholarship@UMMS. It has been accepted for inclusion in Open Access Articles by an authorized administrator of eScholarship@UMMS. For more information, please contact [email protected]. MOLECULAR AND CELLULAR BIOLOGY, Mar. 1999, p. 2300–2307 Vol. 19, No. 3 0270-7306/99/$04.0010 Copyright © 1999, American Society for Microbiology. All Rights Reserved. Requirement for Transcription Factor NFAT in Interleukin-2 Expression 1 2 1 CHI-WING CHOW, MERCEDES RINCO´ N, AND ROGER J. DAVIS * Howard Hughes Medical Institute and Program in Molecular Medicine, Department of Biochemistry and Molecular Biology, University of Massachusetts Medical School, Worcester, Massachusetts 01605,1 and Program in Immunobiology, Department of Medicine, University of Vermont, Burlington, Vermont 054052 Received 23 July 1998/Returned for modification 5 October 1998/Accepted 24 November 1998 The nuclear factor of activated T cells (NFAT) transcription factor is implicated in expression of the cytokine interleukin-2 (IL-2). Binding sites for NFAT are located in the IL-2 promoter. -

Nephrin Deficiency Activates NF-B and Promotes Glomerular Injury
BASIC RESEARCH www.jasn.org Nephrin Deficiency Activates NF-B and Promotes Glomerular Injury Sagair Hussain,* Leile Romio,* Moin Saleem,† Peter Mathieson,† Manuel Serrano,‡ Jorge Moscat,§ Maria Diaz-Meco,§ Peter Scambler,* and Ania Koziell* *Molecular Medicine Unit, Institute of Child Health, London, and †Academic Renal Unit, University of Bristol, Southmead Hospital, Bristol, United Kingdom; ‡Spanish National Cancer Research Centre, Madrid, Spain; and §Department of Genome Science, Genome Research Institute, University of Cincinnati, Cincinnati, Ohio ABSTRACT Increasing evidence implicates activation of NF-B in a variety of glomerular diseases, but the mechanisms involved are unknown. Here, upregulation of NF-B in the podocytes of transgenic mice resulted in glomer- ulosclerosis and proteinuria. Absence of the podocyte protein nephrin resulted in NF-B activation, suggest- ing that nephrin negatively regulates the NF-B pathway. Signal transduction assays supported a functional relationship between nephrin and NF-B and suggested the involvement of atypical protein kinase C (aPKC//) as an intermediary. We propose that disruption of the slit diaphragm leads to activation of NF-B; subsequent upregulation of NF-B-driven genes results in glomerular damage mediated by NF-B-depen- dent pathways. In summary, nephrin may normally limit NF-B activity in the podocyte, suggesting a mechanism by which it might discourage the evolution of glomerular disease. J Am Soc Nephrol 20: 1733–1743, 2009. doi: 10.1681/ASN.2008111219 NF-B is a transcription factor activated by cell sur- of aPKC/ inhibitory proteins such as Par4 can also face receptor signaling to meet stress and inflam- abrogate NF-B activation.12,13 matory responses, regulating key cellular processes Glomerular disease manifests as urinary protein such as inflammation, innate and adaptive immu- leak resulting from malfunction of the glomerular nity, and cell growth and survival.1 Five mamma- filtration barrier. -

Choose Your Partners: Dimerization in Eukaryotic Transcription Factors
Review Choose your partners: dimerization in eukaryotic transcription factors Grigoris D. Amoutzias1,2, David L. Robertson3, Yves Van de Peer1,2 and Stephen G. Oliver4 1 Department of Plant Systems Biology, VIB, Ghent University, Technologiepark 927, B-9052 Ghent, Belgium 2 Department of Molecular Genetics, Ghent University, Technologiepark 927, B-9052 Ghent, Belgium 3 Faculty of Life Sciences, University of Manchester, Manchester, M13 9PT, UK 4 Department of Biochemistry, University of Cambridge, Sanger Building, 80 Tennis Court Road, Cambridge CB2 1GA, UK In many eukaryotic transcription factor gene families, bZIP: Basic region leucine zipper. This is the second-largest family of proteins require a physical interaction with an identical dimerizing TFs in humans. They are encoded by 51 genes, many of which molecule or with another molecule within the same are well-studied oncogenes. Choanoflagellates: Flagellate unicellular eukaryotes. They are considered to be family to form a functional dimer and bind DNA. Depend- the closest living relatives of the metazoa. The last unicellular ancestors of ing on the choice of partner and the cellular context, each animals are thought to have resembled modern choanoflagellates. dimer triggers a sequence of regulatory events that lead Cnidaria: Animals with radial symmetry and two germ layers, endoderm and ectoderm. Corals, sea anemones and jellyfish are some of the animals that to a particular cellular fate, for example, proliferation or belong to this group. differentiation. Recent syntheses of genomic and func- Enhanceosome: A complex of TFs and other proteins that assemble and bind tional data reveal that partner choice is not random; cooperatively to the enhancer region of a gene.