Overexpression of MLN51 Triggers P-Body Disassembly and Formation of a New Type of RNA Granules N
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Table S1. Upregulated Genes Differentially
Supplementary Table S1. Upregulated genes differentially expressed in athletes (p < 0.05 and 1.3-fold change) Gene Symbol p Value Fold Change 221051_s_at NMRK2 0.01 2.38 236518_at CCDC183 0.00 2.05 218804_at ANO1 0.00 2.05 234675_x_at 0.01 2.02 207076_s_at ASS1 0.00 1.85 209135_at ASPH 0.02 1.81 228434_at BTNL9 0.03 1.81 229985_at BTNL9 0.01 1.79 215795_at MYH7B 0.01 1.78 217979_at TSPAN13 0.01 1.77 230992_at BTNL9 0.01 1.75 226884_at LRRN1 0.03 1.74 220039_s_at CDKAL1 0.01 1.73 236520_at 0.02 1.72 219895_at TMEM255A 0.04 1.72 201030_x_at LDHB 0.00 1.69 233824_at 0.00 1.69 232257_s_at 0.05 1.67 236359_at SCN4B 0.04 1.64 242868_at 0.00 1.63 1557286_at 0.01 1.63 202780_at OXCT1 0.01 1.63 1556542_a_at 0.04 1.63 209992_at PFKFB2 0.04 1.63 205247_at NOTCH4 0.01 1.62 1554182_at TRIM73///TRIM74 0.00 1.61 232892_at MIR1-1HG 0.02 1.61 204726_at CDH13 0.01 1.6 1561167_at 0.01 1.6 1565821_at 0.01 1.6 210169_at SEC14L5 0.01 1.6 236963_at 0.02 1.6 1552880_at SEC16B 0.02 1.6 235228_at CCDC85A 0.02 1.6 1568623_a_at SLC35E4 0.00 1.59 204844_at ENPEP 0.00 1.59 1552256_a_at SCARB1 0.02 1.59 1557283_a_at ZNF519 0.02 1.59 1557293_at LINC00969 0.03 1.59 231644_at 0.01 1.58 228115_at GAREM1 0.01 1.58 223687_s_at LY6K 0.02 1.58 231779_at IRAK2 0.03 1.58 243332_at LOC105379610 0.04 1.58 232118_at 0.01 1.57 203423_at RBP1 0.02 1.57 AMY1A///AMY1B///AMY1C///AMY2A///AMY2B// 208498_s_at 0.03 1.57 /AMYP1 237154_at LOC101930114 0.00 1.56 1559691_at 0.01 1.56 243481_at RHOJ 0.03 1.56 238834_at MYLK3 0.01 1.55 213438_at NFASC 0.02 1.55 242290_at TACC1 0.04 1.55 ANKRD20A1///ANKRD20A12P///ANKRD20A2/// -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -
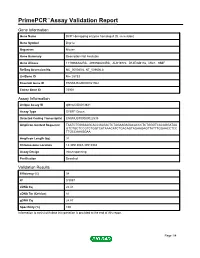
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name DCP1 decapping enzyme homolog A (S. cerevisiae) Gene Symbol Dcp1a Organism Mouse Gene Summary Description Not Available Gene Aliases 1110066A22Rik, 4930568L04Rik, AU019772, D14Ertd817e, Mitc1, SMIF RefSeq Accession No. NC_000080.6, NT_039606.8 UniGene ID Mm.28733 Ensembl Gene ID ENSMUSG00000021962 Entrez Gene ID 75901 Assay Information Unique Assay ID qMmuCID0013841 Assay Type SYBR® Green Detected Coding Transcript(s) ENSMUST00000022535 Amplicon Context Sequence TAATCTGGGAAGCACCGAGACTCTAGAAGAGACACCCTCTGGGTCACAGGATAA GTCTGCTCCGTCTGGTCATAAACATCTGACAGTAGAAGAGTTATTTGGAACCTCC TTGCCAAAGGAA Amplicon Length (bp) 91 Chromosome Location 14:30513043-30518984 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 98 R2 0.9997 cDNA Cq 22.41 cDNA Tm (Celsius) 81 gDNA Cq 24.87 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 1/4 PrimePCR™Assay Validation Report Dcp1a, Mouse Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 2/4 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Mouse Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. Detected Coding Transcript(s) This is a list of the Ensembl transcript ID(s) that this assay will detect. -

Exon Amplification: a Strategy to Isolate Mammalian Genes Based on RNA Splicing (Gene Cloning/Polymerase Chain Reaction) ALAN J
Proc. Natl. Acad. Sci. USA Vol. 88, pp. 4005-4009, May 1991 Genetics Exon amplification: A strategy to isolate mammalian genes based on RNA splicing (gene cloning/polymerase chain reaction) ALAN J. BUCKLER*, DAVID D. CHANG, SHARON L. GRAw, J. DAVID BROOK, DANIEL A. HABER, PHILLIP A. SHARP, AND DAVID E. HOUSMAN Center for Cancer Research, Department of Biology, Massachusetts Institute of Technology, Cambridge, MA 02139 Contributed by Phillip A. Sharp, January 25, 1991 ABSTRACT We have developed a method, exon amplifi- (7, 8). Thus, this method may be generally applicable for the cation, for fast and efficient isolation of coding sequences from selection of exon sequences from any gene. The method is complex mammalian genomic DNA. This method is based on also both rapid and easily adapted to large scale experiments. the selection of RNA sequences, exons, which are flanked by A series of cloned genomic DNA fragments can be screened functional 5' and 3' splice sites. Fragments of cloned genomic within 1-2 weeks. The sensitivity of this method is high. DNA are inserted into an intron, which is flanked by 5' and 3' Genomic DNA segments of 20 kilobases (kb) or more can be splice sites of the human immunodeficiency virus 1 tat gene successfully screened in a single transfection by using a set contained within the plasmid pSPL1. COS-7 cells are trans- of pooled subclones. This method thus allows the rapid fected with these constructs, and the resulting RNA transcripts identification of exons in mammalian genomic DNA and are processed in vivo. Splice sites of exons contained within the should facilitate the isolation of a wide spectrum of genes of inserted genomic fragment are paired with splice sites of the significance in physiology and development. -

Mrna Turnover Philip Mitchell* and David Tollervey†
320 mRNA turnover Philip Mitchell* and David Tollervey† Nuclear RNA-binding proteins can record pre-mRNA are cotransported to the cytoplasm with the mRNP. These processing events in the structure of messenger proteins may preserve a record of the nuclear history of the ribonucleoprotein particles (mRNPs). During initial rounds of pre-mRNA in the cytoplasmic mRNP structure. This infor- translation, the mature mRNP structure is established and is mation can strongly influence the cytoplasmic fate of the monitored by mRNA surveillance systems. Competition for the mRNA and is used by mRNA surveillance systems that act cap structure links translation and subsequent mRNA as a checkpoint of mRNP integrity, particularly in the identi- degradation, which may also involve multiple deadenylases. fication of premature translation termination codons (PTCs). Addresses Cotransport of nuclear mRNA-binding proteins with mRNA Wellcome Trust Centre for Cell Biology, ICMB, University of Edinburgh, from the nucleus to the cytoplasm (nucleocytoplasmic shut- Kings’ Buildings, Edinburgh EH9 3JR, UK tling) was first observed for the heterogeneous nuclear *e-mail: [email protected] ribonucleoprotein (hnRNP) proteins. Some hnRNP proteins †e-mail: [email protected] are stripped from the mRNA at export [1], but hnRNP A1, Current Opinion in Cell Biology 2001, 13:320–325 A2, E, I and K are all exported (see [2]). Although roles for 0955-0674/01/$ — see front matter these hnRNP proteins in transport and translation have been © 2001 Elsevier Science Ltd. All rights reserved. reported [3•,4•], their affects on mRNA stability have been little studied. More is known about hnRNP D/AUF1 and Abbreviations AREs AU-rich sequence elements another nuclear RNA-binding protein, HuR, which act CBC cap-binding complex antagonistically to modulate the stability of a range of DAN deadenylating nuclease mRNAs containing AU-rich sequence elements (AREs) DSEs downstream sequence elements (reviewed in [2]). -

Supplemental Information
Supplemental information Dissection of the genomic structure of the miR-183/96/182 gene. Previously, we showed that the miR-183/96/182 cluster is an intergenic miRNA cluster, located in a ~60-kb interval between the genes encoding nuclear respiratory factor-1 (Nrf1) and ubiquitin-conjugating enzyme E2H (Ube2h) on mouse chr6qA3.3 (1). To start to uncover the genomic structure of the miR- 183/96/182 gene, we first studied genomic features around miR-183/96/182 in the UCSC genome browser (http://genome.UCSC.edu/), and identified two CpG islands 3.4-6.5 kb 5’ of pre-miR-183, the most 5’ miRNA of the cluster (Fig. 1A; Fig. S1 and Seq. S1). A cDNA clone, AK044220, located at 3.2-4.6 kb 5’ to pre-miR-183, encompasses the second CpG island (Fig. 1A; Fig. S1). We hypothesized that this cDNA clone was derived from 5’ exon(s) of the primary transcript of the miR-183/96/182 gene, as CpG islands are often associated with promoters (2). Supporting this hypothesis, multiple expressed sequences detected by gene-trap clones, including clone D016D06 (3, 4), were co-localized with the cDNA clone AK044220 (Fig. 1A; Fig. S1). Clone D016D06, deposited by the German GeneTrap Consortium (GGTC) (http://tikus.gsf.de) (3, 4), was derived from insertion of a retroviral construct, rFlpROSAβgeo in 129S2 ES cells (Fig. 1A and C). The rFlpROSAβgeo construct carries a promoterless reporter gene, the β−geo cassette - an in-frame fusion of the β-galactosidase and neomycin resistance (Neor) gene (5), with a splicing acceptor (SA) immediately upstream, and a polyA signal downstream of the β−geo cassette (Fig. -

Sequences at the Exon-Intron Boundaries* (Split Gene/Mrna Splicing/Eukaryotic Gene Structure) R
Proc. Nati. Acad. Sci. USA Vol. 75, No. 10, pp. 4853-4857, October 1978 Biochemistry Ovalbumin gene: Evidence for a leader sequence in mRNA and DNA sequences at the exon-intron boundaries* (split gene/mRNA splicing/eukaryotic gene structure) R. BREATHNACH, C. BENOIST, K. O'HARE, F. GANNON, AND P. CHAMBON Laboratoire de Genetique Mol6culaire des Eucaryotes du Centre National de la Recherche Scientifique, Unite 44 de l'Institut National de la Sant6 et de la Recherche MWdicale, Institut de Chimie Biologique, Facult6 de Melecine, Strasbourg 67085, France Communicated by A. Frey-Wyssling, July 31, 1978 ABSTRACT Selected regions of cloned EcoRI fragments the 5' end of ov-mRNA and have revealed some interesting of the chicken ovalbumin gene have been sequenced. The po- features in the DNA sequences at exon-intron boundaries. sitions where the sequences coding for ovalbumin mRNA (ov- mRNA) are interrupted in the genome have been determined, and a previously unreported interruption in the DNA sequences MATERIALS AND METHODS coding for the 5' nontranslated region of the messenger has been discovered. Because directly repeated sequences are found at Plasmid pCR1 ov 2.1 containing the ov-ds-cDNA insert was exon-intron boundaries, the nucleotide sequence alone cannot prepared as described (9). EcoRI fragments "b," "c," and "d" define unique excision-ligation points for the processing of a previously cloned in X vectors (3) were transferred to the plas- possible ov-mRNA precursor. However, the sequences in these mid pBR 322. An EcoRI/HindIII of the EcoRI boundary regions share common features; this leads to the fragment proposal that there are, in fact, unique excision-ligation points fragment "a" containing the entirety of exon 7 (Fig. -

Nuclear PTEN Safeguards Pre-Mrna Splicing to Link Golgi Apparatus for Its Tumor Suppressive Role
ARTICLE DOI: 10.1038/s41467-018-04760-1 OPEN Nuclear PTEN safeguards pre-mRNA splicing to link Golgi apparatus for its tumor suppressive role Shao-Ming Shen1, Yan Ji2, Cheng Zhang1, Shuang-Shu Dong2, Shuo Yang1, Zhong Xiong1, Meng-Kai Ge1, Yun Yu1, Li Xia1, Meng Guo1, Jin-Ke Cheng3, Jun-Ling Liu1,3, Jian-Xiu Yu1,3 & Guo-Qiang Chen1 Dysregulation of pre-mRNA alternative splicing (AS) is closely associated with cancers. However, the relationships between the AS and classic oncogenes/tumor suppressors are 1234567890():,; largely unknown. Here we show that the deletion of tumor suppressor PTEN alters pre-mRNA splicing in a phosphatase-independent manner, and identify 262 PTEN-regulated AS events in 293T cells by RNA sequencing, which are associated with significant worse outcome of cancer patients. Based on these findings, we report that nuclear PTEN interacts with the splicing machinery, spliceosome, to regulate its assembly and pre-mRNA splicing. We also identify a new exon 2b in GOLGA2 transcript and the exon exclusion contributes to PTEN knockdown-induced tumorigenesis by promoting dramatic Golgi extension and secretion, and PTEN depletion significantly sensitizes cancer cells to secretion inhibitors brefeldin A and golgicide A. Our results suggest that Golgi secretion inhibitors alone or in combination with PI3K/Akt kinase inhibitors may be therapeutically useful for PTEN-deficient cancers. 1 Department of Pathophysiology, Key Laboratory of Cell Differentiation and Apoptosis of Chinese Ministry of Education, Shanghai Jiao Tong University School of Medicine (SJTU-SM), Shanghai 200025, China. 2 Institute of Health Sciences, Shanghai Institutes for Biological Sciences of Chinese Academy of Sciences and SJTU-SM, Shanghai 200025, China. -

The Sub-Nuclear Localization of RNA-Binding Proteins in KSHV-Infected Cells
cells Article The Sub-Nuclear Localization of RNA-Binding Proteins in KSHV-Infected Cells Ella Alkalay, Chen Gam Ze Letova Refael, Irit Shoval, Noa Kinor, Ronit Sarid and Yaron Shav-Tal * The Mina & Everard Goodman Faculty of Life Sciences and The Institute of Nanotechnology and Advanced Materials, Bar-Ilan University, Ramat Gan 5290002, Israel; [email protected] (E.A.); [email protected] (C.G.Z.L.R.); [email protected] (I.S.); [email protected] (N.K.); [email protected] (R.S.) * Correspondence: [email protected] Received: 14 August 2020; Accepted: 21 August 2020; Published: 25 August 2020 Abstract: RNA-binding proteins, particularly splicing factors, localize to sub-nuclear domains termed nuclear speckles. During certain viral infections, as the nucleus fills up with replicating virus compartments, host cell chromatin distribution changes, ending up condensed at the nuclear periphery. In this study we wished to determine the fate of nucleoplasmic RNA-binding proteins and nuclear speckles during the lytic cycle of the Kaposi’s sarcoma associated herpesvirus (KSHV). We found that nuclear speckles became fewer and dramatically larger, localizing at the nuclear periphery, adjacent to the marginalized chromatin. Enlarged nuclear speckles contained splicing factors, whereas other proteins were nucleoplasmically dispersed. Polyadenylated RNA, typically found in nuclear speckles under regular conditions, was also found in foci separated from nuclear speckles in infected cells. Poly(A) foci did not contain lncRNAs known to colocalize with nuclear speckles but contained the poly(A)-binding protein PABPN1. Examination of the localization of spliced viral RNAs revealed that some spliced transcripts could be detected within the nuclear speckles. -

The Nuclear Poly(A) Binding Protein of Mammals, but Not of Fission Yeast, Participates in Mrna Polyadenylation
Downloaded from rnajournal.cshlp.org on September 30, 2021 - Published by Cold Spring Harbor Laboratory Press REPORT The nuclear poly(A) binding protein of mammals, but not of fission yeast, participates in mRNA polyadenylation UWE KÜHN, JULIANE BUSCHMANN, and ELMAR WAHLE Institute of Biochemistry and Biotechnology, Martin Luther University Halle-Wittenberg, 06099 Halle, Germany ABSTRACT The nuclear poly(A) binding protein (PABPN1) has been suggested, on the basis of biochemical evidence, to play a role in mRNA polyadenylation by strongly increasing the processivity of poly(A) polymerase. While experiments in metazoans have tended to support such a role, the results were not unequivocal, and genetic data show that the S. pombe ortholog of PABPN1, Pab2, is not involved in mRNA polyadenylation. The specific model in which PABPN1 increases the rate of poly(A) tail elongation has never been examined in vivo. Here, we have used 4-thiouridine pulse-labeling to examine the lengths of newly synthesized poly(A) tails in human cells. Knockdown of PABPN1 strongly reduced the synthesis of full-length tails of ∼250 nucleotides, as predicted from biochemical data. We have also purified S. pombe Pab2 and the S. pombe poly(A) polymerase, Pla1, and examined their in vitro activities. Whereas PABPN1 strongly increases the activity of its cognate poly(A) polymerase in vitro, Pab2 was unable to stimulate Pla1 to any significant extent. Thus, in vitro and in vivo data are consistent in supporting a role of PABPN1 but not S. pombe Pab2 in the polyadenylation of mRNA precursors. Keywords: poly(A) binding protein; poly(A) polymerase; mRNA polyadenylation; pre-mRNA 3′; processing INTRODUCTION by poly(A) polymerase with the help of the cleavage and poly- adenylation specificity factor (CPSF), which binds the polya- The poly(A) tails of eukaryotic mRNAs are covered by specif- denylation signal AAUAAA. -

Interaction Profiling of RNA-Binding Ubiquitin Ligases Reveals A
www.nature.com/scientificreports OPEN Interaction profling of RNA- binding ubiquitin ligases reveals a link between posttranscriptional Received: 7 March 2017 Accepted: 14 September 2017 regulation and the ubiquitin system Published: xx xx xxxx Andrea Hildebrandt1, Gregorio Alanis-Lobato1,2, Andrea Voigt1, Kathi Zarnack3, Miguel A. Andrade-Navarro1,2, Petra Beli1 & Julian König1 RNA-binding ubiquitin ligases (RBULs) have the potential to link RNA-mediated mechanisms to protein ubiquitylation. Despite this, the cellular functions, substrates and interaction partners of most RBULs remain poorly characterized. Afnity purifcation (AP) combined with quantitative mass spectrometry (MS)-based proteomics is a powerful approach for analyzing protein functions. Mapping the physiological interaction partners of RNA-binding proteins has been hampered by their intrinsic properties, in particular the existence of low-complexity regions, which are prone to engage in non- physiological interactions. Here, we used an adapted AP approach to identify the interaction partners of human RBULs harboring diferent RNA-binding domains. To increase the likelihood of recovering physiological interactions, we combined control and bait-expressing cells prior to lysis. In this setup, only stable interactions that were originally present in the cell will be identifed. We exploit gene function similarity between the bait proteins and their interactors to benchmark our approach in its ability to recover physiological interactions. We reveal that RBULs engage in stable interactions with RNA-binding proteins involved in diferent steps of RNA metabolism as well as with components of the ubiquitin conjugation machinery and ubiquitin-binding proteins. Our results thus demonstrate their capacity to link posttranscriptional regulation with the ubiquitin system. -

PABPN1 Shuts Down Alternative Poly(A) Sites
Cell Research (2012) 22:1419-1421. npg © 2012 IBCB, SIBS, CAS All rights reserved 1001-0602/12 $ 32.00 RESEARCH HIGHLIGHT www.nature.com/cr PABPN1 shuts down alternative poly(A) sites Martine Simonelig1 1mRNA Regulation and Development, Institute of Human Genetics, CNRS UPR1142, 141 rue de la Cardonille, 34396 Montpellier Cedex 5, France Cell Research (2012) 22:1419-1421. doi:10.1038/cr.2012.86; published online 29 May 2012 Although overlooked for many by two sequences, the canonical poly(A) It has been known for many years years, alternative cleavage and poly- signal AAUAAA localized upstream that, in addition to specific regula- adenylation (APA) is now emerging of the cleavage site and a downstream tors of APA, an important mechanism as a major mechanism of gene regula- U/GU-rich motif; both motifs coop- underlying APA involves changes in tion. A recent study identifies poly(A)- eratively recruit the polyadenylation the concentration of general cleavage binding protein nuclear 1 (PABPN1), machinery through direct interactions and polyadenylation factors, CstF be- a general factor of polyadenylation, with CPSF and CstF, respectively. ing the first complex to be implicated as a suppressor of alternative poly(A) Recent studies using genome-wide in this regulation [4-7]. From these sites. approaches have revealed alternative analyses, a general view emerged that mRNA 3′-end processing is a cotran- cleavage and polyadenylation (APA) promoter-proximal poly(A) sites tend to scriptional reaction that leads to the to be very widespread. More than 50% be weaker than distal poly(A) sites, and addition of a poly(A) tail – polyade- of human genes generate multiple tran- that increased levels of the core cleav- nylation – to virtually all eukaryotic scripts with different 3′ UTRs resulting age and polyadenylation machinery mRNAs.