Targeting Altered Nme Heterooligomerization in Disease?
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

In Silico Prediction of High-Resolution Hi-C Interaction Matrices
ARTICLE https://doi.org/10.1038/s41467-019-13423-8 OPEN In silico prediction of high-resolution Hi-C interaction matrices Shilu Zhang1, Deborah Chasman 1, Sara Knaack1 & Sushmita Roy1,2* The three-dimensional (3D) organization of the genome plays an important role in gene regulation bringing distal sequence elements in 3D proximity to genes hundreds of kilobases away. Hi-C is a powerful genome-wide technique to study 3D genome organization. Owing to 1234567890():,; experimental costs, high resolution Hi-C datasets are limited to a few cell lines. Computa- tional prediction of Hi-C counts can offer a scalable and inexpensive approach to examine 3D genome organization across multiple cellular contexts. Here we present HiC-Reg, an approach to predict contact counts from one-dimensional regulatory signals. HiC-Reg pre- dictions identify topologically associating domains and significant interactions that are enri- ched for CCCTC-binding factor (CTCF) bidirectional motifs and interactions identified from complementary sources. CTCF and chromatin marks, especially repressive and elongation marks, are most important for HiC-Reg’s predictive performance. Taken together, HiC-Reg provides a powerful framework to generate high-resolution profiles of contact counts that can be used to study individual locus level interactions and higher-order organizational units of the genome. 1 Wisconsin Institute for Discovery, 330 North Orchard Street, Madison, WI 53715, USA. 2 Department of Biostatistics and Medical Informatics, University of Wisconsin-Madison, Madison, WI 53715, USA. *email: [email protected] NATURE COMMUNICATIONS | (2019) 10:5449 | https://doi.org/10.1038/s41467-019-13423-8 | www.nature.com/naturecommunications 1 ARTICLE NATURE COMMUNICATIONS | https://doi.org/10.1038/s41467-019-13423-8 he three-dimensional (3D) organization of the genome has Results Temerged as an important component of the gene regulation HiC-Reg for predicting contact count using Random Forests. -

Download Download
Supplementary Figure S1. Results of flow cytometry analysis, performed to estimate CD34 positivity, after immunomagnetic separation in two different experiments. As monoclonal antibody for labeling the sample, the fluorescein isothiocyanate (FITC)- conjugated mouse anti-human CD34 MoAb (Mylteni) was used. Briefly, cell samples were incubated in the presence of the indicated MoAbs, at the proper dilution, in PBS containing 5% FCS and 1% Fc receptor (FcR) blocking reagent (Miltenyi) for 30 min at 4 C. Cells were then washed twice, resuspended with PBS and analyzed by a Coulter Epics XL (Coulter Electronics Inc., Hialeah, FL, USA) flow cytometer. only use Non-commercial 1 Supplementary Table S1. Complete list of the datasets used in this study and their sources. GEO Total samples Geo selected GEO accession of used Platform Reference series in series samples samples GSM142565 GSM142566 GSM142567 GSM142568 GSE6146 HG-U133A 14 8 - GSM142569 GSM142571 GSM142572 GSM142574 GSM51391 GSM51392 GSE2666 HG-U133A 36 4 1 GSM51393 GSM51394 only GSM321583 GSE12803 HG-U133A 20 3 GSM321584 2 GSM321585 use Promyelocytes_1 Promyelocytes_2 Promyelocytes_3 Promyelocytes_4 HG-U133A 8 8 3 GSE64282 Promyelocytes_5 Promyelocytes_6 Promyelocytes_7 Promyelocytes_8 Non-commercial 2 Supplementary Table S2. Chromosomal regions up-regulated in CD34+ samples as identified by the LAP procedure with the two-class statistics coded in the PREDA R package and an FDR threshold of 0.5. Functional enrichment analysis has been performed using DAVID (http://david.abcc.ncifcrf.gov/) -

Identification of Conserved Genes Triggering Puberty in European Sea
Blázquez et al. BMC Genomics (2017) 18:441 DOI 10.1186/s12864-017-3823-2 RESEARCHARTICLE Open Access Identification of conserved genes triggering puberty in European sea bass males (Dicentrarchus labrax) by microarray expression profiling Mercedes Blázquez1,2* , Paula Medina1,2,3, Berta Crespo1,4, Ana Gómez1 and Silvia Zanuy1* Abstract Background: Spermatogenesisisacomplexprocesscharacterized by the activation and/or repression of a number of genes in a spatio-temporal manner. Pubertal development in males starts with the onset of the first spermatogenesis and implies the division of primary spermatogonia and their subsequent entry into meiosis. This study is aimed at the characterization of genes involved in the onset of puberty in European sea bass, and constitutes the first transcriptomic approach focused on meiosis in this species. Results: European sea bass testes collected at the onset of puberty (first successful reproduction) were grouped in stage I (resting stage), and stage II (proliferative stage). Transition from stage I to stage II was marked by an increase of 11ketotestosterone (11KT), the main fish androgen, whereas the transcriptomic study resulted in 315 genes differentially expressed between the two stages. The onset of puberty induced 1) an up-regulation of genes involved in cell proliferation, cell cycle and meiosis progression, 2) changes in genes related with reproduction and growth, and 3) a down-regulation of genes included in the retinoic acid (RA) signalling pathway. The analysis of GO-terms and biological pathways showed that cell cycle, cell division, cellular metabolic processes, and reproduction were affected, consistent with the early events that occur during the onset of puberty. -

Characterization of the Small RNA Transcriptomes of Cell Protrusions and Cell Bodies of Highly Metastatic Hepatocellular Carcinoma Cells Via RNA Sequencing
ONCOLOGY LETTERS 22: 568, 2021 Characterization of the small RNA transcriptomes of cell protrusions and cell bodies of highly metastatic hepatocellular carcinoma cells via RNA sequencing WENPIN CAI1*, JINGZHANG JI2*, BITING WU2*, KAIXUAN HAO2, PING REN2, YU JIN2, LIHONG YANG2, QINGCHAO TONG2 and ZHIFA SHEN2 1Department of Laboratory Medicine, Wen Zhou Traditional Chinese Medicine Hospital; 2Zhejiang Provincial Key Laboratory of Medical Genetics, Key Laboratory of Laboratory Medicine, Ministry of Education, School of Laboratory Medicine and Life Sciences, Wenzhou Medical University, Wenzhou, Zhejiang 325035, P.R. China Received July 11, 2020; Accepted February 23, 2021 DOI: 10.3892/ol.2021.12829 Abstract. Increasing evidence suggest that hepatocellular differentially expressed miRNAs and circRNAs. The interac‑ carcinoma (HCC) HCCLM3 cells initially develop pseudo‑ tion maps between miRNAs and circRNAs were constructed, podia when they metastasize, and microRNAs (miRNAs/miRs) and signaling pathway maps were analyzed to determine the and circular RNAs (circRNAs) have been demonstrated to molecular mechanism and regulation of the differentially serve important roles in the development, progression and expressed miRNAs and circRNAs. Taken together, the results metastasis of cancer. The present study aimed to isolate the of the present study suggest that the Boyden chamber assay cell bodies (CBs) and cell protrusions (CPs) from HCCLM3 can be used to effectively isolate the somatic CBs and CPs of cells, and screen the miRNAs and circRNAs associated with HCC, which can be used to screen the miRNAs and circRNAs HCC infiltration and metastasis in CBs and CPs. The Boyden associated with invasion and metastasis of HCC. chamber assay has been confirmed to effectively isolate the CBs and CPs from HCCLM3 cells via observation of microtu‑ Introduction bule immunofluorescence, DAPI staining and nuclear protein H3 western blotting. -

Supplementary Materials
Supplementary materials Supplementary Table S1: MGNC compound library Ingredien Molecule Caco- Mol ID MW AlogP OB (%) BBB DL FASA- HL t Name Name 2 shengdi MOL012254 campesterol 400.8 7.63 37.58 1.34 0.98 0.7 0.21 20.2 shengdi MOL000519 coniferin 314.4 3.16 31.11 0.42 -0.2 0.3 0.27 74.6 beta- shengdi MOL000359 414.8 8.08 36.91 1.32 0.99 0.8 0.23 20.2 sitosterol pachymic shengdi MOL000289 528.9 6.54 33.63 0.1 -0.6 0.8 0 9.27 acid Poricoic acid shengdi MOL000291 484.7 5.64 30.52 -0.08 -0.9 0.8 0 8.67 B Chrysanthem shengdi MOL004492 585 8.24 38.72 0.51 -1 0.6 0.3 17.5 axanthin 20- shengdi MOL011455 Hexadecano 418.6 1.91 32.7 -0.24 -0.4 0.7 0.29 104 ylingenol huanglian MOL001454 berberine 336.4 3.45 36.86 1.24 0.57 0.8 0.19 6.57 huanglian MOL013352 Obacunone 454.6 2.68 43.29 0.01 -0.4 0.8 0.31 -13 huanglian MOL002894 berberrubine 322.4 3.2 35.74 1.07 0.17 0.7 0.24 6.46 huanglian MOL002897 epiberberine 336.4 3.45 43.09 1.17 0.4 0.8 0.19 6.1 huanglian MOL002903 (R)-Canadine 339.4 3.4 55.37 1.04 0.57 0.8 0.2 6.41 huanglian MOL002904 Berlambine 351.4 2.49 36.68 0.97 0.17 0.8 0.28 7.33 Corchorosid huanglian MOL002907 404.6 1.34 105 -0.91 -1.3 0.8 0.29 6.68 e A_qt Magnogrand huanglian MOL000622 266.4 1.18 63.71 0.02 -0.2 0.2 0.3 3.17 iolide huanglian MOL000762 Palmidin A 510.5 4.52 35.36 -0.38 -1.5 0.7 0.39 33.2 huanglian MOL000785 palmatine 352.4 3.65 64.6 1.33 0.37 0.7 0.13 2.25 huanglian MOL000098 quercetin 302.3 1.5 46.43 0.05 -0.8 0.3 0.38 14.4 huanglian MOL001458 coptisine 320.3 3.25 30.67 1.21 0.32 0.9 0.26 9.33 huanglian MOL002668 Worenine -

Two Separate Functions of NME3 Critical for Cell Survival Underlie a Neurodegenerative Disorder
Two separate functions of NME3 critical for cell survival underlie a neurodegenerative disorder Chih-Wei Chena, Hong-Ling Wanga, Ching-Wen Huangb, Chang-Yu Huangc, Wai Keong Lima, I-Chen Tua, Atmaja Koorapatia, Sung-Tsang Hsiehd, Hung-Wei Kand, Shiou-Ru Tzenge, Jung-Chi Liaof, Weng Man Chongf, Inna Naroditzkyg, Dvora Kidronh,i, Ayelet Eranj, Yousif Nijimk, Ella Selak, Hagit Baris Feldmanl, Limor Kalfonm, Hadas Raveh-Barakm, Tzipora C. Falik-Zaccaim,n, Orly Elpelego, Hanna Mandelm,p,1, and Zee-Fen Changa,q,1 aInstitute of Molecular Medicine, College of Medicine, National Taiwan University, 10002 Taipei, Taiwan; bDepartment of Medicine, College of Medicine, National Taiwan University, 10002 Taipei, Taiwan; cInstitute of Biochemistry and Molecular Biology, National Yang-Ming University, 11221 Taipei, Taiwan; dInstitute of Anatomy and Cell Biology, College of Medicine, National Taiwan University, 10002 Taipei, Taiwan; eInstitute of Biochemistry and Molecular Biology, College of Medicine, National Taiwan University, 10002 Taipei, Taiwan; fInstitute of Atomic and Molecular Sciences, Academia Sinica, 10617 Taipei, Taiwan; gDepartment of Pathology, Rambam Health Care Campus, 31096 Haifa, Israel; hDepartment of Pathology, Meir Hospital, 44100 Kfar Saba, Israel; iSackler School of Medicine, Tel Aviv University, 69978 Tel Aviv, Israel; jDepartment of Radiology, Rambam Health Care Campus, 31096 Haifa, Israel; kPediatric and Neonatal Unit, Nazareth Hospital EMMS, 17639 Nazareth, Israel; lThe Genetics Institute, Rambam Health Care Campus, 31096 Haifa, Israel; mInstitute of Human Genetics, Galilee Medical Center, 22100 Nahariya, Israel; nThe Azrieli Faculty of Medicine, Bar Ilan University, 13100 Safed, Israel; oDepartment of Genetic and Metabolic Diseases, Hadassah Hebrew University Medical Center, 91120 Jerusalem, Israel; pMetabolic Unit, Technion Faculty of Medicine, Rambam Health Care Campus, 31096 Haifa, Israel; and qCenter of Precision Medicine, College of Medicine, National Taiwan University, 10002 Taipei, Taiwan Edited by Christopher K. -
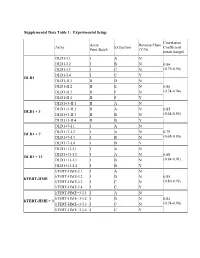
Table 3: Average Gene Expression Profiles by Chromosome
Supplemental Data Table 1: Experimental Setup Correlation Array Reverse Fluor Array Extraction Coefficient Print Batch (Y/N) mean (range) DLD1-I.1 I A N DLD1-I.2 I B N 0.86 DLD1-I.3 I C N (0.79-0.90) DLD1-I.4 I C Y DLD1 DLD1-II.1 II D N DLD1-II.2 II E N 0.86 DLD1-II.3 II F N (0.74-0.94) DLD1-II.4 II F Y DLD1+3-II.1 II A N DLD1+3-II.2 II A N 0.85 DLD1 + 3 DLD1+3-II.3 II B N (0.64-0.95) DLD1+3-II.4 II B Y DLD1+7-I.1 I A N DLD1+7-I.2 I A N 0.79 DLD1 + 7 DLD1+7-I.3 I B N (0.68-0.90) DLD1+7-I.4 I B Y DLD1+13-I.1 I A N DLD1+13-I.2 I A N 0.88 DLD1 + 13 DLD1+13-I.3 I B N (0.84-0.91) DLD1+13-I.4 I B Y hTERT-HME-I.1 I A N hTERT-HME-I.2 I B N 0.85 hTERT-HME hTERT-HME-I.3 I C N (0.80-0.92) hTERT-HME-I.4 I C Y hTERT-HME+3-I.1 I A N hTERT-HME+3-I.2 I B N 0.84 hTERT-HME + 3 hTERT-HME+3-I.3 I C N (0.74-0.90) hTERT-HME+3-I.4 I C Y Supplemental Data Table 2: Average gene expression profiles by chromosome arm DLD1 hTERT-HME Ratio.7 Ratio.1 Ratio.3 Ratio.3 Chrom. -

Datasheet: VMA00345KT Product Details
Datasheet: VMA00345KT Description: NUCLEOSIDE DIPHOSPHATE KINASE A ANTIBODY WITH CONTROL LYSATE Specificity: NUCLEOSIDE DIPHOSPHATE KINASE A Format: Purified Product Type: PrecisionAb™ Monoclonal Isotype: IgG2b Quantity: 2 Westerns Product Details Applications This product has been reported to work in the following applications. This information is derived from testing within our laboratories, peer-reviewed publications or personal communications from the originators. Please refer to references indicated for further information. For general protocol recommendations, please visit www.bio-rad-antibodies.com/protocols. Yes No Not Determined Suggested Dilution Western Blotting 1/1000 PrecisionAb antibodies have been extensively validated for the western blot application. The antibody has been validated at the suggested dilution. Where this product has not been tested for use in a particular technique this does not necessarily exclude its use in such procedures. Further optimization may be required dependant on sample type. Target Species Human Species Cross Reacts with: Mouse, Rat Reactivity N.B. Antibody reactivity and working conditions may vary between species. Product Form Purified IgG - liquid Preparation 20μl Mouse monoclonal antibody prepared by affinity chromatography on Protein G Buffer Solution Phosphate buffered saline Preservative 0.09% Sodium Azide (NaN3) Stabilisers Immunogen Recombinant human nucleoside diphosphate kinase A External Database Links UniProt: P15531 Related reagents Entrez Gene: 4830 NME1 Related reagents Synonyms NDPKA, NM23 Page 1 of 3 Specificity Mouse anti Human nucleoside diphosphate kinase A antibody recognizes nucleoside diphosphate kinase A, also known as NDP kinase A, granzyme A-activated DNase, metastasis inhibition factor nm23, non-metastatic cells 1 and tumor metastatic process-associated protein. The NME1 gene was identified because of its reduced mRNA transcript levels in highly metastatic cells. -

Databases Featuring Genomic Sequence Alignment
D466–D470 Nucleic Acids Research, 2005, Vol. 33, Database issue doi:10.1093/nar/gki045 Improvements to GALA and dbERGE II: databases featuring genomic sequence alignment, annotation and experimental results Laura Elnitski1,2,*, Belinda Giardine1, Prachi Shah1, Yi Zhang1, Cathy Riemer1, Matthew Weirauch4, Richard Burhans1, Webb Miller1,3 and Ross C. Hardison2 1Department of Computer Science and Engineering, 2Department of Biochemistry and Molecular Biology and 3Department of Biology, The Pennsylvania State University, University Park, PA 16802, USA and 4Department of Computer Science, University of California, Santa Cruz, CA 95064, USA Received August 14, 2004; Revised and Accepted September 28, 2004 ABSTRACT gene expression patterns, requires bioinformatic tools for organization and interpretation. Genome browsers, such as We describe improvements to two databases that give the UCSC Genome Browser (1,2), Ensembl (3) and Map access to information on genomic sequence similar- Viewer at NCBI (4), provide views of genes and genomic ities, functional elements in DNA and experimental regions with user-selected annotations. results that demonstrate those functions. GALA, the To provide querying capacity across data types, we devel- database of Genome ALignments and Annotations, is oped GALA, a Genome Alignment and Annotation database. now a set of interlinked relational databases for five The first release recorded whole-genome human–mouse align- vertebrate species, human, chimpanzee, mouse, rat ments along with extensive annotation of the human genome and chicken. For each species, GALA records pair- in a relational database (5). An example of the use of GALA is wise and multiple sequence alignments, scores to find highly conserved regions that do not code for proteins; derived from those alignments that reflect the likeli- some of these could have novel functions. -

2014-Platform-Abstracts.Pdf
American Society of Human Genetics 64th Annual Meeting October 18–22, 2014 San Diego, CA PLATFORM ABSTRACTS Abstract Abstract Numbers Numbers Saturday 41 Statistical Methods for Population 5:30pm–6:50pm: Session 2: Plenary Abstracts Based Studies Room 20A #198–#205 Featured Presentation I (4 abstracts) Hall B1 #1–#4 42 Genome Variation and its Impact on Autism and Brain Development Room 20BC #206–#213 Sunday 43 ELSI Issues in Genetics Room 20D #214–#221 1:30pm–3:30pm: Concurrent Platform Session A (12–21): 44 Prenatal, Perinatal, and Reproductive 12 Patterns and Determinants of Genetic Genetics Room 28 #222–#229 Variation: Recombination, Mutation, 45 Advances in Defining the Molecular and Selection Hall B1 Mechanisms of Mendelian Disorders Room 29 #230–#237 #5-#12 13 Genomic Studies of Autism Room 6AB #13–#20 46 Epigenomics of Normal Populations 14 Statistical Methods for Pedigree- and Disease States Room 30 #238–#245 Based Studies Room 6CF #21–#28 15 Prostate Cancer: Expression Tuesday Informing Risk Room 6DE #29–#36 8:00pm–8:25am: 16 Variant Calling: What Makes the 47 Plenary Abstracts Featured Difference? Room 20A #37–#44 Presentation III Hall BI #246 17 New Genes, Incidental Findings and 10:30am–12:30pm:Concurrent Platform Session D (49 – 58): Unexpected Observations Revealed 49 Detailing the Parts List Using by Exome Sequencing Room 20BC #45–#52 Genomic Studies Hall B1 #247–#254 18 Type 2 Diabetes Genetics Room 20D #53–#60 50 Statistical Methods for Multigene, 19 Genomic Methods in Clinical Practice Room 28 #61–#68 Gene Interaction -

Global Screening of Sentrin-Specific Protease Family Substrates in Sumoylation
bioRxiv preprint doi: https://doi.org/10.1101/2020.02.25.964072; this version posted February 26, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Global Screening of Sentrin-Specific Protease Family Substrates in SUMOylation Yunzhi Wang, 1, 4 Xiaohui Wu,1, 4 Rui Ge,1, 4 Lei Song, 2 Kai Li, 2 Sha Tian,1 Lili Cai, 1 Mingwei Liu, 2 Wenhao Shi, 2Guoying Yu, 3 Bei Zhen, 2 Yi Wang, 2 Fuchu He, 1, 2, * Jun Qin, 1, 2, * Chen Ding1, 2, * 1State Key Laboratory of Genetic Engineering and Collaborative Innovation Center for Genetics and Development, School of Life Sciences, Institutes of Biomedical Sciences, Zhongshan Hospital, Fudan University, Shanghai, 200432, China 2State Key Laboratory of Proteomics, Beijing Proteome Research Center, National Center for Protein Sciences (Beijing), Beijing Institute of Lifeomics, Beijing, 102206, China 3State Key Laboratory of Cell Differentiation and Regulation, College of Life Science, Henan Normal University, Xinxiang, Henan 453007 4Co-first author *Correspondence: [email protected] (C.D.); [email protected] (J.Q.); [email protected] (F.H.); Key Words 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.02.25.964072; this version posted February 26, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Posttranslational modification; SUMO; Sentrin/SUMO-specific proteases; Mass Spectrometry; Innate Immunity Abstract Post-translational modification of proteins by the addition of small ubiquitin-related modifier (SUMO) is a dynamic process, in which deSUMOylation is carried out by members of the Sentrin/SUMO-specific protease (SENP) family. -

Identi Cation of Crucial Genes in Metastatic Osteosarcoma And
Identication of crucial genes in metastatic osteosarcoma and prognostic biomarker in osteosarcoma patients Weihao Wang First Aliated Hospital of Shantou University Medical College Zhaoyong Liu First Aliated Hospital of Shantou University Medical College Weiqing Lu First Aliated Hospital of Shantou University Medical College Huancheng Guo First Aliated Hospital of Shantou University Medical College Chunbin Zhou First Aliated Hospital of Shantou University Medical College Youbin Lin First Aliated Hospital of Shantou University Medical College Hu Wang ( [email protected] ) the First Aliated Hospital of Shantou University Medical Collegethe First Aliated Hospital of Shantou University Medical Collegethe First Aliated Hospital of Shantou University Medical Collegethe First Aliated Hospital of Shantou University Med Research article Keywords: osteosarcoma, metastasis, crucial genes, prognostic biomarker Posted Date: July 22nd, 2020 DOI: https://doi.org/10.21203/rs.3.rs-44119/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/19 Abstract Background Osteosarcoma is still a challenging cancer that poses a huge threat to human health worldwide, especially in young people. The aim of this study was to identify prognostic genes associated with neoplasm metastasis in osteosarcoma, and construct a prognostic nomogram to help the orthopedist to assess prognosis of osteosarcoma patients. Methods Gene expression and clinical information were extracted from the TARGET database. Differentially expressed genes (DEGs) associated with osteosarcoma metastasis were identied and subjected to GO functional and KEGG pathway enrichment analyses. Survival and Cox analyses were used to identify prognostic genes. Results First, 214 DEGs were identied as crucial genes associated with osteosarcoma metastasis.