Anti-NME4 Monoclonal Antibody (DCABH-12587) This Product Is for Research Use Only and Is Not Intended for Diagnostic Use
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

In Silico Prediction of High-Resolution Hi-C Interaction Matrices
ARTICLE https://doi.org/10.1038/s41467-019-13423-8 OPEN In silico prediction of high-resolution Hi-C interaction matrices Shilu Zhang1, Deborah Chasman 1, Sara Knaack1 & Sushmita Roy1,2* The three-dimensional (3D) organization of the genome plays an important role in gene regulation bringing distal sequence elements in 3D proximity to genes hundreds of kilobases away. Hi-C is a powerful genome-wide technique to study 3D genome organization. Owing to 1234567890():,; experimental costs, high resolution Hi-C datasets are limited to a few cell lines. Computa- tional prediction of Hi-C counts can offer a scalable and inexpensive approach to examine 3D genome organization across multiple cellular contexts. Here we present HiC-Reg, an approach to predict contact counts from one-dimensional regulatory signals. HiC-Reg pre- dictions identify topologically associating domains and significant interactions that are enri- ched for CCCTC-binding factor (CTCF) bidirectional motifs and interactions identified from complementary sources. CTCF and chromatin marks, especially repressive and elongation marks, are most important for HiC-Reg’s predictive performance. Taken together, HiC-Reg provides a powerful framework to generate high-resolution profiles of contact counts that can be used to study individual locus level interactions and higher-order organizational units of the genome. 1 Wisconsin Institute for Discovery, 330 North Orchard Street, Madison, WI 53715, USA. 2 Department of Biostatistics and Medical Informatics, University of Wisconsin-Madison, Madison, WI 53715, USA. *email: [email protected] NATURE COMMUNICATIONS | (2019) 10:5449 | https://doi.org/10.1038/s41467-019-13423-8 | www.nature.com/naturecommunications 1 ARTICLE NATURE COMMUNICATIONS | https://doi.org/10.1038/s41467-019-13423-8 he three-dimensional (3D) organization of the genome has Results Temerged as an important component of the gene regulation HiC-Reg for predicting contact count using Random Forests. -

Download Download
Supplementary Figure S1. Results of flow cytometry analysis, performed to estimate CD34 positivity, after immunomagnetic separation in two different experiments. As monoclonal antibody for labeling the sample, the fluorescein isothiocyanate (FITC)- conjugated mouse anti-human CD34 MoAb (Mylteni) was used. Briefly, cell samples were incubated in the presence of the indicated MoAbs, at the proper dilution, in PBS containing 5% FCS and 1% Fc receptor (FcR) blocking reagent (Miltenyi) for 30 min at 4 C. Cells were then washed twice, resuspended with PBS and analyzed by a Coulter Epics XL (Coulter Electronics Inc., Hialeah, FL, USA) flow cytometer. only use Non-commercial 1 Supplementary Table S1. Complete list of the datasets used in this study and their sources. GEO Total samples Geo selected GEO accession of used Platform Reference series in series samples samples GSM142565 GSM142566 GSM142567 GSM142568 GSE6146 HG-U133A 14 8 - GSM142569 GSM142571 GSM142572 GSM142574 GSM51391 GSM51392 GSE2666 HG-U133A 36 4 1 GSM51393 GSM51394 only GSM321583 GSE12803 HG-U133A 20 3 GSM321584 2 GSM321585 use Promyelocytes_1 Promyelocytes_2 Promyelocytes_3 Promyelocytes_4 HG-U133A 8 8 3 GSE64282 Promyelocytes_5 Promyelocytes_6 Promyelocytes_7 Promyelocytes_8 Non-commercial 2 Supplementary Table S2. Chromosomal regions up-regulated in CD34+ samples as identified by the LAP procedure with the two-class statistics coded in the PREDA R package and an FDR threshold of 0.5. Functional enrichment analysis has been performed using DAVID (http://david.abcc.ncifcrf.gov/) -
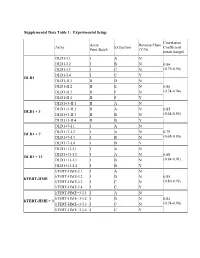
Table 3: Average Gene Expression Profiles by Chromosome
Supplemental Data Table 1: Experimental Setup Correlation Array Reverse Fluor Array Extraction Coefficient Print Batch (Y/N) mean (range) DLD1-I.1 I A N DLD1-I.2 I B N 0.86 DLD1-I.3 I C N (0.79-0.90) DLD1-I.4 I C Y DLD1 DLD1-II.1 II D N DLD1-II.2 II E N 0.86 DLD1-II.3 II F N (0.74-0.94) DLD1-II.4 II F Y DLD1+3-II.1 II A N DLD1+3-II.2 II A N 0.85 DLD1 + 3 DLD1+3-II.3 II B N (0.64-0.95) DLD1+3-II.4 II B Y DLD1+7-I.1 I A N DLD1+7-I.2 I A N 0.79 DLD1 + 7 DLD1+7-I.3 I B N (0.68-0.90) DLD1+7-I.4 I B Y DLD1+13-I.1 I A N DLD1+13-I.2 I A N 0.88 DLD1 + 13 DLD1+13-I.3 I B N (0.84-0.91) DLD1+13-I.4 I B Y hTERT-HME-I.1 I A N hTERT-HME-I.2 I B N 0.85 hTERT-HME hTERT-HME-I.3 I C N (0.80-0.92) hTERT-HME-I.4 I C Y hTERT-HME+3-I.1 I A N hTERT-HME+3-I.2 I B N 0.84 hTERT-HME + 3 hTERT-HME+3-I.3 I C N (0.74-0.90) hTERT-HME+3-I.4 I C Y Supplemental Data Table 2: Average gene expression profiles by chromosome arm DLD1 hTERT-HME Ratio.7 Ratio.1 Ratio.3 Ratio.3 Chrom. -

Databases Featuring Genomic Sequence Alignment
D466–D470 Nucleic Acids Research, 2005, Vol. 33, Database issue doi:10.1093/nar/gki045 Improvements to GALA and dbERGE II: databases featuring genomic sequence alignment, annotation and experimental results Laura Elnitski1,2,*, Belinda Giardine1, Prachi Shah1, Yi Zhang1, Cathy Riemer1, Matthew Weirauch4, Richard Burhans1, Webb Miller1,3 and Ross C. Hardison2 1Department of Computer Science and Engineering, 2Department of Biochemistry and Molecular Biology and 3Department of Biology, The Pennsylvania State University, University Park, PA 16802, USA and 4Department of Computer Science, University of California, Santa Cruz, CA 95064, USA Received August 14, 2004; Revised and Accepted September 28, 2004 ABSTRACT gene expression patterns, requires bioinformatic tools for organization and interpretation. Genome browsers, such as We describe improvements to two databases that give the UCSC Genome Browser (1,2), Ensembl (3) and Map access to information on genomic sequence similar- Viewer at NCBI (4), provide views of genes and genomic ities, functional elements in DNA and experimental regions with user-selected annotations. results that demonstrate those functions. GALA, the To provide querying capacity across data types, we devel- database of Genome ALignments and Annotations, is oped GALA, a Genome Alignment and Annotation database. now a set of interlinked relational databases for five The first release recorded whole-genome human–mouse align- vertebrate species, human, chimpanzee, mouse, rat ments along with extensive annotation of the human genome and chicken. For each species, GALA records pair- in a relational database (5). An example of the use of GALA is wise and multiple sequence alignments, scores to find highly conserved regions that do not code for proteins; derived from those alignments that reflect the likeli- some of these could have novel functions. -

2014-Platform-Abstracts.Pdf
American Society of Human Genetics 64th Annual Meeting October 18–22, 2014 San Diego, CA PLATFORM ABSTRACTS Abstract Abstract Numbers Numbers Saturday 41 Statistical Methods for Population 5:30pm–6:50pm: Session 2: Plenary Abstracts Based Studies Room 20A #198–#205 Featured Presentation I (4 abstracts) Hall B1 #1–#4 42 Genome Variation and its Impact on Autism and Brain Development Room 20BC #206–#213 Sunday 43 ELSI Issues in Genetics Room 20D #214–#221 1:30pm–3:30pm: Concurrent Platform Session A (12–21): 44 Prenatal, Perinatal, and Reproductive 12 Patterns and Determinants of Genetic Genetics Room 28 #222–#229 Variation: Recombination, Mutation, 45 Advances in Defining the Molecular and Selection Hall B1 Mechanisms of Mendelian Disorders Room 29 #230–#237 #5-#12 13 Genomic Studies of Autism Room 6AB #13–#20 46 Epigenomics of Normal Populations 14 Statistical Methods for Pedigree- and Disease States Room 30 #238–#245 Based Studies Room 6CF #21–#28 15 Prostate Cancer: Expression Tuesday Informing Risk Room 6DE #29–#36 8:00pm–8:25am: 16 Variant Calling: What Makes the 47 Plenary Abstracts Featured Difference? Room 20A #37–#44 Presentation III Hall BI #246 17 New Genes, Incidental Findings and 10:30am–12:30pm:Concurrent Platform Session D (49 – 58): Unexpected Observations Revealed 49 Detailing the Parts List Using by Exome Sequencing Room 20BC #45–#52 Genomic Studies Hall B1 #247–#254 18 Type 2 Diabetes Genetics Room 20D #53–#60 50 Statistical Methods for Multigene, 19 Genomic Methods in Clinical Practice Room 28 #61–#68 Gene Interaction -

ATR16 Syndrome: Mechanisms Linking Monosomy to Phenotype 2 3 4 Christian Babbs,*1 Jill Brown,1 Sharon W
bioRxiv preprint doi: https://doi.org/10.1101/768895; this version posted October 7, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 ATR16 Syndrome: Mechanisms Linking Monosomy to Phenotype 2 3 4 Christian Babbs,*1 Jill Brown,1 Sharon W. Horsley,1 Joanne Slater,1 Evie Maifoshie,1 5 Shiwangini Kumar,2 Paul Ooijevaar,3 Marjolein Kriek,3 Amanda Dixon-McIver,2 6 Cornelis L. Harteveld,3 Joanne Traeger-Synodinos,4 Douglas Higgs1 and Veronica 7 Buckle*1 8 9 10 11 12 1. MRC Molecular Haematology Unit, MRC Weatherall Institute of Molecular 13 Medicine, University of Oxford, Oxford, UK. 14 15 2. IGENZ Ltd, Auckland, New Zealand. 16 17 3. Department of Clinical Genetics, Leiden University Medical Center, Leiden, The 18 Netherlands. 19 20 4. Department of Medical Genetics, National and Kapodistrian University of Athens, 21 Greece. 22 23 24 25 *Authors for correspondence 26 [email protected] 27 [email protected] 28 29 1 bioRxiv preprint doi: https://doi.org/10.1101/768895; this version posted October 7, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Abstract 2 3 Background: Sporadic deletions removing 100s-1000s kb of DNA, and 4 variable numbers of poorly characterised genes, are often found in patients 5 with a wide range of developmental abnormalities. In such cases, 6 understanding the contribution of the deletion to an individual’s clinical 7 phenotype is challenging. -

Adventitious Changes in Long-Range Gene Expression Caused by Polymorphic Structural Variation and Promoter Competition
Adventitious changes in long-range gene expression caused by polymorphic structural variation and promoter competition Karen M. Lowera, Jim R. Hughesa, Marco De Gobbia, Shirley Hendersonb, Vip Viprakasitc, Chris Fishera, Anne Gorielya, Helena Ayyuba, Jackie Sloane-Stanleya, Douglas Vernimmena, Cordelia Langfordd, David Garricka, Richard J. Gibbonsa, and Douglas R. Higgsa,1 aMedical Research Council Molecular Haematology Unit, Weatherall Institute of Molecular Medicine, The John Radcliffe Hospital, Headington, Oxford, OX3 9DS, United Kingdom; bNational Haemoglobinopathy Reference Laboratory, Oxford Radcliffe Hospitals NHS Trust, Oxford, OX3 7LJ, United Kingdom; cDepartment of Paediatrics, Faculty of Medicine, Siriaj Hospital, Mahidol University, Bangkok, 10700, Thailand; and dMicroarray Facility, The Wellcome Trust Sanger Institute, Cambridge, CB10 1SA, United Kingdom Edited by Mark T. Groudine, Fred Hutchinson Cancer Research Center, Seattle, WA, and approved October 14, 2009 (received for review August 17, 2009) It is well established that all of the cis-acting sequences required for 70 kb upstream the ␣-like genes (4). In addition, we have also shown fully regulated human ␣-globin expression are contained within a that a 120-kb region of conserved synteny containing the human region of Ϸ120 kb of conserved synteny. Here, we show that activa- ␣-like globin genes, together with their major upstream regulatory tion of this cluster in erythroid cells dramatically affects expression of element (MCS-R2, also called HS-40), is sufficient to obtain apparently unrelated and noncontiguous genes in the 500 kb sur- optimal tissue- and developmental stage-specific expression in a rounding this domain, including a gene (NME4) located 300 kb from mouse model (6). However, here we have asked whether globin the ␣-globin cluster. -

The NDPK/NME Superfamily: State of the Art Mathieu Boissan1,2, Uwe Schlattner3 and Marie-Lise Lacombe1
Laboratory Investigation (2018) 98, 164–174 © 2018 USCAP, Inc All rights reserved 0023-6837/18 EDITORIAL The NDPK/NME superfamily: state of the art Mathieu Boissan1,2, Uwe Schlattner3 and Marie-Lise Lacombe1 Nucleoside diphosphate kinases (NDPK) are nucleotide metabolism enzymes encoded by NME genes (also called NM23). Given the fact that not all NME-encoded proteins are catalytically active NDPKs and that NM23 generally refers to clinical studies on metastasis, we use here NME/NDPK to denote the proteins. Since their discovery in the 1950’s, NMEs/NDPKs have been shown to be involved in multiple physiological and pathological cellular processes, but the molecular mechanisms have not been fully determined. Recent progress in elucidating these underlying mechanisms has been presented by experts in the field at the 10th International Congress on the NDPK/NME/AWD protein family in October 2016 in Dubrovnik, Croatia, and is summarized in review articles or original research in this and an upcoming issue of Laboratory Investigation. Within this editorial, we discuss three major cellular processes that involve members of the multi-functional NME/NDPK family: (i) cancer and metastasis dissemination, (ii) membrane remodeling and nucleotide channeling, and iii) protein histidine phosphorylation. Laboratory Investigation (2018) 98, 164–174; doi:10.1038/labinvest.2017.137 ucleoside diphosphate kinases mitochondria (Lacombe et al, upcoming issue8). (NDPK) are nucleotide metabolism NME1, NME2, NME3, and NME4 form hexamers enzymes encoded by NME (also as the catalytically active form (Ćetković et al, next named NM23) genes. NDPKs are issue9). Group II is comprised of more divergent ubiquitous enzymes catalyzing the proteins sharing only 22 to 44% identity with Ntransfer of a phosphate from nucleoside group I enzymes and between each other. -

NME3 Regulates Mitochondria to Reduce ROS-Mediated Genome Instability
International Journal of Molecular Sciences Article NME3 Regulates Mitochondria to Reduce ROS-Mediated Genome Instability Chih-Wei Chen 1 , Ning Tsao 2, Wei Zhang 3 and Zee-Fen Chang 1,4,5,* 1 Institute of Molecular Medicine, College of Medicine, National Taiwan University, Taipei 10002, Taiwan; [email protected] 2 Institute of Biochemistry and Molecular Biology, College of Medicine, National Taiwan University, Taipei 10002, Taiwan; [email protected] 3 State Key Laboratory of Quality Research in Chinese Medicines, Macau University of Science and Technology, Taipa, Macau 999078, China; [email protected] 4 Center of Precision Medicine, College of Medicine, National Taiwan University, Taipei 10002, Taiwan 5 Genomics Research Center, Academia Sinica, Taipei 11529, Taiwan * Correspondence: [email protected]; Tel.: +886-2-23123456 (ext. 88590); Fax: +886-2-2826-0919 Received: 27 April 2020; Accepted: 15 July 2020; Published: 17 July 2020 Abstract: NME3 is a member of the nucleoside diphosphate kinase (NDPK) family that binds to the mitochondrial outer membrane to stimulate mitochondrial fusion. In this study, we showed that NME3 knockdown delayed DNA repair without reducing the cellular levels of nucleotide triphosphates. Further analyses revealed that NME3 knockdown increased fragmentation of mitochondria, which in turn led to mitochondrial oxidative stress-mediated DNA single-strand breaks (SSBs) in nuclear DNA. Re-expression of wild-type NME3 or inhibition of mitochondrial fission markedly reduced SSBs and facilitated DNA repair in NME3 knockdown cells, while expression of N-terminal deleted mutant defective in mitochondrial binding had no rescue effect. We further showed that disruption of mitochondrial fusion by knockdown of NME4 or MFN1 also caused mitochondrial oxidative stress-mediated genome instability. -

Two Separate Functions of NME3 Critical for Cell Survival Underlie a Neurodegenerative Disorder
Two separate functions of NME3 critical for cell survival underlie a neurodegenerative disorder Chih-Wei Chena, Hong-Ling Wanga, Ching-Wen Huangb, Chang-Yu Huangc, Wai Keong Lima, I-Chen Tua, Atmaja Koorapatia, Sung-Tsang Hsiehd, Hung-Wei Kand, Shiou-Ru Tzenge, Jung-Chi Liaof, Weng Man Chongf, Inna Naroditzkyg, Dvora Kidronh,i, Ayelet Eranj, Yousif Nijimk, Ella Selak, Hagit Baris Feldmanl, Limor Kalfonm, Hadas Raveh-Barakm, Tzipora C. Falik-Zaccaim,n, Orly Elpelego, Hanna Mandelm,p,1, and Zee-Fen Changa,q,1 aInstitute of Molecular Medicine, College of Medicine, National Taiwan University, 10002 Taipei, Taiwan; bDepartment of Medicine, College of Medicine, National Taiwan University, 10002 Taipei, Taiwan; cInstitute of Biochemistry and Molecular Biology, National Yang-Ming University, 11221 Taipei, Taiwan; dInstitute of Anatomy and Cell Biology, College of Medicine, National Taiwan University, 10002 Taipei, Taiwan; eInstitute of Biochemistry and Molecular Biology, College of Medicine, National Taiwan University, 10002 Taipei, Taiwan; fInstitute of Atomic and Molecular Sciences, Academia Sinica, 10617 Taipei, Taiwan; gDepartment of Pathology, Rambam Health Care Campus, 31096 Haifa, Israel; hDepartment of Pathology, Meir Hospital, 44100 Kfar Saba, Israel; iSackler School of Medicine, Tel Aviv University, 69978 Tel Aviv, Israel; jDepartment of Radiology, Rambam Health Care Campus, 31096 Haifa, Israel; kPediatric and Neonatal Unit, Nazareth Hospital EMMS, 17639 Nazareth, Israel; lThe Genetics Institute, Rambam Health Care Campus, 31096 Haifa, Israel; mInstitute of Human Genetics, Galilee Medical Center, 22100 Nahariya, Israel; nThe Azrieli Faculty of Medicine, Bar Ilan University, 13100 Safed, Israel; oDepartment of Genetic and Metabolic Diseases, Hadassah Hebrew University Medical Center, 91120 Jerusalem, Israel; pMetabolic Unit, Technion Faculty of Medicine, Rambam Health Care Campus, 31096 Haifa, Israel; and qCenter of Precision Medicine, College of Medicine, National Taiwan University, 10002 Taipei, Taiwan Edited by Christopher K. -

Generation of Bivalent Chromatin Domains During Cell Fate Decisions
De Gobbi et al. Epigenetics & Chromatin 2011, 4:9 http://www.epigeneticsandchromatin.com/content/4/1/9 RESEARCH Open Access Generation of bivalent chromatin domains during cell fate decisions Marco De Gobbi1, David Garrick1, Magnus Lynch1, Douglas Vernimmen1, Jim R Hughes1, Nicolas Goardon1, Sidinh Luc2, Karen M Lower1, Jacqueline A Sloane-Stanley1, Cristina Pina1, Shamit Soneji1, Raffaele Renella1, Tariq Enver1, Stephen Taylor3, Sten Eirik W Jacobsen2, Paresh Vyas1,4, Richard J Gibbons1 and Douglas R Higgs1* Abstract Background: In self-renewing, pluripotent cells, bivalent chromatin modification is thought to silence (H3K27me3) lineage control genes while ‘poising’ (H3K4me3) them for subsequent activation during differentiation, implying an important role for epigenetic modification in directing cell fate decisions. However, rather than representing an equivalently balanced epigenetic mark, the patterns and levels of histone modifications at bivalent genes can vary widely and the criteria for identifying this chromatin signature are poorly defined. Results: Here, we initially show how chromatin status alters during lineage commitment and differentiation at a single well characterised bivalent locus. In addition we have determined how chromatin modifications at this locus change with gene expression in both ensemble and single cell analyses. We also show, on a global scale, how mRNA expression may be reflected in the ratio of H3K4me3/H3K27me3. Conclusions: While truly ‘poised’ bivalently modified genes may exist, the original hypothesis that all bivalent genes are epigenetically premarked for subsequent expression might be oversimplistic. In fact, from the data presented in the present work, it is equally possible that many genes that appear to be bivalent in pluripotent and multipotent cells may simply be stochastically expressed at low levels in the process of multilineage priming. -
NME/NM23/NDPK and Histidine Phosphorylation
International Journal of Molecular Sciences Review NME/NM23/NDPK and Histidine Phosphorylation Kevin Adam y , Jia Ning y, Jeffrey Reina y and Tony Hunter * Molecular and Cell Biology Laboratory, Salk Institute for Biological Studies, La Jolla, CA 92037, USA; [email protected] (K.A.); [email protected] (J.N.); [email protected] (J.R.) * Correspondence: [email protected] These authors contributed equally to this work. y Received: 17 June 2020; Accepted: 7 August 2020; Published: 14 August 2020 Abstract: The NME (Non-metastatic) family members, also known as NDPKs (nucleoside diphosphate kinases), were originally identified and studied for their nucleoside diphosphate kinase activities. This family of kinases is extremely well conserved through evolution, being found in prokaryotes and eukaryotes, but also diverges enough to create a range of complexity, with homologous members having distinct functions in cells. In addition to nucleoside diphosphate kinase activity, some family members are reported to possess protein-histidine kinase activity, which, because of the lability of phosphohistidine, has been difficult to study due to the experimental challenges and lack of molecular tools. However, over the past few years, new methods to investigate this unstable modification and histidine kinase activity have been reported and scientific interest in this area is growing rapidly. This review presents a global overview of our current knowledge of the NME family and histidine phosphorylation, highlighting the underappreciated protein-histidine kinase activity of NME family members, specifically in human cells. In parallel, information about the structural and functional aspects of the NME family, and the knowns and unknowns of histidine kinase involvement in cell signaling are summarized.