Feeding Intensity and Molecular Prey Identification of The
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
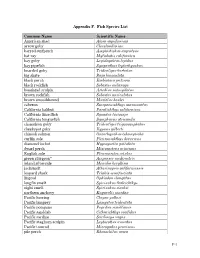
Appendix E: Fish Species List
Appendix F. Fish Species List Common Name Scientific Name American shad Alosa sapidissima arrow goby Clevelandia ios barred surfperch Amphistichus argenteus bat ray Myliobatis californica bay goby Lepidogobius lepidus bay pipefish Syngnathus leptorhynchus bearded goby Tridentiger barbatus big skate Raja binoculata black perch Embiotoca jacksoni black rockfish Sebastes melanops bonehead sculpin Artedius notospilotus brown rockfish Sebastes auriculatus brown smoothhound Mustelus henlei cabezon Scorpaenichthys marmoratus California halibut Paralichthys californicus California lizardfish Synodus lucioceps California tonguefish Symphurus atricauda chameleon goby Tridentiger trigonocephalus cheekspot goby Ilypnus gilberti chinook salmon Oncorhynchus tshawytscha curlfin sole Pleuronichthys decurrens diamond turbot Hypsopsetta guttulata dwarf perch Micrometrus minimus English sole Pleuronectes vetulus green sturgeon* Acipenser medirostris inland silverside Menidia beryllina jacksmelt Atherinopsis californiensis leopard shark Triakis semifasciata lingcod Ophiodon elongatus longfin smelt Spirinchus thaleichthys night smelt Spirinchus starksi northern anchovy Engraulis mordax Pacific herring Clupea pallasi Pacific lamprey Lampetra tridentata Pacific pompano Peprilus simillimus Pacific sanddab Citharichthys sordidus Pacific sardine Sardinops sagax Pacific staghorn sculpin Leptocottus armatus Pacific tomcod Microgadus proximus pile perch Rhacochilus vacca F-1 plainfin midshipman Porichthys notatus rainwater killifish Lucania parva river lamprey Lampetra -

Genetic Identification of Octopodidae Species in Southern California Seafood Markets: Species Diversity and Resource Implications
Genetic Identification of Octopodidae Species in Southern California Seafood Markets: Species Diversity and Resource Implications Chase Martin Center for Marine Biodiversity and Conservation Scripps Institution of Oceanography University of California San Diego Abstract Various species of Octopodidae are commonly found in seafood markets throughout Southern California. Most of the octopus available for purchase is imported, with the majority of imports coming from various Asian nations. Despite the diversity of global octopus species, products are most commonly labeled as simply “octopus,” with some distinctions being made in size, e.g., “baby” or “little octopus.” In efforts to characterize species diversity, this study genetically tested 59 octopus samples from a variety of seafood markets in Los Angeles, Orange, and San Diego Counties. Universal 16S rRNA primers (ref) and CO1 primers developed by Folmer et al. (1994) were used for PCR amplification and sequencing of mtDNA. In all, 105 sequences were acquired. Seven species were identified with some confidence. Amphioctopus aegina was the most prevalent species, while two additional species were undetermined. Little available data exists pertaining to octopus fisheries of the countries of production of the samples. Most available information on octopus fisheries pertains to those of Mediterranean and North African nations, and identifies the Octopus vulgaris as the fished species. Characterizing octopus diversity in Southern California seafood markets and assessing labeling and countries of production provides the necessary first step for assessing the possible management implications of these fisheries and seafood supply chain logistics for this group of cephalopods. Introduction Octopuses are exclusively marine cephalopod mollusks that form the order Octopoda. -

Pharaoh Cuttlefish, Sepia Pharaonis, Genome Reveals Unique Reflectin
fmars-08-639670 February 9, 2021 Time: 18:18 # 1 ORIGINAL RESEARCH published: 15 February 2021 doi: 10.3389/fmars.2021.639670 Pharaoh Cuttlefish, Sepia pharaonis, Genome Reveals Unique Reflectin Camouflage Gene Set Weiwei Song1,2, Ronghua Li1,2,3, Yun Zhao1,2, Herve Migaud1,2,3, Chunlin Wang1,2* and Michaël Bekaert3* 1 Key Laboratory of Applied Marine Biotechnology, Ministry of Education, Ningbo University, Ningbo, China, 2 Collaborative Innovation Centre for Zhejiang Marine High-Efficiency and Healthy Aquaculture, Ningbo University, Ningbo, China, 3 Institute of Aquaculture, Faculty of Natural Sciences, University of Stirling, Stirling, United Kingdom Sepia pharaonis, the pharaoh cuttlefish, is a commercially valuable cuttlefish species across the southeast coast of China and an important marine resource for the world fisheries. Research efforts to develop linkage mapping, or marker-assisted selection have been hampered by the absence of a high-quality reference genome. To address this need, we produced a hybrid reference genome of S. pharaonis using a long-read Edited by: platform (Oxford Nanopore Technologies PromethION) to assemble the genome and Andrew Stanley Mount, short-read, high quality technology (Illumina HiSeq X Ten) to correct for sequencing Clemson University, United States errors. The genome was assembled into 5,642 scaffolds with a total length of 4.79 Gb Reviewed by: and a scaffold N of 1.93 Mb. Annotation of the S. pharaonis genome assembly Simo Njabulo Maduna, 50 Norwegian Institute of Bioeconomy identified a total of 51,541 genes, including 12 copies of the reflectin gene, that enable Research (NIBIO), Norway cuttlefish to control their body coloration. -

Cephalopoda: Octopodidae): the Smallest Southwestern Atlantic Octopod, Found in Sea Debris
A new species of pygmy Paroctopus (Cephalopoda: Octopodidae): the smallest southwestern Atlantic octopod, found in sea debris Tatiana S. Leite ( [email protected] ) Universidade Federal de Santa Catarina Centro de Ciencias Biologicas https://orcid.org/0000-0001-9117-9648 Erica A.G. Vidal Universidade Federal do Parana Setor de Ciencias da Terra Françoise Dantas Lima Universidade Federal do Rio Grande do Norte Centro de Biociencias Sergio M.Q. Lima Universidade Federal do Rio Grande do Norte Centro de Biociencias Ricardo M Dias Universidade Federal do Sul da Bahia Giulia A. Giuberti Universidade Federal do Estado do Rio de Janeiro Davi De Vasconcellos Universidade Federal do Rio Grande Jennifer A. Mather University of Lethbridge Manuel Haimovici Universidade Federal do Rio Grande Original Paper Keywords: Paroctopus, octopus Posted Date: January 29th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-172910/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Version of Record: A version of this preprint was published at Marine Biodiversity on July 27th, 2021. See the published version at https://doi.org/10.1007/s12526-021-01201-z. Loading [MathJax]/jax/output/CommonHTML/fonts/TeX/fontdata.js Page 1/27 Abstract The new species, Paroctopus cthulu sp. nov. Leite, Haimovici, Lima and Lima, was recorded from very shallow coastal waters on sandy/muddy and shelter- poor bottoms with natural and human-origin debris. It is a small octopus, adults are less than 35 mm mantle length (ML) and weigh around 15 g. It has short to medium sized arms, enlarged suckers on the arms of both males and females, large posterior salivary glands (25 %ML), a relatively large beak (9 % ML) and medium to large mature eggs (3.5 to > 9 mm). -

Report on the Monitoring of Radionuclides in Fishery Products (March 2011 - January 2015)
Report on the Monitoring of Radionuclides in Fishery Products (March 2011 - January 2015) April 2015 Fisheries Agency of Japan 0 1 Table of Contents Overview…………………………………………………………………………………………………. 8 The Purpose of this Report………………………………………………………………………………9 Part One. Efforts to Guarantee the Safety of Fishery Products………………………………………..11 Chapter 1. Monitoring of Radioactive Materials in Food; Restrictions on Distribution and Other Countermeasures………...…………………………………………………………………11 1-1-1 Standard Limits for Radioactive Materials in Food………………………………………...……11 1-1-2 Methods of Testing for Radioactive Materials………………………………………...…………12 1-1-3 Inspections of Fishery Products for Radioactive Materials…………………………...…………14 1-1-4 Restrictions and Suspensions on Distribution and Shipping ……………………………………..18 1-1-5 Cancellation of Restrictions on Shipping and Distribution………………………………………20 Box 1 Calculation of the Limits for Human Consumption……..………………………………………23 Box 2 Survey of Radiation Dose from Radionuclides in Foods Calculation of the Limits…………….24 Box 3 Examples of Local Government Monitoring Plan………………………………...…………….25 Chapter 2. Results of Radioactive Cesium Inspections for Fishery Products…………………………26 1-2-1 Inspection Results for Nationwide Fishery Products in Japan (in total)…………………………26 1-2-2 Inspection Results for Fukushima Prefecture Fishery Products (all)…………………………….27 1-2-3 Inspection Results for Fishery Products (all) from Outside Fukushima Prefecture……………...30 1-2-4 Trends within Fish Species……………………………………………………………………….32 1-2-5 Inspection Results for Main Target Fish Species of Fishing and Farming by Fiscal Year……….42 1-2-6 Radioactive Material Concentrations within Fish within 20 km of the Fukushima Daiichi NPS.46 Box 4 Fukushima Fishing Trials………………………………...……………………………………...47 1-2-7 Screening Test by Prefectural and Municipal Governments……………………………………..48 Chapter 3. Inspection for Radionuclides Other Than Radioactive Cesium……………………………49 1-3-1 Inspections for Radioactive Strontium etc. -

Fishing Performance of an Octopus Minor Net Pot Made of Biodegradable Twines
www.trjfas.org ISSN 1303-2712 Turkish Journal of Fisheries and Aquatic Sciences 14: 21-30 (2014) DOI: 10.4194/1303-2712-v14_1_03 Fishing Performance of an Octopus minor Net Pot Made of Biodegradable Twines 1,* 1 1 Seonghun Kim , Seongwook Park , Kyounghoon Lee 1 National Fisheries Research and Development Institute, Fisheries Engineering Division, 216 Gijanghaean-ro Gijang-gun Gijang-eup Busan, Republic of Korea, 619-705. * Corresponding Author: Tel.: +82.51 7202584; Fax: +82.51 7202586; Received 17 April 2013 E-mail: [email protected] Accepted 17 December 2013 Abstract Gillnets and net pots are made of synthetic fiber as polyester (PE) and polyamide (PA). These are often lost by heavy weather or trawling of the active fishing gears. Lost gears result in the ghost fishing because these are non-degradable in seawater and damage to spawning grounds or habitats. To address these problems, biodegradable nets composed of aliphatic polyester were developed. This study describes four types of biodegradable net pots for capturing Octopus minor in Southern Korea, which is an area associated with high rates of net pot loss. The fishing performance of the biodegradable pots was compared to that of commercial net pots. The net pot with a synthetic body and biodegradable funnel (PE/Bio) produced approximately 50% of the catch collected using a commercial net pot (PE/PA). Conversely, net pots with a biodegradable body and a synthetic funnel (Bio/PA) produced the same amount of catch as the commercial net pot. The completely biodegradable net pot (Bio/Bio) caught 60% of that using the commercial net pot. -

A Dissertation Entitled Evolution, Systematics
A Dissertation Entitled Evolution, systematics, and phylogeography of Ponto-Caspian gobies (Benthophilinae: Gobiidae: Teleostei) By Matthew E. Neilson Submitted as partial fulfillment of the requirements for The Doctor of Philosophy Degree in Biology (Ecology) ____________________________________ Adviser: Dr. Carol A. Stepien ____________________________________ Committee Member: Dr. Christine M. Mayer ____________________________________ Committee Member: Dr. Elliot J. Tramer ____________________________________ Committee Member: Dr. David J. Jude ____________________________________ Committee Member: Dr. Juan L. Bouzat ____________________________________ College of Graduate Studies The University of Toledo December 2009 Copyright © 2009 This document is copyrighted material. Under copyright law, no parts of this document may be reproduced without the expressed permission of the author. _______________________________________________________________________ An Abstract of Evolution, systematics, and phylogeography of Ponto-Caspian gobies (Benthophilinae: Gobiidae: Teleostei) Matthew E. Neilson Submitted as partial fulfillment of the requirements for The Doctor of Philosophy Degree in Biology (Ecology) The University of Toledo December 2009 The study of biodiversity, at multiple hierarchical levels, provides insight into the evolutionary history of taxa and provides a framework for understanding patterns in ecology. This is especially poignant in invasion biology, where the prevalence of invasiveness in certain taxonomic groups could -

Appendix A. Species List and Threatened Or Endangered Species
Appendix A Great egrets rely on wetlands for feeding and nesting. © Mark Wilson Species Lists and Threatened or Endangered Species ■ Bird Species of the Complex ■ Mammal Species of the Complex ■ Reptile and Amphibian Species of the Complex ■ Fish Species of the Complex ■ Butterfly Species of the Complex ■ Threatened or Endanged Species Appendix A Bird Species of the Complex Conscience Lido Oyster Target Bird Species Amagansett Morton Sayville Seatuck Wertheim Point Beach Bay Rock s=Spring (Mar–May) S=Summer (Jun–Aug) A=Autumn (Sep–Nov) W=Winter (Dec–Feb) *=Birds documented breeding at the Complex Red-Throated Loon s AW s AW s AW s AW s AW s AW s AW s AW Gavia stellata Common Loon (Sc) s AW s AW s AW sSAW s AW s AW s AW sSAW Gavia immer Horned Grebe s AW s AW s AW s AW s AW s AW s AW s AW Podiceps auritus Red Necked Grebe s AW s AW Podiceps grisegena Eared Grebe s AW Podiceps nigricollis Pied-billed Grebe*(St) s AW s AW s AW s AW sSAW sSAW sSAW* Podilymbus podiceps Great Cormorant s AW s AW s AW s AW s AW s AW s AW s AW Phalacrocorax carbo Double-crested Cormorant sSAW sSAW sSAW sSAW sSAW sSAW sSAW sSAW Phalacrocorax auritus Brown Pelican S S Pelecanus occidentalis Northern Gannet s AW s AW s AW s AW s AW Morus bassanus Brown Booby S Sula leucogaster American Bittern* (Sc) s AW s AW s AW s AW sSAW* s AW sSAW* Botaurus lentiginosus Least Bittern*(St) sSA* sSAW* Ixobrychus exilis Great Blue Heron s AW sSAW sSAW sSAW sSAW sSAW sSAW sSAW Ardea herodias Great Egret sSA sSA sSAW sSA sSA sSA sSA sSAW Casmerodius albus Snowy Egret sSA sSA sSA sSA -

Inland Fishes of California
Inland Fishes of California Revise d and Expanded PETER B. MO YL E Illustrations by Chris Ma ri van Dyck and Joe Tome ller i NIVERS ITyor ALfFORNJA PRESS Ikrkd cr I.", ..\ n~d e ' Lon don Universit }, 0 Ca lifornia Press Herkdey and Los Angeles, Ca lifornia Uni ve rsity of alifornia Press, Ltd. Lundun, England ~ 2002 by the Regents of the Unive rsi ty of Ca lifornia Library of Cungress ataloging-in -Publ ica tion Data j\·[oyk, Pen: r B. Inland fis hes of California / Peter B. Moyle ; illustrations by Chris Mari van D)'ck and Joe Tomell eri.- Rev. and expanded. p. cm. In lu de> bibl iographical refe rences (p. ). ISBN 0- 20-2.2754 -'1 (cl ot h: alk. papa) I. rreshw:ltcr lishes-Cali(ornia. I. Title. QL62S C2 M6H 2002 597 .17/i'097Q4-dc21 20010 27680 1\!UlIl.Ifaclu rcd in Canacla II 10 Q9 00 07 06 0 04 03 02 10 ' 1\ 7 b '; -\ 3 2 1 Th paper u!)ed in thi> public.ltiu(] 111l'd., the minimum requirements "fA SI / i': ISO Z39.4H-1992 (R 199;) ( Pmlllllh'/l e ofPa pcr) . e Special Thanks The illustrations for this book were made possible by gra nts from the following : California-Nevada Chapter, American Fi she ries Soc iety Western Di vision, Am erican Fi she ries Society California Department of Fish and Game Giles W. and El ise G. Mead Foundation We appreciate the generous funding support toward the publication of this book by the United Sta tes Environme ntal Protection Agency, Region IX, San Francisco Contents Pre(acc ix Salmon and Trout, Salmonidae 242 Ackl10 11'ledgl11 el1ts Xlll Silversides, Atherinopsidae 307 COlll'er,<iol1 ['actors xv -

Octopus Insularis</Italic> As a New Marine Model for Evolutionary
© 2019. Published by The Company of Biologists Ltd | Biology Open (2019) 8, bio046086. doi:10.1242/bio.046086 RESEARCH ARTICLE Octopus insularis as a new marine model for evolutionary developmental biology Ernesto Maldonado1,*, Emma Rangel-Huerta1,2, Roberto González-Gómez3,4, Gabriel Fajardo-Alvarado3,4 and Piedad S. Morillo-Velarde4,5,* ABSTRACT of aquatic animal eggs and embryos guarantees the observation of Octopuses are intriguing organisms that, together with squids and every developmental stage using microscopy and allows detailed cuttlefishes, form the extant coleoid cephalopods. This group includes experimental analysis from the first cell division through to the many species that can potentially be used as models in the fields of formation of embryonic germ layers and organogenesis (Boletzky biomedicine, developmental biology, evolution, neuroscience and et al., 2006). Finally, small embryos allow reasonable sample sizes even for robotics research. The purpose of this work is to first to be tested together using multi-well plates to provide multiple present a simple method for maintaining Octopus insularis embryos experimental replicates at the same time, making them cost- under a laboratory setup. Second, we show that these embryos are effective animal models (Hill et al., 2005). suitable for detailed analyses of specific traits that appear during Coleoid cephalopods (octopus, squid and cuttlefish) exhibit the developmental stages, including the eyes, hearts, arms, suckers, largest nervous systems found among invertebrates (Young, 1971) chromatophores and Kölliker’s organs. Similar complex traits between and a sophisticated visual system controlling body color changes for cephalopods and vertebrates such as the visual, cardiovascular, communication, camouflage and mimicry (Hanlon et al., 2011; neural and pigmentation systems are generally considered to be a Robin et al., 2014). -

2013-2015 Cherry Point Final Report
Intertidal Biota Monitoring in the Cherry Point Aquatic Reserve 2013-2015 Monitoring Report Prepared for: Cherry Point Aquatic Reserve Citizen Stewardship Committee Prepared by: Michael Kyte Independent Marine Biologist and Wendy Steffensen and Eleanor Hines RE Sources for Sustainable Communities September 2016 Publication Information This Monitoring Report describes the research and monitoring study of intertidal biota conducted in the summers of 2013-2015 in the Cherry Point Aquatic Reserve. Copies of this Monitoring Report will be available at https://sites.google.com/a/re-sources.org/main- 2/programs/cleanwater/whatcom-and-skagit-county-aquatic-reserves. Author and Contact Information Wendy Steffensen North Sound Baykeeper, RE Sources for Sustainable Communities Eleanor Hines Lead Scientist, Clean Water Program RE Sources for Sustainable Communities 2309 Meridian Street Bellingham, WA 98225 [email protected] Michael Kyte Independent Marine Biologist [email protected] The report template was provided by Jerry Joyce for the Cherry Point and Fidalgo Bay Aquatic Reserves Citizen Stewardship Committees, and adapted here. Jerry Joyce Washington Environmental Council 1402 Third Avenue Seattle, WA 98101 206-440-8688 [email protected] i Acknowledgments Most of the sampling protocols and procedures are based on the work of the Island County/WSU Beach Watchers (currently known as the Sound Water Stewards). We thank them for the use of their materials and assistance. In particular, we thank Barbara Bennett, project coordinator for her assistance. We also thank our partners at WDNR and especially Betty Bookheim for her assistance in refining the procedures. We thank Dr. Megan Dethier of University of Washington for her assistance in helping us resolve some of the theoretical issues in the sampling protocol Surveys, data entry, quality control assistance and report writing were made possible by a vast array of interns and volunteers. -

Octopus Minor
https://doi.org/10.5607/en.2018.27.4.257 Exp Neurobiol. 2018 Aug;27(4):257-266. pISSN 1226-2560 • eISSN 2093-8144 Original Article A Brain Atlas of the Long Arm Octopus, Octopus minor Seung-Hyun Jung†, Ha Yeun Song†, Young Se Hyun, Yu-Cheol Kim, Ilson Whang, Tae-Young Choi and Seonmi Jo* Department of Genetic Resources Research, National Marine Biodiversity Institute of Korea (MABIK), Seocheon 33662, Korea Cephalopods have the most advanced nervous systems and intelligent behavior among all invertebrates. Their brains provide com- parative insights for understanding the molecular and functional origins of the human brain. Although brain maps that contain information on the organization of each subregion are necessary for a study on the brain, no whole brain atlas for adult cephalopods has been constructed to date. Here, we obtained sagittal and coronal sections covering the entire brain of adult Octopus minor (Sa- saki), which belongs to the genus with the most species in the class Cephalopoda and is commercially available in East Asia through- out the year. Sections were stained using Hematoxylin and Eosin (H&E) to visualize the cellular nuclei and subregions. H&E images of the serial sections were obtained at 30~70-μm intervals for the sagittal plain and at 40~80-μm intervals for the coronal plain. Set- ting the midline point of the posterior end as the fiducial point, we also established the distance coordinates of each image. We found that the brain had the typical brain structure of the Octopodiformes. A number of subregions were discriminated by a Hematoxylin- positive layer, the thickness and neuronal distribution pattern of which varied markedly depending upon the region.