Ab Initio Protein Structure Prediction Algorithms
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

An Effective Computational Method Incorporating Multiple Secondary
Old Dominion University ODU Digital Commons Computer Science Faculty Publications Computer Science 2017 An Effective Computational Method Incorporating Multiple Secondary Structure Predictions in Topology Determination for Cryo-EM Images Abhishek Biswas Old Dominion University Desh Ranjan Old Dominion University Mohammad Zubair Old Dominion University Stephanie Zeil Old Dominion University Kamal Al Nasr See next page for additional authors Follow this and additional works at: https://digitalcommons.odu.edu/computerscience_fac_pubs Part of the Biochemistry Commons, Computer Sciences Commons, Molecular Biology Commons, and the Statistics and Probability Commons Repository Citation Biswas, Abhishek; Ranjan, Desh; Zubair, Mohammad; Zeil, Stephanie; Al Nasr, Kamal; and He, Jing, "An Effective Computational Method Incorporating Multiple Secondary Structure Predictions in Topology Determination for Cryo-EM Images" (2017). Computer Science Faculty Publications. 81. https://digitalcommons.odu.edu/computerscience_fac_pubs/81 Original Publication Citation Biswas, A., Ranjan, D., Zubair, M., Zeil, S., Al Nasr, K., & He, J. (2017). An effective computational method incorporating multiple secondary structure predictions in topology determination for cryo-em images. IEEE/ACM Transactions on Computational Biology and Bioinformatics, 14(3), 578-586. doi:10.1109/tcbb.2016.2543721 This Article is brought to you for free and open access by the Computer Science at ODU Digital Commons. It has been accepted for inclusion in Computer Science Faculty Publications by an authorized administrator of ODU Digital Commons. For more information, please contact [email protected]. Authors Abhishek Biswas, Desh Ranjan, Mohammad Zubair, Stephanie Zeil, Kamal Al Nasr, and Jing He This article is available at ODU Digital Commons: https://digitalcommons.odu.edu/computerscience_fac_pubs/81 HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author IEEE/ACM Manuscript Author Trans Comput Manuscript Author Biol Bioinform. -

Protein Structure & Folding
6 Protein Structure & Folding To understand protein folding In the last chapter we learned that proteins are composed of amino acids Goal as a chemical equilibrium. linked together by peptide bonds. We also learned that the twenty amino acids display a wide range of chemical properties. In this chapter we will see Objectives that how a protein folds is determined by its amino acid sequence and that the three-dimensional shape of a folded protein determines its function by After this chapter, you should be able to: the way it positions these amino acids. Finally, we will see that proteins fold • describe the four levels of protein because doing so minimizes Gibbs free energy and that this minimization structure and the thermodynamic involves both making the most favorable bonds and maximizing disorder. forces that stabilize them. • explain how entropy (S) and enthalpy Proteins exhibit four levels of structure (H) contribute to Gibbs free energy. • use the equation ΔG = ΔH – TΔS The structure of proteins can be broken down into four levels of to determine the dependence of organization. The first is primary structure, the linear sequence of amino the favorability of a reaction on acids in the polypeptide chain. By convention, the primary sequence is temperature. written in order from the amino acid at the N-terminus (by convention • explain the hydrophobic effect and its usually on the left) to the amino acid at the C-terminus. The second level role in protein folding. of protein structure, secondary structure, is the local conformation adopted by stretches of contiguous amino acids. -

DNA Glycosylase Exercise - Levels 1 & 2: Answer Key
Name________________________ StarBiochem DNA Glycosylase Exercise - Levels 1 & 2: Answer Key Background In this exercise, you will explore the structure of a DNA repair protein found in most species, including bacteria. DNA repair proteins move along DNA strands, checking for mistakes or damage. DNA glycosylases, a specific type of DNA repair protein, recognize DNA bases that have been chemically altered and remove them, leaving a site in the DNA without a base. Other proteins then come along to fill in the missing DNA base. Learning objectives We will explore the relationship between a protein’s structure and its function in a human DNA glycosylase called human 8-oxoguanine glycosylase (hOGG1). Getting started We will begin this exercise by exploring the structure of hOGG1 using a molecular 3-D viewer called StarBiochem. In this particular structure, the repair protein is bound to a segment of DNA that has been damaged. We will first focus on the structure hOGG1 and then on how this protein interacts with DNA to repair a damaged DNA base. • To begin using StarBiochem, please navigate to: http://mit.edu/star/biochem/. • Click on the Start button Click on the Start button for StarBiochem. • Click Trust when a prompt appears asking if you trust the certificate. • In the top menu, click on Samples à Select from Samples. Within the Amino Acid/Proteins à Protein tab, select “DNA glycosylase hOGG1 w/ DNA – H. sapiens (1EBM)”. “1EBM” is the four character unique ID for this structure. Take a moment to look at the structure from various angles by rotating and zooming on the structure. -

POLYMER STRUCTURE and CHARACTERIZATION Professor
POLYMER STRUCTURE AND CHARACTERIZATION Professor John A. Nairn Fall 2007 TABLE OF CONTENTS 1 INTRODUCTION 1 1.1 Definitions of Terms . 2 1.2 Course Goals . 5 2 POLYMER MOLECULAR WEIGHT 7 2.1 Introduction . 7 2.2 Number Average Molecular Weight . 9 2.3 Weight Average Molecular Weight . 10 2.4 Other Average Molecular Weights . 10 2.5 A Distribution of Molecular Weights . 11 2.6 Most Probable Molecular Weight Distribution . 12 3 MOLECULAR CONFORMATIONS 21 3.1 Introduction . 21 3.2 Nomenclature . 23 3.3 Property Calculation . 25 3.4 Freely-Jointed Chain . 27 3.4.1 Freely-Jointed Chain Analysis . 28 3.4.2 Comment on Freely-Jointed Chain . 34 3.5 Equivalent Freely Jointed Chain . 37 3.6 Vector Analysis of Polymer Conformations . 38 3.7 Freely-Rotating Chain . 41 3.8 Hindered Rotating Chain . 43 3.9 More Realistic Analysis . 45 3.10 Theta (Θ) Temperature . 47 3.11 Rotational Isomeric State Model . 48 4 RUBBER ELASTICITY 57 4.1 Introduction . 57 4.2 Historical Observations . 57 4.3 Thermodynamics . 60 4.4 Mechanical Properties . 62 4.5 Making Elastomers . 68 4.5.1 Diene Elastomers . 68 0 4.5.2 Nondiene Elastomers . 69 4.5.3 Thermoplastic Elastomers . 70 5 AMORPHOUS POLYMERS 73 5.1 Introduction . 73 5.2 The Glass Transition . 73 5.3 Free Volume Theory . 73 5.4 Physical Aging . 73 6 SEMICRYSTALLINE POLYMERS 75 6.1 Introduction . 75 6.2 Degree of Crystallization . 75 6.3 Structures . 75 Chapter 1 INTRODUCTION The topic of polymer structure and characterization covers molecular structure of polymer molecules, the arrangement of polymer molecules within a bulk polymer material, and techniques used to give information about structure or properties of polymers. -

Ideal Chain Conformations and Statistics
ENAS 606 : Polymer Physics Chinedum Osuji 01.24.2013:HO2 Ideal Chain Conformations and Statistics 1 Overview Consideration of the structure of macromolecules starts with a look at the details of chain level chemical details which can impact the conformations adopted by the polymer. In the case of saturated carbon ◦ backbones, while maintaining the desired Ci−1 − Ci − Ci+1 bond angle of 112 , the placement of the final carbon in the triad above can occur at any point along the circumference of a circle, defining a torsion angle '. We can readily recognize the energetic differences as a function this angle, U(') such that there are 3 minima - a deep minimum corresponding the the trans state, for which ' = 0 and energetically equivalent gauche− and gauche+ states at ' = ±120 degrees, as shown in Fig 1. Figure 1: Trans, gauche- and gauche+ configurations, and their energetic sates 1.1 Static Flexibility The static flexibility of the chain in equilibrium is determined by the difference between the levels of the energy minima corresponding the gauche and trans states, ∆. If ∆ < kT , the g+, g− and t states occur with similar probability, and so the chain can change direction and appears as a random coil. If ∆ takes on a larger value, then the t conformations will be enriched, so the chain will be rigid locally, but on larger length scales, the eventual occurrence of g+ and g− conformations imparts a random conformation. Overall, if we ignore details on some length scale smaller than lp, the persistence length, the polymer appears as a continuous flexible chain where 1 lp = l0 exp(∆/kT ) (1) where l0 is something like a monomer length. -

Proteasomes on the Chromosome Cell Research (2017) 27:602-603
602 Cell Research (2017) 27:602-603. © 2017 IBCB, SIBS, CAS All rights reserved 1001-0602/17 $ 32.00 RESEARCH HIGHLIGHT www.nature.com/cr Proteasomes on the chromosome Cell Research (2017) 27:602-603. doi:10.1038/cr.2017.28; published online 7 March 2017 Targeted proteolysis plays an this process, both in mediating homolog ligases, respectively, as RN components important role in the execution and association and in providing crossovers that impact crossover formation [5, 6]. regulation of many cellular events. that tether homologs and ensure their In the first of the two papers, Ahuja et Two recent papers in Science identify accurate segregation. Meiotic recom- al. [7] report that budding yeast pre9∆ novel roles for proteasome-mediated bination is initiated by DNA double- mutants, which lack a nonessential proteolysis in homologous chromo- strand breaks (DSBs) at many sites proteasome subunit, display defects in some pairing, recombination, and along chromosomes, and the multiple meiotic DSB repair, in chromosome segregation during meiosis. interhomolog interactions formed by pairing and synapsis, and in crossover Protein degradation by the 26S pro- DSB repair drive homolog association, formation. Similar defects are seen, teasome drives a variety of processes culminating in the end-to-end homolog but to a lesser extent, in cells treated central to the cell cycle, growth, and dif- synapsis by a protein structure called with the proteasome inhibitor MG132. ferentiation. Proteins are targeted to the the synaptonemal complex (SC) [3]. These defects all can be ascribed to proteasome by covalently ligated chains SC-associated focal protein complexes, a failure to remove nonhomologous of ubiquitin, a small (8.5 kDa) protein. -

Chapter 13 Protein Structure Learning Objectives
Chapter 13 Protein structure Learning objectives Upon completing this material you should be able to: ■ understand the principles of protein primary, secondary, tertiary, and quaternary structure; ■use the NCBI tool CN3D to view a protein structure; ■use the NCBI tool VAST to align two structures; ■explain the role of PDB including its purpose, contents, and tools; ■explain the role of structure annotation databases such as SCOP and CATH; and ■describe approaches to modeling the three-dimensional structure of proteins. Outline Overview of protein structure Principles of protein structure Protein Data Bank Protein structure prediction Intrinsically disordered proteins Protein structure and disease Overview: protein structure The three-dimensional structure of a protein determines its capacity to function. Christian Anfinsen and others denatured ribonuclease, observed rapid refolding, and demonstrated that the primary amino acid sequence determines its three-dimensional structure. We can study protein structure to understand problems such as the consequence of disease-causing mutations; the properties of ligand-binding sites; and the functions of homologs. Outline Overview of protein structure Principles of protein structure Protein Data Bank Protein structure prediction Intrinsically disordered proteins Protein structure and disease Protein primary and secondary structure Results from three secondary structure programs are shown, with their consensus. h: alpha helix; c: random coil; e: extended strand Protein tertiary and quaternary structure Quarternary structure: the four subunits of hemoglobin are shown (with an α 2β2 composition and one beta globin chain high- lighted) as well as four noncovalently attached heme groups. The peptide bond; phi and psi angles The peptide bond; phi and psi angles in DeepView Protein secondary structure Protein secondary structure is determined by the amino acid side chains. -

Native-Like Mean Structure in the Unfolded Ensemble of Small Proteins
B doi:10.1016/S0022-2836(02)00888-4 available online at http://www.idealibrary.com on w J. Mol. Biol. (2002) 323, 153–164 Native-like Mean Structure in the Unfolded Ensemble of Small Proteins Bojan Zagrovic1, Christopher D. Snow1, Siraj Khaliq2 Michael R. Shirts2 and Vijay S. Pande1,2* 1Biophysics Program The nature of the unfolded state plays a great role in our understanding of Stanford University, Stanford proteins. However, accurately studying the unfolded state with computer CA 94305-5080, USA simulation is difficult, due to its complexity and the great deal of sampling required. Using a supercluster of over 10,000 processors we 2Department of Chemistry have performed close to 800 ms of molecular dynamics simulation in Stanford University, Stanford atomistic detail of the folded and unfolded states of three polypeptides CA 94305-5080, USA from a range of structural classes: the all-alpha villin headpiece molecule, the beta hairpin tryptophan zipper, and a designed alpha-beta zinc finger mimic. A comparison between the folded and the unfolded ensembles reveals that, even though virtually none of the individual members of the unfolded ensemble exhibits native-like features, the mean unfolded structure (averaged over the entire unfolded ensemble) has a native-like geometry. This suggests several novel implications for protein folding and structure prediction as well as new interpretations for experiments which find structure in ensemble-averaged measurements. q 2002 Elsevier Science Ltd. All rights reserved Keywords: mean-structure hypothesis; unfolded state of proteins; *Corresponding author distributed computing; conformational averaging Introduction under folding conditions, with some notable exceptions.14 – 16 This is understandable since under Historically, the unfolded state of proteins has such conditions the unfolded state is an unstable, received significantly less attention than the folded fleeting species making any kind of quantitative state.1 The reasons for this are primarily its struc- experimental measurement very difficult. -

Increasing the Accuracy of Single Sequence Prediction Methods Using a Deep Semi-Supervised Learning Framework Lewis Moffat 1,∗ and David T
Structural Bioinformatics Downloaded from https://academic.oup.com/bioinformatics/advance-article/doi/10.1093/bioinformatics/btab491/6313164 by guest on 07 July 2021 Increasing the Accuracy of Single Sequence Prediction Methods Using a Deep Semi-Supervised Learning Framework Lewis Moffat 1,∗ and David T. Jones 1,∗ 1Department of Computer Science, University College London, Gower Street, London WC1E 6BT, UK; and Francis Crick Institute, 1 Midland Road, London NW1 1AT, UK ∗To whom correspondence should be addressed. Associate Editor: XXXXXXX Received on XXXXX; revised on XXXXX; accepted on XXXXX Abstract Motivation: Over the past 50 years, our ability to model protein sequences with evolutionary information has progressed in leaps and bounds. However, even with the latest deep learning methods, the modelling of a critically important class of proteins, single orphan sequences, remains unsolved. Results: By taking a bioinformatics approach to semi-supervised machine learning, we develop Profile Augmentation of Single Sequences (PASS), a simple but powerful framework for building accurate single- sequence methods. To demonstrate the effectiveness of PASS we apply it to the mature field of secondary structure prediction. In doing so we develop S4PRED, the successor to the open-source PSIPRED-Single method, which achieves an unprecedented Q3 score of 75.3% on the standard CB513 test. PASS provides a blueprint for the development of a new generation of predictive methods, advancing our ability to model individual protein sequences. Availability: The S4PRED model is available as open source software on the PSIPRED GitHub repository (https://github.com/psipred/s4pred), along with documentation. It will also be provided as a part of the PSIPRED web service (http://bioinf.cs.ucl.ac.uk/psipred/) Contact: [email protected], [email protected] Supplementary information: Supplementary data are available at Bioinformatics online. -

Introduction to Proteins and Amino Acids Introduction
Introduction to Proteins and Amino Acids Introduction • Twenty percent of the human body is made up of proteins. Proteins are the large, complex molecules that are critical for normal functioning of cells. • They are essential for the structure, function, and regulation of the body’s tissues and organs. • Proteins are made up of smaller units called amino acids, which are building blocks of proteins. They are attached to one another by peptide bonds forming a long chain of proteins. Amino acid structure and its classification • An amino acid contains both a carboxylic group and an amino group. Amino acids that have an amino group bonded directly to the alpha-carbon are referred to as alpha amino acids. • Every alpha amino acid has a carbon atom, called an alpha carbon, Cα ; bonded to a carboxylic acid, –COOH group; an amino, –NH2 group; a hydrogen atom; and an R group that is unique for every amino acid. Classification of amino acids • There are 20 amino acids. Based on the nature of their ‘R’ group, they are classified based on their polarity as: Classification based on essentiality: Essential amino acids are the amino acids which you need through your diet because your body cannot make them. Whereas non essential amino acids are the amino acids which are not an essential part of your diet because they can be synthesized by your body. Essential Non essential Histidine Alanine Isoleucine Arginine Leucine Aspargine Methionine Aspartate Phenyl alanine Cystine Threonine Glutamic acid Tryptophan Glycine Valine Ornithine Proline Serine Tyrosine Peptide bonds • Amino acids are linked together by ‘amide groups’ called peptide bonds. -
CD of Proteins Sources Include
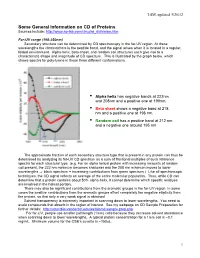
LSM, updated 3/26/12 Some General Information on CD of Proteins Sources include: http://www.ap-lab.com/circular_dichroism.htm Far-UV range (190-250nm) Secondary structure can be determined by CD spectroscopy in the far-UV region. At these wavelengths the chromophore is the peptide bond, and the signal arises when it is located in a regular, folded environment. Alpha-helix, beta-sheet, and random coil structures each give rise to a characteristic shape and magnitude of CD spectrum. This is illustrated by the graph below, which shows spectra for poly-lysine in these three different conformations. • Alpha helix has negative bands at 222nm and 208nm and a positive one at 190nm. • Beta sheet shows a negative band at 218 nm and a positive one at 196 nm. • Random coil has a positive band at 212 nm and a negative one around 195 nm. The approximate fraction of each secondary structure type that is present in any protein can thus be determined by analyzing its far-UV CD spectrum as a sum of fractional multiples of such reference spectra for each structural type. (e.g. For an alpha helical protein with increasing amounts of random coil present, the 222 nm minimum becomes shallower and the 208 nm minimum moves to lower wavelengths ⇒ black spectrum + increasing contributions from green spectrum.) Like all spectroscopic techniques, the CD signal reflects an average of the entire molecular population. Thus, while CD can determine that a protein contains about 50% alpha-helix, it cannot determine which specific residues are involved in the helical portion. -

Bi-Allelic Novel Variants in CLIC5 Identified in a Cameroonian
G C A T T A C G G C A T genes Article Bi-Allelic Novel Variants in CLIC5 Identified in a Cameroonian Multiplex Family with Non-Syndromic Hearing Impairment Edmond Wonkam-Tingang 1, Isabelle Schrauwen 2 , Kevin K. Esoh 1 , Thashi Bharadwaj 2, Liz M. Nouel-Saied 2 , Anushree Acharya 2, Abdul Nasir 3 , Samuel M. Adadey 1,4 , Shaheen Mowla 5 , Suzanne M. Leal 2 and Ambroise Wonkam 1,* 1 Division of Human Genetics, Faculty of Health Sciences, University of Cape Town, Cape Town 7925, South Africa; [email protected] (E.W.-T.); [email protected] (K.K.E.); [email protected] (S.M.A.) 2 Center for Statistical Genetics, Sergievsky Center, Taub Institute for Alzheimer’s Disease and the Aging Brain, and the Department of Neurology, Columbia University Medical Centre, New York, NY 10032, USA; [email protected] (I.S.); [email protected] (T.B.); [email protected] (L.M.N.-S.); [email protected] (A.A.); [email protected] (S.M.L.) 3 Synthetic Protein Engineering Lab (SPEL), Department of Molecular Science and Technology, Ajou University, Suwon 443-749, Korea; [email protected] 4 West African Centre for Cell Biology of Infectious Pathogens (WACCBIP), University of Ghana, Accra LG 54, Ghana 5 Division of Haematology, Department of Pathology, Faculty of Health Sciences, University of Cape Town, Cape Town 7925, South Africa; [email protected] * Correspondence: [email protected]; Tel.: +27-21-4066-307 Received: 28 September 2020; Accepted: 20 October 2020; Published: 23 October 2020 Abstract: DNA samples from five members of a multiplex non-consanguineous Cameroonian family, segregating prelingual and progressive autosomal recessive non-syndromic sensorineural hearing impairment, underwent whole exome sequencing.