Supplementary Table S2. Cpuorfs Extracted from D. Melanogaster , D
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Inherited Monogenic Defects of Ceramide Metabolism Molecular
Clinica Chimica Acta 495 (2019) 457–466 Contents lists available at ScienceDirect Clinica Chimica Acta journal homepage: www.elsevier.com/locate/cca Review Inherited monogenic defects of ceramide metabolism: Molecular bases and diagnoses T ⁎⁎ Patricia Dubota,b, Frédérique Sabourdya,b, Jitka Rybovac,Jeffrey A. Medinc,d, , ⁎ Thierry Levadea,b, a Laboratoire de Biochimie Métabolique, Centre de Référence en Maladies Héréditaires du Métabolisme, Institut Fédératif de Biologie, CHU de Toulouse, Toulouse, France b INSERM UMR1037, CRCT (Cancer Research Center of Toulouse), Université Paul Sabatier, Toulouse, France c Department of Pediatrics, Medical College of Wisconsin, Milwaukee, WI, USA d Department of Biochemistry, Medical College of Wisconsin, Milwaukee, WI, USA ABSTRACT Ceramides are membrane lipids implicated in the regulation of numerous biological functions. Recent evidence suggests that specific subsets of molecular species of ceramide may play distinct physiological roles. The importance of this family of molecules in vertebrates is witnessed by the deleterious consequences of genetic alterations in ceramide metabolism. This brief review summarizes the clinical presentation of human disorders due to the deficiency of enzymes involved either in the biosynthesis or the degradation of ceramides. Information on the possible underlying pathophysiological mechanisms is also provided, based on knowledge gathered from animal models of these inherited rare conditions. When appropriate, tools for chemical and molecular diagnosis of these disorders and therapeutic options are also presented. 1. Introduction/foreword relationships that are relevant for unraveling the biological role of these genes and gene products in humans. Studies on ceramides and sphingolipid metabolism have attracted a lot of attention recently. This is largely related to the multiplicity of 2. -

Lysophosphatidic Acid and Its Receptors: Pharmacology and Therapeutic Potential in Atherosclerosis and Vascular Disease
JPT-107404; No of Pages 13 Pharmacology & Therapeutics xxx (2019) xxx Contents lists available at ScienceDirect Pharmacology & Therapeutics journal homepage: www.elsevier.com/locate/pharmthera Lysophosphatidic acid and its receptors: pharmacology and therapeutic potential in atherosclerosis and vascular disease Ying Zhou a, Peter J. Little a,b, Hang T. Ta a,c, Suowen Xu d, Danielle Kamato a,b,⁎ a School of Pharmacy, University of Queensland, Pharmacy Australia Centre of Excellence, Woolloongabba, QLD 4102, Australia b Department of Pharmacy, Xinhua College of Sun Yat-sen University, Tianhe District, Guangzhou 510520, China c Australian Institute for Bioengineering and Nanotechnology, The University of Queensland, Brisbane, St Lucia, QLD 4072, Australia d Aab Cardiovascular Research Institute, Department of Medicine, University of Rochester School of Medicine and Dentistry, Rochester, NY 14642, USA article info abstract Available online xxxx Lysophosphatidic acid (LPA) is a collective name for a set of bioactive lipid species. Via six widely distributed G protein-coupled receptors (GPCRs), LPA elicits a plethora of biological responses, contributing to inflammation, Keywords: thrombosis and atherosclerosis. There have recently been considerable advances in GPCR signaling especially Lysophosphatidic acid recognition of the extended role for GPCR transactivation of tyrosine and serine/threonine kinase growth factor G-protein coupled receptors receptors. This review covers LPA signaling pathways in the light of new information. The use of transgenic and Atherosclerosis gene knockout animals, gene manipulated cells, pharmacological LPA receptor agonists and antagonists have Gproteins fi β-arrestins provided many insights into the biological signi cance of LPA and individual LPA receptors in the progression Transactivation of atherosclerosis and vascular diseases. -

Genome-Wide Prediction of Small Molecule Binding to Remote
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.04.236729; this version posted August 5, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Genome-wide Prediction of Small Molecule Binding 2 to Remote Orphan Proteins Using Distilled Sequence 3 Alignment Embedding 1 2 3 4 4 Tian Cai , Hansaim Lim , Kyra Alyssa Abbu , Yue Qiu , 5,6 1,2,3,4,7,* 5 Ruth Nussinov , and Lei Xie 1 6 Ph.D. Program in Computer Science, The Graduate Center, The City University of New York, New York, 10016, USA 2 7 Ph.D. Program in Biochemistry, The Graduate Center, The City University of New York, New York, 10016, USA 3 8 Department of Computer Science, Hunter College, The City University of New York, New York, 10065, USA 4 9 Ph.D. Program in Biology, The Graduate Center, The City University of New York, New York, 10016, USA 5 10 Computational Structural Biology Section, Basic Science Program, Frederick National Laboratory for Cancer Research, 11 Frederick, MD 21702, USA 6 12 Department of Human Molecular Genetics and Biochemistry, Sackler School of Medicine, Tel Aviv University, Tel 13 Aviv, Israel 7 14 Helen and Robert Appel Alzheimer’s Disease Research Institute, Feil Family Brain & Mind Research Institute, Weill 15 Cornell Medicine, Cornell University, New York, 10021, USA * 16 [email protected] 17 July 27, 2020 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.08.04.236729; this version posted August 5, 2020. -

Edinburgh Research Explorer
Edinburgh Research Explorer International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list Citation for published version: Davenport, AP, Alexander, SPH, Sharman, JL, Pawson, AJ, Benson, HE, Monaghan, AE, Liew, WC, Mpamhanga, CP, Bonner, TI, Neubig, RR, Pin, JP, Spedding, M & Harmar, AJ 2013, 'International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list: recommendations for new pairings with cognate ligands', Pharmacological reviews, vol. 65, no. 3, pp. 967-86. https://doi.org/10.1124/pr.112.007179 Digital Object Identifier (DOI): 10.1124/pr.112.007179 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: Pharmacological reviews Publisher Rights Statement: U.S. Government work not protected by U.S. copyright General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 02. Oct. 2021 1521-0081/65/3/967–986$25.00 http://dx.doi.org/10.1124/pr.112.007179 PHARMACOLOGICAL REVIEWS Pharmacol Rev 65:967–986, July 2013 U.S. -

Enhancing Skin Health: by Oral Administration of Natural Compounds and Minerals with Implications to the Dermal Microbiome
Review Enhancing Skin Health: By Oral Administration of Natural Compounds and Minerals with Implications to the Dermal Microbiome David L. Vollmer 1, Virginia A. West 1 and Edwin D. Lephart 2,* 1 4Life Research, Scientific Research Division, Sandy, Utah 84070, USA; [email protected] (D.L.V.); [email protected] (V.A.W) 2 Department of Physiology, Developmental Biology and The Neuroscience Center, Brigham Young University, Provo, Utah 84602, USA * Correspondence: [email protected]; Tel.: +1-801-422-2006 Received: 23 August 2018; Accepted: 1 October 2018; Published: 7 October 2018 Abstract: The history of cosmetics goes back to early Egyptian times for hygiene and health benefits while the history of topical applications that provide a medicinal treatment to combat dermal aging is relatively new. For example, the term cosmeceutical was first coined by Albert Kligman in 1984 to describe topical products that afford both cosmetic and therapeutic benefits. However, beauty comes from the inside. Therefore, for some time scientists have considered how nutrition reflects healthy skin and the aging process. The more recent link between nutrition and skin aging began in earnest around the year 2000 with the demonstrated increase in peer-reviewed scientific journal reports on this topic that included biochemical and molecular mechanisms of action. Thus, the application of: (a) topical administration from outside into the skin and (b) inside by oral consumption of nutritionals to the outer skin layers is now common place and many journal reports exhibit significant improvement for both on a variety of dermal parameters. Therefore, this review covers, where applicable, the history, chemical structure, and sources such as biological and biomedical properties in the skin along with animal and clinical data on the oral applications of: (a) collagen, (b) ceramide, (c) β-carotene, (d) astaxanthin, (e) coenzyme Q10, (f) colostrum, (g) zinc, and (h) selenium in their mode of action or function in improving dermal health by various quantified endpoints. -
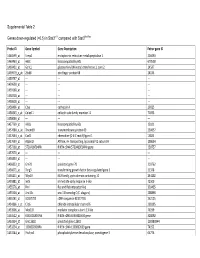
0.5) in Stat3∆/∆ Compared with Stat3flox/Flox
Supplemental Table 2 Genes down-regulated (<0.5) in Stat3∆/∆ compared with Stat3flox/flox Probe ID Gene Symbol Gene Description Entrez gene ID 1460599_at Ermp1 endoplasmic reticulum metallopeptidase 1 226090 1460463_at H60c histocompatibility 60c 670558 1460431_at Gcnt1 glucosaminyl (N-acetyl) transferase 1, core 2 14537 1459979_x_at Zfp68 zinc finger protein 68 24135 1459747_at --- --- --- 1459608_at --- --- --- 1459168_at --- --- --- 1458718_at --- --- --- 1458618_at --- --- --- 1458466_at Ctsa cathepsin A 19025 1458345_s_at Colec11 collectin sub-family member 11 71693 1458046_at --- --- --- 1457769_at H60a histocompatibility 60a 15101 1457680_a_at Tmem69 transmembrane protein 69 230657 1457644_s_at Cxcl1 chemokine (C-X-C motif) ligand 1 14825 1457639_at Atp6v1h ATPase, H+ transporting, lysosomal V1 subunit H 108664 1457260_at 5730409E04Rik RIKEN cDNA 5730409E04Rik gene 230757 1457070_at --- --- --- 1456893_at --- --- --- 1456823_at Gm70 predicted gene 70 210762 1456671_at Tbrg3 transforming growth factor beta regulated gene 3 21378 1456211_at Nlrp10 NLR family, pyrin domain containing 10 244202 1455881_at Ier5l immediate early response 5-like 72500 1455576_at Rinl Ras and Rab interactor-like 320435 1455304_at Unc13c unc-13 homolog C (C. elegans) 208898 1455241_at BC037703 cDNA sequence BC037703 242125 1454866_s_at Clic6 chloride intracellular channel 6 209195 1453906_at Med13l mediator complex subunit 13-like 76199 1453522_at 6530401N04Rik RIKEN cDNA 6530401N04 gene 328092 1453354_at Gm11602 predicted gene 11602 100380944 1453234_at -
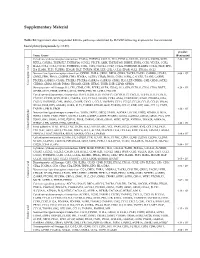
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R, -

The Effect of Hypoxia on the Expression of CXC Chemokines and CXC Chemokine Receptors—A Review of Literature
International Journal of Molecular Sciences Review The Effect of Hypoxia on the Expression of CXC Chemokines and CXC Chemokine Receptors—A Review of Literature Jan Korbecki 1 , Klaudyna Kojder 2, Patrycja Kapczuk 1, Patrycja Kupnicka 1 , Barbara Gawro ´nska-Szklarz 3 , Izabela Gutowska 4 , Dariusz Chlubek 1 and Irena Baranowska-Bosiacka 1,* 1 Department of Biochemistry and Medical Chemistry, Pomeranian Medical University in Szczecin, Powsta´nców Wielkopolskich 72 Av., 70-111 Szczecin, Poland; [email protected] (J.K.); [email protected] (P.K.); [email protected] (P.K.); [email protected] (D.C.) 2 Department of Anaesthesiology and Intensive Care, Pomeranian Medical University in Szczecin, Unii Lubelskiej 1, 71-281 Szczecin, Poland; [email protected] 3 Department of Pharmacokinetics and Therapeutic Drug Monitoring, Pomeranian Medical University in Szczecin, Powsta´nców Wielkopolskich 72 Av., 70-111 Szczecin, Poland; [email protected] 4 Department of Medical Chemistry, Pomeranian Medical University in Szczecin, Powsta´nców Wlkp. 72 Av., 70-111 Szczecin, Poland; [email protected] * Correspondence: [email protected]; Tel.: +48-914661515 Abstract: Hypoxia is an integral component of the tumor microenvironment. Either as chronic or cycling hypoxia, it exerts a similar effect on cancer processes by activating hypoxia-inducible factor-1 (HIF-1) and nuclear factor (NF-κB), with cycling hypoxia showing a stronger proinflammatory influ- ence. One of the systems affected by hypoxia is the CXC chemokine system. This paper reviews all available information on hypoxia-induced changes in the expression of all CXC chemokines (CXCL1, CXCL2, CXCL3, CXCL4, CXCL5, CXCL6, CXCL7, CXCL8 (IL-8), CXCL9, CXCL10, CXCL11, CXCL12 Citation: Korbecki, J.; Kojder, K.; Kapczuk, P.; Kupnicka, P.; (SDF-1), CXCL13, CXCL14, CXCL15, CXCL16, CXCL17) as well as CXC chemokine receptors— Gawro´nska-Szklarz,B.; Gutowska, I.; CXCR1, CXCR2, CXCR3, CXCR4, CXCR5, CXCR6, CXCR7 and CXCR8. -

G Protein-Coupled Receptors
S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2015/16: G protein-coupled receptors. British Journal of Pharmacology (2015) 172, 5744–5869 THE CONCISE GUIDE TO PHARMACOLOGY 2015/16: G protein-coupled receptors Stephen PH Alexander1, Anthony P Davenport2, Eamonn Kelly3, Neil Marrion3, John A Peters4, Helen E Benson5, Elena Faccenda5, Adam J Pawson5, Joanna L Sharman5, Christopher Southan5, Jamie A Davies5 and CGTP Collaborators 1School of Biomedical Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK, 2Clinical Pharmacology Unit, University of Cambridge, Cambridge, CB2 0QQ, UK, 3School of Physiology and Pharmacology, University of Bristol, Bristol, BS8 1TD, UK, 4Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK, 5Centre for Integrative Physiology, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2015/16 provides concise overviews of the key properties of over 1750 human drug targets with their pharmacology, plus links to an open access knowledgebase of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. The full contents can be found at http://onlinelibrary.wiley.com/doi/ 10.1111/bph.13348/full. G protein-coupled receptors are one of the eight major pharmacological targets into which the Guide is divided, with the others being: ligand-gated ion channels, voltage-gated ion channels, other ion channels, nuclear hormone receptors, catalytic receptors, enzymes and transporters. These are presented with nomenclature guidance and summary information on the best available pharmacological tools, alongside key references and suggestions for further reading. -

1 Supplemental Material Maresin 1 Activates LGR6 Receptor
Supplemental Material Maresin 1 Activates LGR6 Receptor Promoting Phagocyte Immunoresolvent Functions Nan Chiang, Stephania Libreros, Paul C. Norris, Xavier de la Rosa, Charles N. Serhan Center for Experimental Therapeutics and Reperfusion Injury, Department of Anesthesiology, Perioperative and Pain Medicine, Brigham and Women’s Hospital and Harvard Medical School, Boston, Massachusetts 02115, USA. 1 Supplemental Table 1. Screening of orphan GPCRs with MaR1 Vehicle Vehicle MaR1 MaR1 mean RLU > GPCR ID SD % Activity Mean RLU Mean RLU + 2 SD Mean RLU Vehicle mean RLU+2 SD? ADMR 930920 33283 997486.5381 863760 -7% BAI1 172580 18362 209304.1828 176160 2% BAI2 26390 1354 29097.71737 26240 -1% BAI3 18040 758 19555.07976 18460 2% CCRL2 15090 402 15893.6583 13840 -8% CMKLR2 30080 1744 33568.954 28240 -6% DARC 119110 4817 128743.8016 126260 6% EBI2 101200 6004 113207.8197 105640 4% GHSR1B 3940 203 4345.298244 3700 -6% GPR101 41740 1593 44926.97349 41580 0% GPR103 21413 1484 24381.25067 23920 12% NO GPR107 366800 11007 388814.4922 360020 -2% GPR12 77980 1563 81105.4653 76260 -2% GPR123 1485190 46446 1578081.986 1342640 -10% GPR132 860940 17473 895885.901 826560 -4% GPR135 18720 1656 22032.6827 17540 -6% GPR137 40973 2285 45544.0809 39140 -4% GPR139 438280 16736 471751.0542 413120 -6% GPR141 30180 2080 34339.2307 29020 -4% GPR142 105250 12089 129427.069 101020 -4% GPR143 89390 5260 99910.40557 89380 0% GPR146 16860 551 17961.75617 16240 -4% GPR148 6160 484 7128.848113 7520 22% YES GPR149 50140 934 52008.76073 49720 -1% GPR15 10110 1086 12282.67884 -

G Protein‐Coupled Receptors
S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2019/20: G protein-coupled receptors. British Journal of Pharmacology (2019) 176, S21–S141 THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: G protein-coupled receptors Stephen PH Alexander1 , Arthur Christopoulos2 , Anthony P Davenport3 , Eamonn Kelly4, Alistair Mathie5 , John A Peters6 , Emma L Veale5 ,JaneFArmstrong7 , Elena Faccenda7 ,SimonDHarding7 ,AdamJPawson7 , Joanna L Sharman7 , Christopher Southan7 , Jamie A Davies7 and CGTP Collaborators 1School of Life Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK 2Monash Institute of Pharmaceutical Sciences and Department of Pharmacology, Monash University, Parkville, Victoria 3052, Australia 3Clinical Pharmacology Unit, University of Cambridge, Cambridge, CB2 0QQ, UK 4School of Physiology, Pharmacology and Neuroscience, University of Bristol, Bristol, BS8 1TD, UK 5Medway School of Pharmacy, The Universities of Greenwich and Kent at Medway, Anson Building, Central Avenue, Chatham Maritime, Chatham, Kent, ME4 4TB, UK 6Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK 7Centre for Discovery Brain Sciences, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2019/20 is the fourth in this series of biennial publications. The Concise Guide provides concise overviews of the key properties of nearly 1800 human drug targets with an emphasis on selective pharmacology (where available), plus links to the open access knowledgebase source of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. Although the Concise Guide represents approximately 400 pages, the material presented is substantially reduced compared to information and links presented on the website. -

Initial, Transient, and Specific Interaction Between G Protein
Sato T. et al. Medical Research Archives, vol. 6, issue 9, September 2018 Page 1 of 25 ARTICLE Initial, transient, and specific interaction between G protein-coupled receptor and target G protein in parallel signal processing: a case of olfactory discrimination of cancer-induced odors Takaaki Sato1, Mutsumi Matsukawa2, Yoichi Mizutani3, Toshio Iijima4, Hiroyoshi Matsumura5 Authors’ affiliations: 1 Biomedical Research Institute, National Institute of Advanced Industrial Science and Technology, Osaka, Japan 2 Division of Anatomical Science, Department of Functional Morphology, Nihon University School of Medicine, Tokyo, Japan 3 Department of Medical Engineering, Faculty of Health Science, Aino University, Osaka, Japan 4 Graduate School of Life Sciences, Tohoku University, Sendai, Japan 5 College of Life Sciences, Ritsumeikan University, Kusatsu, Japan * Corresponding author: Takaaki Sato, Biomedical Research Institute, National Institute of Ad- vanced Industrial Science and Technology, 1-8-31 Midorioka, Ikeda, Osaka 563-8577, Japan, E-mail: [email protected] Abstract: G protein-coupled receptors (GPCRs) detect and distinguish between various subtypes of extracellular sig- nals, such as neurotransmitters, hormones, light, and odorous chemicals. As determinants for robust and appropriate cellular responses, common and unique features of interactions between GPCRs and their target G proteins provide insights into structure-based drug design for treatment of GPCR-related diseases. Re- cently, we found that the hydrophobic core buried between GPCR helix 8 and TM1–2 is essential for acti- vation of both specific and nonspecific G proteins. Furthermore, the 2nd residue of helix 8 is responsible for initial, transient, and specific interaction with a target G protein. Analysis of human and murine olfactory receptors (ORs) and other class-A GPCRs revealed that several amino acids, such as Glu, Gln, and Asp, are conserved at this position.