Genetic Effects of Local Extinction
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Extinction Patterns, Δ18 O Trends, and Magnetostratigraphy from a Southern High-Latitude Cretaceous–Paleogene Section: Links with Deccan Volcanism
Palaeogeography, Palaeoclimatology, Palaeoecology 350–352 (2012) 180–188 Contents lists available at SciVerse ScienceDirect Palaeogeography, Palaeoclimatology, Palaeoecology journal homepage: www.elsevier.com/locate/palaeo Extinction patterns, δ18 O trends, and magnetostratigraphy from a southern high-latitude Cretaceous–Paleogene section: Links with Deccan volcanism Thomas S. Tobin a,⁎, Peter D. Ward a, Eric J. Steig a, Eduardo B. Olivero b, Isaac A. Hilburn c, Ross N. Mitchell d, Matthew R. Diamond c, Timothy D. Raub e, Joseph L. Kirschvink c a University of Washington, Earth and Space Sciences, Box 351310, Seattle WA 98195, United States b CADIC-CONICET, Bernardo Houssay, V9410CAB, Ushuaia, Tierra del Fuego, Argentina c California Institute of Technology, Geological and Planetary Sciences, 1200 E. California Blvd. Pasadena CA 91125, United States d Yale University, Geology & Geophysics, 230 Whitney Ave. New Haven, CT 06511, United States e University of St. Andrews, Department of Earth Sciences, St. Andrews KY16 9AL, UK article info abstract Article history: Although abundant evidence now exists for a massive bolide impact coincident with the Cretaceous–Paleogene Received 19 April 2012 (K–Pg) mass extinction event (~65.5 Ma), the relative importance of this impact as an extinction mechanism is Received in revised form 27 May 2012 still the subject of debate. On Seymour Island, Antarctic Peninsula, the López de Bertodano Formation yields one Accepted 8 June 2012 of the most expanded K–Pg boundary sections known. Using a new chronology from magnetostratigraphy, and Available online 10 July 2012 isotopic data from carbonate-secreting macrofauna, we present a high-resolution, high-latitude paleotemperature record spanning this time interval. -

Estimating Rates of Local Species Extinction, Colonization, and Turnover in Animal Communities
Ecological Applications, 8(4), 1998, pp. 1213±1225 q 1998 by the Ecological Society of America ESTIMATING RATES OF LOCAL SPECIES EXTINCTION, COLONIZATION, AND TURNOVER IN ANIMAL COMMUNITIES JAMES D. NICHOLS,1 THIERRY BOULINIER,2 JAMES E. HINES,1 KENNETH H. POLLOCK,3 AND JOHN R. SAUER1 1U.S. Geological Survey, Biological Resources Division, Patuxent Wildlife Research Center, Laurel, Maryland 20708 USA 2North Carolina Cooperative Fish and Wildlife Research Unit, North Carolina State University, Raleigh, North Carolina 27695 USA 3Institute of Statistics, North Carolina State University, Box 8203, Raleigh, North Carolina 27695-8203 USA Abstract. Species richness has been identi®ed as a useful state variable for conservation and management purposes. Changes in richness over time provide a basis for predicting and evaluating community responses to management, to natural disturbance, and to changes in factors such as community composition (e.g., the removal of a keystone species). Prob- abilistic capture±recapture models have been used recently to estimate species richness from species count and presence±absence data. These models do not require the common assumption that all species are detected in sampling efforts. We extend this approach to the development of estimators useful for studying the vital rates responsible for changes in animal communities over time: rates of local species extinction, turnover, and coloni- zation. Our approach to estimation is based on capture±recapture models for closed animal populations that permit heterogeneity in detection probabilities among the different species in the sampled community. We have developed a computer program, COMDYN, to compute many of these estimators and associated bootstrap variances. -

Addressing the Mammoth in the Room: the Ethical and Political Implications of De- Extinction
Addressing the Mammoth in the room: The ethical and political implications of de- extinction (Ashlock, 2013) Lowieke Vermeulen (S4374452) Political Science: Political Theory Radboud University, Nijmegen, Netherlands Supervisor: prof. dr. Marcel Wissenburg Date: August 12, 2019 Word count: 23590 1 Table of Contents Chapter 1: Introduction...............................................................................................................3 1.2 Thesis structure............................................................................................................................6 Chapter 2: De-extinction and species selection..........................................................8 2.1 Extinction........................................................................................................................................9 2.2 Approaches to de-extinction.................................................................................................10 2.2.1 Back-breeding.........................................................................................................................10 2.2.2 Cloning.......................................................................................................................................12 2.2.3 Genetic engineering..............................................................................................................12 2.2.4 Mixed approaches..................................................................................................................13 2.3 -
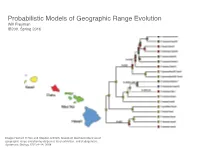
Probabilistic Models of Geographic Range Evolution Will Freyman 10 SYSTEMATIC BIOLOGY VOL
Probabilistic Models of Geographic Range Evolution Will Freyman 10 SYSTEMATIC BIOLOGY VOL. 57 IB200, Spring 2016 Downloaded from http://sysbio.oxfordjournals.org/ at University of California School Law (Boalt Hall) on April 12, 2016 Image: Richard H Ree and Stephen A Smith. Maximum likelihood inference of geographic range evolution by dispersal, local extinction, and cladogenesis. Systematic Biology, 57(1):4–14, 2008. FIGURE 3. Copyedited by: TRJ MANUSCRIPT CATEGORY: Article 2013 LANDIS ET AL.—BAYESIAN BIOGEOGRAPHY FOR MANY AREAS 3 A) B) There are an infinite number of biogeographic 1234011001 010 011 histories that can explain the observed geographic 011 001 ranges. When calculating the probability of the observed Biogeographic histories on geographic ranges at the tips of the phylogenetic tree, 010 a phylogeny: 5 011 it is unreasonable to condition on a specific history 001 101 of biogeographic change. After all, the past history 101 of biogeographic change is not observable. Instead, 6 111 011 the usual approach is to marginalize over all possible 111 histories of biogeographic change that could give rise 7 to the observed geographic ranges. The standard way 111 Downloaded from 101 to do this is to assume that events of colonization or local extinction occur according to a continuous- 8 101 time Markov chain (Ree et al. 2005). Marginalizing over histories of biogeographic change is accomplished C) D) 011101 010 011 011001 010 011 using two procedures. First, exponentiation of the 010 instantaneous-rate matrix, Q, gives the probability http://sysbio.oxfordjournals.org/ 011 010 011 011 011 011 111 111 001 density of all possible biogeographic changes along a 011 010 branch 111 010 101 011 011 111 011 001 Qt 111 101 011 p(y z t,Q) e− , 111 101 → ; = yz 001 % & 101 111 111 011 where y is the ancestral geographic range, z is the 011 001 current geographic range, and t is the duration of the 001 011 branch on the tree. -

Determining Species Expansion and Extinction Possibilities Using Probabilistic and Graphical Models
Ekológia (Bratislava) Vol. 34, No. 2, p. 111–120, 2015 DOI:10.1515/eko-2015-0012 DetermininG species EXpansion AND EXtinction possibilities UsinG probabilistic AND Graphical models RAJESH CHATURVEDI, P. RAMA CHANDRA PRASAD Lab for Spatial Informatics, International Institute of Information Technology, Gachibowli, Hyderabad - 500032, India; e-mail: [email protected], [email protected] Abstract Chaturvedi R., Prasad R.Ch.: Determining species expansion and extinction possibilities using probabilistic and graphical models. Ekológia (Bratislava), Vol. 34, No. 2, p. 111–120, 2015. Survival of plant species is governed by a number of functions. The participation of each function in species survival and the impact of the contrary behaviour of the species vary from function to function. The probability of extinction of species varies in all such scenarios and has to be calculated separately. Secondly, species follow different patterns of dispersal and localisation at different stages of occupancy state of the site, therefore, the scenarios of competition for resources with climatic shifts leading to deterioration and loss of biodiversity resulting in extinction needs to be studied. Furthermore, most po- ssible deviations of species from climax community states needs to be calculated before species become extinct due to sudden environmental disruption. Globally, various types of anthropogenic disturbances threaten the diversity of biological systems. The impact of these anthropogenic activities needs to be analysed to identify extinction patterns with respect to these activities. All the analyses mentioned abo- ve have been tried to be achieved through probabilistic or graphical models in this study. Key words: biodiversity, disturbances, climate change, endangered, risks, ecological models. -

Mammalian Evolution During the Cretaceous-Tertiary Transition; Evidence Fo~
Acta Palaeontologica Polonica Vol. 28, No. 1-2 pp, 55-61 Warszawa, 1983 Second Symposium on Mesozoic T errestial Ecosystems, Jadwisin 1981 WILLIAM A. CLEMENS MAMMALIAN EVOLUTION DURING THE CRETACEOUS-TERTIARY TRANSITION; EVIDENCE FO~ . GRADUAL, NON-CATASTROPHIC PATTERNS OF BIOTIC CHANGE CLEMENS WILLIAM A. : Mammalian evolution during the Cretaceous-Tertiary transition; ev idence for gradual, non-catastrophic patterns of biotic change. Acta Palaeont. Polonlca, 28, 1-2, 55-61, 1983, Hypotheses Invoking catastrophic causal factors for terminal Cretaceous extinctions call for, 1) Instantaneous, global extinctions, 2) resulting from a single or dominant causal factor. 'c u rr entl y available methods of correlation are not precise enough to test the first element of these hypotheses. Patterns of mammalian extinction and survival In northeastern Montana, USA - as well as records of other organisms - argue against the second element. In Montana most of the lineages of marsupials were decimated. Extinction of multltuber curates was not as severe, while only one lineage of eutherlans appears to have become extinct. Groups that apparently Immigrated Into northeastern Montana just prior to the end of the Cretaceous had a greater probability of survival than the residents. The prObability of extinction was Independent of Individual body size. Different patterns of mammalian extinction In different environments also appear to defy explanation In terms of one common, catastrophic causal factor. Key w 0 r d s: Mammalia, Cretaceous, Tertiary, extinction, Montana, USA . Wltltam A . Clemens. Department Of Paleontology. University 01 Callfornla. Berkeley, Callfornla. 94720. USA. Recei ved: September 1981. INTRODUCTION Scientific and popular publications continue to be flooded with articles concerning the causes of the extinction of dinosaurs and other organisms used to mark the end of the Cretaceous. -

Evidence for Local and Global Redox Conditions at an Early Ordovician (Tremadocian) Mass Extinction ∗ ∗∗ Cole T
Earth and Planetary Science Letters 481 (2018) 125–135 Contents lists available at ScienceDirect Earth and Planetary Science Letters www.elsevier.com/locate/epsl Evidence for local and global redox conditions at an Early Ordovician (Tremadocian) mass extinction ∗ ∗∗ Cole T. Edwards a,b, , David A. Fike b, Matthew R. Saltzman c, Wanyi Lu d, Zunli Lu d, a Department of Geological and Environmental Sciences, Appalachian State University, Boone, NC 28608, USA b Department of Earth and Planetary Sciences, Washington University, St. Louis, MO 63130, USA c School of Earth Sciences, The Ohio State University, Columbus, OH 43214, USA d Department of Earth Sciences, Syracuse University, Syracuse, NY 13244, USA a r t i c l e i n f o a b s t r a c t Article history: Profound changes in environmental conditions, particularly atmospheric oxygen levels, are thought to be Received 29 May 2017 important drivers of several major biotic events (e.g. mass extinctions and diversifications). The early Received in revised form 20 September Paleozoic represents a key interval in the oxygenation of the ocean–atmosphere system and evolution 2017 of the biosphere. Global proxies (e.g. carbon (δ13C) and sulfur (δ34S) isotopes) are used to diagnose Accepted 1 October 2017 potential changes in oxygenation and infer causes of environmental change and biotic turnover. The Available online xxxx Cambrian–Ordovician contains several trilobite extinctions (some are apparently local, but others are Editor: D. Vance 13 34 globally correlative) that are attributed to anoxia based on coeval positive δ C and δ S excursions. These Keywords: extinction and excursion events have yet to be coupled with more recently developed proxies thought Ordovician to be more reflective of local redox conditions in the water column (e.g. -

The Indirect Paths to Cascading Effects of Extinctions in Mutualistic Networks
The indirect paths to cascading effects of extinctions in mutualistic networks Item Type Article Authors Pires, Mathias M.; O'Donnell, James L.; Burkle, Laura A.; Díaz- Castelazo, Cecilia; Hembry, David H.; Yeakel, Justin D.; Newman, Erica A.; Medeiros, Lucas P.; de Aguiar, Marcus A. M.; Guimarães, Paulo R. Jr Citation Pires, M. M., O'Donnell, J. L., Burkle, L. A., DíazCastelazo, C., Hembry, D. H., Yeakel, J. D., ... & Guimarães Jr, P. R. (2020). The indirect paths to cascading effects of extinctions in mutualistic networks. Ecology, e03080. DOI 10.1002/ecy.3080 Publisher WILEY Journal ECOLOGY Rights © 2020 by the Ecological Society of America. Download date 05/10/2021 23:12:03 Item License http://rightsstatements.org/vocab/InC/1.0/ Version Final published version Link to Item http://hdl.handle.net/10150/648146 Reports Ecology, 101(7), 2020, e03080 © 2020 by the Ecological Society of America The indirect paths to cascading effects of extinctions in mutualistic networks 1,11 2 3 4 MATHIAS M. PIRES , JAMES L. O'DONNELL, LAURA A. BURKLE , CECILIA DIAZ-CASTELAZO , 5,6 7 6 8 DAVID H. HEMBRY , JUSTIN D. YEAKEL , ERICA A. NEWMAN , LUCAS P. M EDEIROS , 9 10 MARCUS A. M. DE AGUIAR , AND PAULO R. GUIMARAES~ JR. 1Departamento de Biologia Animal, Instituto de Biologia, Universidade Estadual de Campinas, Campinas,13.083-862, Sao~ Paulo, Brazil 2School of Marine and Environmental Affairs, University of Washington, Seattle, WA 98105, Washington, USA 3Department of Ecology, Montana State University, Bozeman, MT 59717, Montana, USA 4Red de Interacciones -

A Review on the Effect of Habitat Fragmentation on Ecosystem
CORE Metadata, citation and similar papers at core.ac.uk Provided by International Institute for Science, Technology and Education (IISTE): E-Journals Journal of Natural Sciences Research www.iiste.org ISSN 2224-3186 (Paper) ISSN 2225-0921 (Online) Vol.6, No.15, 2016 A Review on the Effect of Habitat Fragmentation on Ecosystem Dagnachew Mullu Department of Biology, College of Natural Sciences, Arba Minch, University PO Box 21, Arba Minch Ethiopia Abstract Habitat fragmentation is considered a primary issue in conservation biology. This concern centers around the disruption of once large continuous blocks of habitat into less continuous habitat, primarily by human disturbances such as land clearing and conversion of vegetation from one type to another. Habitat loss and fragmentation are the primary causes of species extinction worldwide. The largest single threat to biological diversity worldwide is the outright destruction of habitat, along with habitat alteration and fragmentation of large habitats into smaller patches. Fragmentation is caused by both natural forces and human activities, each acting over various time frames and spatial scales. Physical Features of the landscape, associated with very slow geomorphic processes (e.g., erosion) may also cause some patches to remain isolated over evolutionary time- scales. The effects of fragmentation also vary depending on the cause of fragmentation (for example, fragmentation of agriculture versus for logging). As a result, there is necessity to take effective actions to maintain biodiversity in fragmented landscapes. Keywords: corridors, fragmentation, island, habitat, metapopulation. 1. Introduction Landscapes all over the world are being fragmented at an unprecedented rate due to fragmentation of the natural habitats of wildlife. -

Downloaded on 15 July 2016)
bioRxiv preprint doi: https://doi.org/10.1101/566406; this version posted March 4, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Integrating behaviour and ecology into global 2 biodiversity conservation strategies 3 4 Joseph A. Tobias1, Alex L. Pigot2 5 6 1Department of Life Sciences, Imperial College London, Silwood Park, Buckhurst Road, Ascot, Berkshire, 7 SL5 7PY, UK. 8 2Centre for Biodiversity and Environment Research, Department of Genetics, Evolution and Environment, 9 University College London, Gower Street, London, WC1E 6BT, UK. 10 11 Keywords: behavioural ecology, birds, indicators, latent risk, macroecology, priority-setting 12 13 14 Summary 15 16 Insights into animal behaviour play an increasingly central role in species-focused conservation practice. 17 However, progress towards incorporating behaviour into regional or global conservation strategies has been 18 far more limited, not least because standardised datasets of behavioural traits are generally lacking at wider 19 taxonomic or spatial scales. Here we make use of the recent expansion of global datasets for birds to assess the 20 prospects for including behavioural traits in systematic conservation priority-setting and monitoring 21 programmes. Using IUCN Red List classification for >9500 bird species, we show that the incidence of threat 22 can vary substantially across different behavioural syndromes, and that some types of behaviour—including 23 particular foraging, mating and migration strategies—are significantly more threatened than others. -

Ecological Response of Plankton to Environmental Change – Thresholds for Extinction
1 This manuscript has been submitted for publication in Annual Reviews of Earth and Planetary Sciences. It 2 will probably change somewhat before it is accepted. If accepted, the final version of the manuscript will be 3 available via the “Peer-reviewed Publication DOI” link on the PaleorXiv page. Please feel free to contact 4 the authors with feedback. 5 6 When citing this paper, please use the following: Lowery, C.M., Bown, P., Fraass, A.J., Hull, P.M., 7 (2019). Ecological Response of Plankton to Environmental Change – Thresholds for Extinction. 8 Submitted to Annual Review of Earth and Planetary Sciences v. 48. 9 10 Ecological Response of Plankton to Environmental Change – Thresholds for 11 Extinction 12 Christopher M. Lowery1*, Paul Bown2, Andrew J. Fraass3, Pincelli M. Hull4 13 14 1University of Texas Institute for Geophysics, University of Texas, Austin 15 2Department of Earth Sciences, University College, London 16 3School of Earth Sciences, University of Bristol 17 4Department of Geology and Geophysics, Yale University 18 19 *Corresponding Author: [email protected] 20 21 Abstract 22 Severe climatic and environmental changes are far more prevalent in Earth history than major 23 extinction events, and the relationship between environmental change and extinction severity has 24 important implications for the outcome of the ongoing anthropogenic extinction event. The response of 25 mineralized marine plankton to environmental change offers an interesting contrast to the overall record 1 26 of marine biota, which is dominated by benthic invertebrates. Here, we summarize changes in the species 27 diversity of planktic foraminifera and calcareous nannoplankton over the Mesozoic-Cenozoic and that of 28 radiolarians and diatoms over the Cenozoic. -

Effects of Habitat Loss and Fragmentation on Amphibians: a Review and Prospectus
BIOLOGICAL CONSERVATION 128 (2006) 231– 240 available at www.sciencedirect.com journal homepage: www.elsevier.com/locate/biocon Effects of habitat loss and fragmentation on amphibians: A review and prospectus Samuel A. Cushman* USDA Forest Service, Rocky Mountain Research Station, P.O. Box 8089, Missoula, MT 59801, USA ARTICLE INFO ABSTRACT Article history: Habitat loss and fragmentation are among the largest threats to amphibian populations. Received 17 March 2005 However, most studies have not provided clear insights into their population-level implica- Received in revised form tions. There is a critical need to investigate the mechanisms that underlie patterns of distri- 26 September 2005 bution and abundance. In order to understand the population- and species-level implications Accepted 29 September 2005 of habitat loss and fragmentation, it is necessary to move from site-specific inferences to Available online 7 November 2005 assessments of how the influences of multiple factors interact across extensive landscapes to influence population size and population connectivity. The goal of this paper is to summa- Keywords: rize the state of knowledge, identify information gaps and suggest research approaches to Habitat loss provide reliable knowledge and effective conservation of amphibians in landscapes experi- Habitat fragmentation encing habitat loss and fragmentation. Reliable inferences require attention to species- Amphibians specific ecological characteristics and their interactions with environmental conditions at Dispersal a range of spatial scales. Habitat connectivity appears to play a key role in regional viability Persistence of amphibian populations. In amphibians, population connectivity is predominantly effected Extinction through juvenile dispersal. The preponderance of evidence suggests that the short-term Review impact of habitat loss and fragmentation increases with dispersal ability.