First Insight Into the Viral Community of the Cnidarian Model Metaorganism Aiptasia Using RNA-Seq Data
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Download Animal List
MICHIGAN'S SPECIAL ANIMALS Endangered, Threatened, Special Concern, and Probably Extirpated This list presents the Endangered (E), Threatened (T), and Probably Extirpated (X) animal species of Michigan, which are protected under the Endangered Species Act of the State of Michigan (Part 365 of PA 451, 1994 Michigan Natural Resources and Environmental Protection Act). The current list became effective on April 9, 2009, after extensive review by technical advisors to the Michigan Department of Natural Resources and the citizenry of the state. Also included in this list are animal species of Special Concern (SC). While not afforded legal protection under the Act, many of these species are of concern because of declining or relict populations in the state. Should these species continue to decline, they would be recommended for Threatened or Endangered status. Protection of Special Concern species now, before they reach dangerously low population levels, would prevent the need to list them in the future by maintaining adequate numbers of self-sustaining populations within Michigan. Some other potentially rare species are listed as Special Concern pending more precise information on their status in the state; when such information becomes available, they could be moved to threatened or endangered status or deleted from the list. This list was produced by the Endangered Species Program of the Michigan Department of Natural Resources and the Michigan Natural Features Inventory. English names in common usage or from published sources have been incorporated, when possible, to promote public understanding of and participation in the Endangered Species Program. To comment on the list or request additional copies, or for information on the Endangered Species Program, contact the Endangered Species Coordinator, Wildlife Division, Michigan Department of Natural Resources, P.O. -

The Viruses of Wild Pigeon Droppings
The Viruses of Wild Pigeon Droppings Tung Gia Phan1,2, Nguyen Phung Vo1,3,A´ kos Boros4,Pe´ter Pankovics4,Ga´bor Reuter4, Olive T. W. Li6, Chunling Wang5, Xutao Deng1, Leo L. M. Poon6, Eric Delwart1,2* 1 Blood Systems Research Institute, San Francisco, California, United States of America, 2 Department of Laboratory Medicine, University of California San Francisco, San Francisco, California, United States of America, 3 Pharmacology Department, School of Pharmacy, Ho Chi Minh City University of Medicine and Pharmacy, Ho Chi Minh, Vietnam, 4 Regional Laboratory of Virology, National Reference Laboratory of Gastroenteric Viruses, A´ NTSZ Regional Institute of State Public Health Service, Pe´cs, Hungary, 5 Stanford Genome Technology Center, Stanford, California, United States of America, 6 Centre of Influenza Research and School of Public Health, University of Hong Kong, Hong Kong SAR Abstract Birds are frequent sources of emerging human infectious diseases. Viral particles were enriched from the feces of 51 wild urban pigeons (Columba livia) from Hong Kong and Hungary, their nucleic acids randomly amplified and then sequenced. We identified sequences from known and novel species from the viral families Circoviridae, Parvoviridae, Picornaviridae, Reoviridae, Adenovirus, Astroviridae, and Caliciviridae (listed in decreasing number of reads), as well as plant and insect viruses likely originating from consumed food. The near full genome of a new species of a proposed parvovirus genus provisionally called Aviparvovirus contained an unusually long middle ORF showing weak similarity to an ORF of unknown function from a fowl adenovirus. Picornaviruses found in both Asia and Europe that are distantly related to the turkey megrivirus and contained a highly divergent 2A1 region were named mesiviruses. -

Computational Exploration of Virus Diversity on Transcriptomic Datasets
Computational Exploration of Virus Diversity on Transcriptomic Datasets Digitaler Anhang der Dissertation zur Erlangung des Doktorgrades (Dr. rer. nat.) der Mathematisch-Naturwissenschaftlichen Fakultät der Rheinischen Friedrich-Wilhelms-Universität Bonn vorgelegt von Simon Käfer aus Andernach Bonn 2019 Table of Contents 1 Table of Contents 1 Preliminary Work - Phylogenetic Tree Reconstruction 3 1.1 Non-segmented RNA Viruses ........................... 3 1.2 Segmented RNA Viruses ............................. 4 1.3 Flavivirus-like Superfamily ............................ 5 1.4 Picornavirus-like Viruses ............................. 6 1.5 Togavirus-like Superfamily ............................ 7 1.6 Nidovirales-like Viruses .............................. 8 2 TRAVIS - True Positive Details 9 2.1 INSnfrTABRAAPEI-14 .............................. 9 2.2 INSnfrTADRAAPEI-16 .............................. 10 2.3 INSnfrTAIRAAPEI-21 ............................... 11 2.4 INSnfrTAORAAPEI-35 .............................. 13 2.5 INSnfrTATRAAPEI-43 .............................. 14 2.6 INSnfrTBERAAPEI-19 .............................. 15 2.7 INSytvTABRAAPEI-11 .............................. 16 2.8 INSytvTALRAAPEI-35 .............................. 17 2.9 INSytvTBORAAPEI-47 .............................. 18 2.10 INSswpTBBRAAPEI-21 .............................. 19 2.11 INSeqtTAHRAAPEI-88 .............................. 20 2.12 INShkeTCLRAAPEI-44 .............................. 22 2.13 INSeqtTBNRAAPEI-11 .............................. 23 2.14 INSeqtTCJRAAPEI-20 -

Dorset Moth Group
Melwood Moths Database last trap recording 2004 Shortcut Code Taxon Vernacular First Record Recorder Latest Record Recorder Method Comment Hep sylv 15 Hepialus sylvina Orange Swift 20/08/1989 JR Cilix glauc 1651 Cilix glaucata Chinese Character 07/07/1989 JR Habros pyrit 1653 Habrosyne pyritoides Buff Arches 06/07/1987 JR 31/07/1998 JR 80w sheet Teth oc 1654 Tethea ocularis Figure of Eighty 06/07/1987 JR Als aesc 1663 Alsophila aescularia March Moth 01/04/2004 JR 01/04/2004 JR 6w actinic trap 1673 1673 Hemistola chrysoprasaria Small Emerald <2000 JR beat for larvae Larvae on Clematis 1682 1682 Timandra comae Blood-vein 06/07/1987 JR id bis 1702 Idaea biselata Small Fan-footed Wave 06/07/1987 JR Id avers 1713 Idaea aversata Riband Wave 06/07/1987 JR 31/07/1998 JR 80w Sheet Xanth ferrug 1725 Xanthorhoe ferrugata Dark-barred Twin-spot Carpet 20/08/1989 JR Xanth fluct 1728 Xanthorhoe fluctuata Garden Carpet 20/08/1989 JR Lamp suffum 1750 Lampropteryx suffumata Water Carpet 01/04/2004 JR 01/04/2004 JR 6w actinic trap 1738 1738 Epirrhoe alternata Common Carpet 07/05/1988 JR Eul pyral 1758 Eulithis pyraliata Barred Straw 06/07/1987 JR Chloro trunc 1764 Chloroclysta truncata Common Marbled Carpet 19/10/2004 JR 80w sheet Cid fulv 1765 Cidaria fulvata Barred Yellow 06/07/1987 JR Colo pect 1776 Colostygia pectinataria Green Carpet 31/07/1998 JR 15/05/2004 JR 6w actinic trap Horis vitalb 1781 Horisme vitalbata Small Waved Umber 18/06/2000 JR Hydrio furc 1777 Hydriomena furcata July Highflyer 06/07/1987 JR 31/07/1998 JR 80w sheet Epirrit dil 1795 -

Zoogeography of the Holarctic Species of the Noctuidae (Lepidoptera): Importance of the Bering Ian Refuge
© Entomologica Fennica. 8.XI.l991 Zoogeography of the Holarctic species of the Noctuidae (Lepidoptera): importance of the Bering ian refuge Kauri Mikkola, J, D. Lafontaine & V. S. Kononenko Mikkola, K., Lafontaine, J.D. & Kononenko, V. S. 1991 : Zoogeography of the Holarctic species of the Noctuidae (Lepidoptera): importance of the Beringian refuge. - En to mol. Fennica 2: 157- 173. As a result of published and unpublished revisionary work, literature compi lation and expeditions to the Beringian area, 98 species of the Noctuidae are listed as Holarctic and grouped according to their taxonomic and distributional history. Of the 44 species considered to be "naturall y" Holarctic before this study, 27 (61 %) are confirmed as Holarctic; 16 species are added on account of range extensions and 29 because of changes in their taxonomic status; 17 taxa are deleted from the Holarctic list. This brings the total of the group to 72 species. Thirteen species are considered to be introduced by man from Europe, a further eight to have been transported by man in the subtropical areas, and five migrant species, three of them of Neotropical origin, may have been assisted by man. The m~jority of the "naturally" Holarctic species are associated with tundra habitats. The species of dry tundra are frequently endemic to Beringia. In the taiga zone, most Holarctic connections consist of Palaearctic/ Nearctic species pairs. The proportion ofHolarctic species decreases from 100 % in the High Arctic to between 40 and 75 % in Beringia and the northern taiga zone, and from between 10 and 20 % in Newfoundland and Finland to between 2 and 4 % in southern Ontario, Central Europe, Spain and Primorye. -

Viruses in Transplantation - Not Always Enemies
Viruses in transplantation - not always enemies Virome and transplantation ECCMID 2018 - Madrid Prof. Laurent Kaiser Head Division of Infectious Diseases Laboratory of Virology Geneva Center for Emerging Viral Diseases University Hospital of Geneva ESCMID eLibrary © by author Conflict of interest None ESCMID eLibrary © by author The human virome: definition? Repertoire of viruses found on the surface of/inside any body fluid/tissue • Eukaryotic DNA and RNA viruses • Prokaryotic DNA and RNA viruses (phages) 25 • The “main” viral community (up to 10 bacteriophages in humans) Haynes M. 2011, Metagenomic of the human body • Endogenous viral elements integrated into host chromosomes (8% of the human genome) • NGS is shaping the definition Rascovan N et al. Annu Rev Microbiol 2016;70:125-41 Popgeorgiev N et al. Intervirology 2013;56:395-412 Norman JM et al. Cell 2015;160:447-60 ESCMID eLibraryFoxman EF et al. Nat Rev Microbiol 2011;9:254-64 © by author Viruses routinely known to cause diseases (non exhaustive) Upper resp./oropharyngeal HSV 1 Influenza CNS Mumps virus Rhinovirus JC virus RSV Eye Herpes viruses Parainfluenza HSV Measles Coronavirus Adenovirus LCM virus Cytomegalovirus Flaviviruses Rabies HHV6 Poliovirus Heart Lower respiratory HTLV-1 Coxsackie B virus Rhinoviruses Parainfluenza virus HIV Coronaviruses Respiratory syncytial virus Parainfluenza virus Adenovirus Respiratory syncytial virus Coronaviruses Gastro-intestinal Influenza virus type A and B Human Bocavirus 1 Adenovirus Hepatitis virus type A, B, C, D, E Those that cause -

Harper's Island Wetlands Butterflies & Moths (2020)
Introduction Harper’s Island Wetlands (HIW) nature reserve, situated close to the village of Glounthaune on the north shore of Cork Harbour is well known for its birds, many of which come from all over northern Europe and beyond, but there is a lot more to the wildlife at the HWI nature reserve than birds. One of our goals it to find out as much as we can about all aspects of life, both plant and animal, that live or visit HIW. This is a report on the butterflies and moths of HIW. Butterflies After birds, butterflies are probably the one of the best known flying creatures. While there has been no structured study of them on at HIW, 17 of Ireland’s 33 resident and regular migrant species of Irish butterflies have been recorded. Just this summer we added the Comma butterfly to the island list. A species spreading across Ireland in recent years possibly in response to climate change. Hopefully we can set up regular monitoring of the butterflies at HIW in the next couple of years. Butterfly Species Recorded at Harper’s Island Wetlands up to September 2020. Colias croceus Clouded Yellow Pieris brassicae Large White Pieris rapae Small White Pieris napi Green-veined White Anthocharis cardamines Orange-tip Lycaena phlaeas Small Copper Polyommatus icarus Common Blue Celastrina argiolus Holly Blue Vanessa atalanta Red Admiral Vanessa cardui Painted Lady Aglais io Peacock Aglais urticae Small Tortoiseshell Polygonia c-album Comma Speyeria aglaja Dark-green Fritillary Pararge aegeria Speckled Wood Maniola jurtina Meadow Brown Aphantopus hyperantus Ringlet Moths One group of insects that are rarely seen by visitors to HIW is the moths. -

Mini Review Picobirnavirus: a Putative Emerging Threat to Humans And
Advances in Animal and Veterinary Sciences Mini Review Picobirnavirus: A Putative Emerging Threat to Humans and Animals JOBIN JOSE KATTOOR, SHUBHANKAR SIRCAR, SHARAD SAURAB, SHANMUGANATHAN SUBRAMANIYAN, KULDEEP DHAMA, YASHPAL SINGH MALIK* ICAR-Indian Veterinary Research Institute, Izatnagar 243122, Bareilly, Uttar Pradesh, India. Abstract | Diarrheal diseases remain fatal threat to human and animal population with the emergence of new types of pathogens. Among them, viral gastroenteritis plays a lion share with a number ranging over 100 different types including emerging and re-emerging types of viruses. Recent viral metagenomics studies confirm the co-existence of viruses in gastrointestinal tract of several different host species. A Picobirnavirus, consisting of 2 segments, has recently attained attention due to its wide host range and genetic variability. Until 2011, these small viruses were not consid- ered as a separate virus family, when a new family (Picobirnaviridae) was approved by the International Committee on Taxonomy of Viruses (ICTV). Currently two distinct genogroups (GG-I and GG-II) and one predicted genogroup (GG-III) are included in the Picobirnaviridae family. Recently, picobirnavirus infections have been reported from al- most all species including wild animals where persistent infection of the virus is also reported. Picobirnaviruses (PBVs) are also reported as opportunistic pathogens in immuno compromised hosts including HIV infected patients. Presence of atypical picobirnaviruses with shorter genomic segments along with genetic closeness of animal and human PBVs and its ability to infect immuno-compromised hosts pose a heavy threat for all human and animal. Currently RNA dependent RNA polymerase based RT-PCR detection is considered as a rapid and sensitive method for detection of PBV. -

Winter Cutworm: a New Pest Threat in Oregon J
OREGON STATE UNIVERSITY EXTENSION SERVICE Winter Cutworm: A New Pest Threat in Oregon J. Green, A. Dreves, B. McDonald, and E. Peachey Introduction Winter cutworm is the common name for the larval stage of the large yellow underwing moth (Noctua pronuba [Lepidoptera: Noctuidae]). The cutworm has tolerance for cold temperatures, and larval feeding activity persists throughout fall and winter. Adult N. pronuba moths have been detected in Oregon for at least a decade, and the species is common in many different ecological habitats. Epidemic outbreaks of adult moths have occurred periodically in this region, resulting in captures of up to 500 moths per night. However, larval feeding by N. pronuba has not been a problem in Oregon until recently. In 2013 and 2014, there were isolated instances reported, including damage by larvae to sod near Portland and defoliation of herb and flower gardens in Corvallis. In 2015, large numbers of larvae were observed around homes, within golf courses, and in field crops located in Oregon and Washington. Winter cutworms have a wide host range across agricultural, urban, and natural landscapes (Table Photo: Nate McGhee, © Oregon State University. 1, page 2) and are a concern as a potential crop pest that can cause considerable damage in a short highlights general information about winter amount of time. Above-ground damage occurs when cutworm, including identification, scouting recom- larvae chew through tissues near ground level, cut- mendations, and potential control measures. ting the stems off plants. Leaf chewing and root feeding also have been observed. Winter cutworms Jessica Green, faculty research assistant, Department of are gregarious, which means they feed and move in Horticulture; Amy J. -

First Insight Into the Viral Community of the Cnidarian Model Metaorganism Aiptasia Using RNA-Seq Data
First insight into the viral community of the cnidarian model metaorganism Aiptasia using RNA-Seq data Jan D. Brüwer and Christian R. Voolstra Red Sea Research Center, Division of Biological and Environmental Science and Engineering (BESE), King Abdullah University of Science and Technology (KAUST), Thuwal, Makkah, Saudi Arabia ABSTRACT Current research posits that all multicellular organisms live in symbioses with asso- ciated microorganisms and form so-called metaorganisms or holobionts. Cnidarian metaorganisms are of specific interest given that stony corals provide the foundation of the globally threatened coral reef ecosystems. To gain first insight into viruses associated with the coral model system Aiptasia (sensu Exaiptasia pallida), we analyzed an existing RNA-Seq dataset of aposymbiotic, partially populated, and fully symbiotic Aiptasia CC7 anemones with Symbiodinium. Our approach included the selective removal of anemone host and algal endosymbiont sequences and subsequent microbial sequence annotation. Of a total of 297 million raw sequence reads, 8.6 million (∼3%) remained after host and endosymbiont sequence removal. Of these, 3,293 sequences could be assigned as of viral origin. Taxonomic annotation of these sequences suggests that Aiptasia is associated with a diverse viral community, comprising 116 viral taxa covering 40 families. The viral assemblage was dominated by viruses from the families Herpesviridae (12.00%), Partitiviridae (9.93%), and Picornaviridae (9.87%). Despite an overall stable viral assemblage, we found that some viral taxa exhibited significant changes in their relative abundance when Aiptasia engaged in a symbiotic relationship with Symbiodinium. Elucidation of viral taxa consistently present across all conditions revealed a core virome of 15 viral taxa from 11 viral families, encompassing many viruses previously reported as members of coral viromes. -
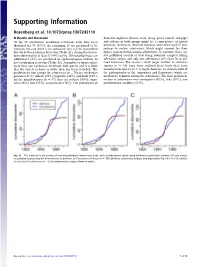
Supporting Information
Supporting Information Rosenberg et al. 10.1073/pnas.1307243110 SI Results and Discussion domestic ungulates (horses, cows, sheep, goats, camels, and pigs) Of the 83 arboviruses, nonhuman vertebrate hosts have been and rodents in both groups might be a consequence of spatial identified for 70 (84%); the remaining 13 are presumed to be proximity to humans. Sentinel monkeys were often used in pro- zoonoses because there is no indication they can be transmitted cedures to isolate arboviruses, which might account for their directly between humans by vectors (Table S1). Animal hosts have higher representation among arboviruses. In contrast, there are been identified for at least 57 (44%) of the 130 nonarboviruses; an few published records of bats being routinely sampled during additional 5 (8%) are presumed on epidemiological evidence to arbovirus studies, and only two arboviruses (3%) have been iso- have nonhuman reservoirs (Table S1). A number of viruses infect lated from bats. The reason a much larger number of arbovirus more than one nonhuman vertebrate host species and it is likely species (n = 16) have been isolated from birds than have that the variety of hosts is wider than has been recorded. The nonarbovirus species (n = 1) might, however, be characteristic of predominant host groups for arboviruses (n = 70) are nonhuman the pathogenicity of the togaviruses and flaviviruses, which are primates (31%), rodents (29%), ungulates (26%), and birds (23%); much more common among the arboviruses. The most prominent for the nonarboviruses (n = 57), they are rodents (30%), ungu- vectors of arboviruses were mosquitoes (67%), ticks (19%), and lates (26%), bats (23%), and primates (16%). -

Libro De Abstract
SPECIAL THANKS EMPRESAS COLABORADORAS SEV VI R O L O G Í A Publicación Oficial de la Sociedad Española de Virología 13th Spanish National Congress Of Virology Madrid 2015 CONGRESO NACIONAL DE Volumen 18 Número 1/2015 EXTRAORDINARIO VIROLOGÍA . PUBLICACIÓN OFICIAL DE LA SOCIEDAD ESPAÑOLA DE VIROLOGÍA 13th Spanish National Congress Of Virology Madrid 2015 Del 7 al 10 de junio de 2015 Auditorio de la Fábrica Nacional de Moneda y Timbre Real Casa de la Moneda de Madrid Volumen 18 Madrid 2015 Número 1/2015 EXTRAORDINARIO Edición y Coordinación: M Angeles Muñoz-Fernández &M. Dolores García-Alonso Diseño y Maquetación: DM&VCH.events.S.L. Diseño Portada: M. Dolores García-Alonso Impresión: Fragma: Servicios de impresión digital en Madrid ISSN (versión digital): 2172-6523 SEV – Sociedad Española de Virología Centro de Biología Molecular “Severo Ochoa” C/ Nicolás Cabrera, 1 28049 Cantoblanco – Madrid [email protected] Página web del congreso: www.congresonacionalvirologia2015.com La responsabilidad del contenido de las colaboraciones publicadas corresponderá a sus autores, quienes autorizan la reproducción de sus artículos a la SEV exclusivamente para esta edición. La SEV no hace necesariamente suyas las opiniones o los criterios expresados por sus colaboradores. Virología. Publicación Oficial de la Sociedad Española de Virología XIII CONGRESO NACIONAL DE VIROLOGÍA B I E N V E N I D A El Comité Organizador del XIII Congreso Nacional de Virología (XIII CNV) tiene el placer de comunicaros que éste se celebrará en Madrid, del 7 al 10 de junio de 2015. En el XIII CNV participan conjuntamente la Sociedad Española de Virología (SEV) y la Sociedad Italiana de Virología (SIV), y estará abierto13th a Spanish la participación National de virólogos de Latinoamérica.