Novel Variants of ELP2 and PIAS1 in the Interferon Gamma Signaling Pathway Are Associated with Non-Small Cell Lung Cancer Survival
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Analysis of Gene Expression Data for Gene Ontology
ANALYSIS OF GENE EXPRESSION DATA FOR GENE ONTOLOGY BASED PROTEIN FUNCTION PREDICTION A Thesis Presented to The Graduate Faculty of The University of Akron In Partial Fulfillment of the Requirements for the Degree Master of Science Robert Daniel Macholan May 2011 ANALYSIS OF GENE EXPRESSION DATA FOR GENE ONTOLOGY BASED PROTEIN FUNCTION PREDICTION Robert Daniel Macholan Thesis Approved: Accepted: _______________________________ _______________________________ Advisor Department Chair Dr. Zhong-Hui Duan Dr. Chien-Chung Chan _______________________________ _______________________________ Committee Member Dean of the College Dr. Chien-Chung Chan Dr. Chand K. Midha _______________________________ _______________________________ Committee Member Dean of the Graduate School Dr. Yingcai Xiao Dr. George R. Newkome _______________________________ Date ii ABSTRACT A tremendous increase in genomic data has encouraged biologists to turn to bioinformatics in order to assist in its interpretation and processing. One of the present challenges that need to be overcome in order to understand this data more completely is the development of a reliable method to accurately predict the function of a protein from its genomic information. This study focuses on developing an effective algorithm for protein function prediction. The algorithm is based on proteins that have similar expression patterns. The similarity of the expression data is determined using a novel measure, the slope matrix. The slope matrix introduces a normalized method for the comparison of expression levels throughout a proteome. The algorithm is tested using real microarray gene expression data. Their functions are characterized using gene ontology annotations. The results of the case study indicate the protein function prediction algorithm developed is comparable to the prediction algorithms that are based on the annotations of homologous proteins. -

Evidence for Differential Alternative Splicing in Blood of Young Boys With
Stamova et al. Molecular Autism 2013, 4:30 http://www.molecularautism.com/content/4/1/30 RESEARCH Open Access Evidence for differential alternative splicing in blood of young boys with autism spectrum disorders Boryana S Stamova1,2,5*, Yingfang Tian1,2,4, Christine W Nordahl1,3, Mark D Shen1,3, Sally Rogers1,3, David G Amaral1,3 and Frank R Sharp1,2 Abstract Background: Since RNA expression differences have been reported in autism spectrum disorder (ASD) for blood and brain, and differential alternative splicing (DAS) has been reported in ASD brains, we determined if there was DAS in blood mRNA of ASD subjects compared to typically developing (TD) controls, as well as in ASD subgroups related to cerebral volume. Methods: RNA from blood was processed on whole genome exon arrays for 2-4–year-old ASD and TD boys. An ANCOVA with age and batch as covariates was used to predict DAS for ALL ASD (n=30), ASD with normal total cerebral volumes (NTCV), and ASD with large total cerebral volumes (LTCV) compared to TD controls (n=20). Results: A total of 53 genes were predicted to have DAS for ALL ASD versus TD, 169 genes for ASD_NTCV versus TD, 1 gene for ASD_LTCV versus TD, and 27 genes for ASD_LTCV versus ASD_NTCV. These differences were significant at P <0.05 after false discovery rate corrections for multiple comparisons (FDR <5% false positives). A number of the genes predicted to have DAS in ASD are known to regulate DAS (SFPQ, SRPK1, SRSF11, SRSF2IP, FUS, LSM14A). In addition, a number of genes with predicted DAS are involved in pathways implicated in previous ASD studies, such as ROS monocyte/macrophage, Natural Killer Cell, mTOR, and NGF signaling. -

ELP2 Antibody (C-Term) Purified Rabbit Polyclonal Antibody (Pab) Catalog # Ap2884b
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 ELP2 Antibody (C-term) Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP2884b Specification ELP2 Antibody (C-term) - Product Information Application IF, WB, FC,E Primary Accession Q6IA86 Reactivity Human Host Rabbit Clonality Polyclonal Isotype Rabbit Ig Calculated MW 92500 Antigen Region 737-765 ELP2 Antibody (C-term) - Additional Information Gene ID 55250 Other Names Elongator complex protein 2, ELP2, SHINC-2, STAT3-interacting protein 1, StIP1, Fluorescent confocal image of SY5Y cells ELP2, STATIP1 stained with ELP2 (C-term) antibody. SY5Y cells were fixed with 4% PFA (20 min), Target/Specificity permeabilized with Triton X-100 (0.2%, 30 This ELP2 antibody is generated from min). Cells were then incubated with rabbits immunized with a KLH conjugated AP2884b ELP2 (C-term) primary antibody synthetic peptide between 737-765 amino (1:100, 2 h at room temperature). For acids from the C-terminal region of human secondary antibody, Alexa Fluor® 488 ELP2. conjugated donkey anti-rabbit antibody (green) was used (1:1000, 1h). Nuclei were Dilution counterstained with Hoechst 33342 (blue) (10 IF~~1:100 WB~~1:1000 μg/ml, 5 min). Note the highly specific FC~~1:10~50 localization of the ELP2 immunosignal mainly to the cytoplasm, supported by Human Format Protein Atlas Data (http://www.proteinatlas.or Purified polyclonal antibody supplied in PBS g/ENSG00000134759). with 0.09% (W/V) sodium azide. This antibody is prepared by Saturated Ammonium Sulfate (SAS) precipitation followed by dialysis against PBS. Storage Maintain refrigerated at 2-8°C for up to 2 weeks. -

Mouse Elp2 Conditional Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Elp2 Conditional Knockout Project (CRISPR/Cas9) Objective: To create a Elp2 conditional knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Elp2 gene (NCBI Reference Sequence: NM_021448 ; Ensembl: ENSMUSG00000024271 ) is located on Mouse chromosome 18. 22 exons are identified, with the ATG start codon in exon 1 and the TGA stop codon in exon 22 (Transcript: ENSMUST00000234266). Exon 2 will be selected as conditional knockout region (cKO region). Deletion of this region should result in the loss of function of the Mouse Elp2 gene. To engineer the targeting vector, homologous arms and cKO region will be generated by PCR using BAC clone RP23-193O18 as template. Cas9, gRNA and targeting vector will be co-injected into fertilized eggs for cKO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Exon 2 starts from about 5.58% of the coding region. The knockout of Exon 2 will result in frameshift of the gene. The size of intron 1 for 5'-loxP site insertion: 2724 bp, and the size of intron 2 for 3'-loxP site insertion: 2694 bp. The size of effective cKO region: ~579 bp. The cKO region does not have any other known gene. Page 1 of 8 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele gRNA region 5' gRNA region 3' 1 2 22 Targeting vector Targeted allele Constitutive KO allele (After Cre recombination) Legends Exon of mouse Elp2 Homology arm cKO region loxP site Page 2 of 8 https://www.alphaknockout.com Overview of the Dot Plot Window size: 10 bp Forward Reverse Complement Sequence 12 Note: The sequence of homologous arms and cKO region is aligned with itself to determine if there are tandem repeats. -

A Yeast Phenomic Model for the Influence of Warburg Metabolism on Genetic Buffering of Doxorubicin Sean M
Santos and Hartman Cancer & Metabolism (2019) 7:9 https://doi.org/10.1186/s40170-019-0201-3 RESEARCH Open Access A yeast phenomic model for the influence of Warburg metabolism on genetic buffering of doxorubicin Sean M. Santos and John L. Hartman IV* Abstract Background: The influence of the Warburg phenomenon on chemotherapy response is unknown. Saccharomyces cerevisiae mimics the Warburg effect, repressing respiration in the presence of adequate glucose. Yeast phenomic experiments were conducted to assess potential influences of Warburg metabolism on gene-drug interaction underlying the cellular response to doxorubicin. Homologous genes from yeast phenomic and cancer pharmacogenomics data were analyzed to infer evolutionary conservation of gene-drug interaction and predict therapeutic relevance. Methods: Cell proliferation phenotypes (CPPs) of the yeast gene knockout/knockdown library were measured by quantitative high-throughput cell array phenotyping (Q-HTCP), treating with escalating doxorubicin concentrations under conditions of respiratory or glycolytic metabolism. Doxorubicin-gene interaction was quantified by departure of CPPs observed for the doxorubicin-treated mutant strain from that expected based on an interaction model. Recursive expectation-maximization clustering (REMc) and Gene Ontology (GO)-based analyses of interactions identified functional biological modules that differentially buffer or promote doxorubicin cytotoxicity with respect to Warburg metabolism. Yeast phenomic and cancer pharmacogenomics data were integrated to predict differential gene expression causally influencing doxorubicin anti-tumor efficacy. Results: Yeast compromised for genes functioning in chromatin organization, and several other cellular processes are more resistant to doxorubicin under glycolytic conditions. Thus, the Warburg transition appears to alleviate requirements for cellular functions that buffer doxorubicin cytotoxicity in a respiratory context. -

Insights Into Functional Connectivity in Mammalian Signal Transduction Pathways by Pairwise
bioRxiv preprint doi: https://doi.org/10.1101/2019.12.30.891200; this version posted December 30, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Insights into Functional Connectivity in Mammalian Signal Transduction Pathways by Pairwise Comparison of Protein Interaction Partners of Critical Signaling Hubs Chilakamarti V. Ramana * Department of Medicine, Dartmouth-Hitchcock Medical Center, Lebanon, NH 03766, USA *Correspondence should be addressed: Chilakamarti V .Ramana, Telephone. (603)-738-2507, E-mail: [email protected] ORCID ID: /0000-0002-5153-8252 1 bioRxiv preprint doi: https://doi.org/10.1101/2019.12.30.891200; this version posted December 30, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Abstract Growth factors and cytokines activate signal transduction pathways and regulate gene expression in eukaryotes. Intracellular domains of activated receptors recruit several protein kinases as well as transcription factors that serve as platforms or hubs for the assembly of multi-protein complexes. The signaling hubs involved in a related biologic function often share common interaction proteins and target genes. This functional connectivity suggests that a pairwise comparison of protein interaction partners of signaling hubs and network analysis of common partners and their expression analysis might lead to the identification of critical nodes in cellular signaling. -

Rabbit Anti-STATIP1/FITC Conjugated Antibody
SunLong Biotech Co.,LTD Tel: 0086-571- 56623320 Fax:0086-571- 56623318 E-mail:[email protected] www.sunlongbiotech.com Rabbit Anti-STATIP1/FITC Conjugated antibody SL12822R-FITC Product Name: Anti-STATIP1/FITC Chinese Name: FITC标记的STAT相互作用蛋白1抗体 AU023723; Elongation protein 2 homolog (S. cerevisiae); Elongator acetyltransferase complex subunit 2; Elongator complex protein 2; elongator protein 2; ELP2; ELP2_HUMAN; Epl2; FLJ10879; SHINC 2; SHINC-2; SHINC2; signal transducer and Alias: activator of transcription 3 interacting protein 1; signal transducer and activator of transcription interacting protein 1; STAT3 interacting protein 1; Stat3 interacting protein; STAT3-interacting protein 1; Stat3-interacting protein; STATIP 1; STATIP1; STATIP1 protein; StIP; StIP1. Organism Species: Rabbit Clonality: Polyclonal React Species: Human,Mouse,Rat,Dog,Pig,Horse, ICC=1:50-200IF=1:50-200 Applications: not yet tested in other applications. optimal dilutions/concentrations should be determined by the end user. Molecular weight: 92kDa Form: Lyophilized or Liquid Concentration: 2mg/1mlwww.sunlongbiotech.com immunogen: KLH conjugated synthetic peptide derived from human STATIP1 Lsotype: IgG Purification: affinity purified by Protein A Storage Buffer: 0.01M TBS(pH7.4) with 1% BSA, 0.03% Proclin300 and 50% Glycerol. Store at -20 °C for one year. Avoid repeated freeze/thaw cycles. The lyophilized antibody is stable at room temperature for at least one month and for greater than a year Storage: when kept at -20°C. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 °C. Function: Regulates the ligand-dependent activation of STAT3. Product Detail: Acts as subunit of the RNA polymerase II elongator complex, which is a histone acetyltransferase component of the RNA polymerase II (Pol II) holoenzyme and is involved in transcriptional elongation. -

Genomic Diagnostics Within a Medically Underserved Population: Efficacy and Implications
© American College of Medical Genetics and Genomics ORIGINAL RESEARCH ARTICLE Genomic diagnostics within a medically underserved population: efficacy and implications Kevin A. Strauss, MD1, Claudia Gonzaga-Jauregui, PhD2, Karlla W. Brigatti, MS1, Katie B. Williams, MD, PhD1, Alejandra K. King, PhD2, Cristopher Van Hout, PhD2, Donna L. Robinson, CRNP1, Millie Young, RNC1, Kavita Praveen, PhD2, Adam D. Heaps, MS1, Mindy Kuebler, MS1, Aris Baras, MD2, Jeffrey G. Reid, PhD2, John D. Overton, PhD2, Frederick E. Dewey, MD2, Robert N. Jinks, PhD3, Ian Finnegan, BA3, Scott J. Mellis, MD, PhD2, Alan R. Shuldiner, MD2 and Erik G. Puffenberger, PhD1 Purpose: We integrated whole-exome sequencing (WES) and Compared to trio analysis, “family” WES (average seven exomes chromosomal microarray analysis (CMA) into a clinical workflow per proband) reduced filtered candidate variants from 22 ± 6to to serve an endogamous, uninsured, agrarian community. 5 ± 3 per proband. Nineteen (51%) alleles were de novo and 17 Methods: Seventy-nine probands (newborn to 49.8 years) who (46%) inherited; the latter added to a population-based diagnostic presented between 1998 and 2015 remained undiagnosed after panel. We found actionable secondary variants in 21 (4.2%) of 502 biochemical and molecular investigations. We generated WES data subjects, all of whom opted to be informed. for probands and family members and vetted variants through Conclusion: CMA and family-based WES streamline and rephenotyping, segregation analyses, and population studies. economize diagnosis of rare genetic disorders, accelerate novel Results: The most common presentation was neurological disease gene discovery, and create new opportunities for community-based (64%). Seven (9%) probands were diagnosed by CMA. -

Journal Pre-Proof
Journal Pre-proof A novel homozygous missense variant in MATN3 causes spondylo-epimetaphyseal dysplasia Matrilin 3 type in a consanguineous family Samina Yasin, Saima Mustafa, Arzoo Ayesha, Muhammad Latif, Mubashir Hassan, Muhammad Faisal, Outi Makitie, Furhan Iqbal, Sadaf Naz PII: S1769-7212(20)30038-0 DOI: https://doi.org/10.1016/j.ejmg.2020.103958 Reference: EJMG 103958 To appear in: European Journal of Medical Genetics Received Date: 22 January 2020 Revised Date: 11 May 2020 Accepted Date: 17 May 2020 Please cite this article as: S. Yasin, S. Mustafa, A. Ayesha, M. Latif, M. Hassan, M. Faisal, O. Makitie, F. Iqbal, S. Naz, A novel homozygous missense variant in MATN3 causes spondylo-epimetaphyseal dysplasia Matrilin 3 type in a consanguineous family, European Journal of Medical Genetics (2020), doi: https://doi.org/10.1016/j.ejmg.2020.103958. This is a PDF file of an article that has undergone enhancements after acceptance, such as the addition of a cover page and metadata, and formatting for readability, but it is not yet the definitive version of record. This version will undergo additional copyediting, typesetting and review before it is published in its final form, but we are providing this version to give early visibility of the article. Please note that, during the production process, errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain. © 2020 Published by Elsevier Masson SAS. Authorship statement Samina Yasin: Methodology, Investigation, Formal analysis, Data curation, -
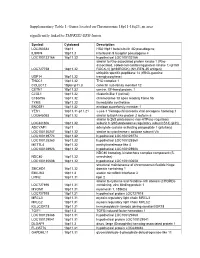
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

Content Based Search in Gene Expression Databases and a Meta-Analysis of Host Responses to Infection
Content Based Search in Gene Expression Databases and a Meta-analysis of Host Responses to Infection A Thesis Submitted to the Faculty of Drexel University by Francis X. Bell in partial fulfillment of the requirements for the degree of Doctor of Philosophy November 2015 c Copyright 2015 Francis X. Bell. All Rights Reserved. ii Acknowledgments I would like to acknowledge and thank my advisor, Dr. Ahmet Sacan. Without his advice, support, and patience I would not have been able to accomplish all that I have. I would also like to thank my committee members and the Biomed Faculty that have guided me. I would like to give a special thanks for the members of the bioinformatics lab, in particular the members of the Sacan lab: Rehman Qureshi, Daisy Heng Yang, April Chunyu Zhao, and Yiqian Zhou. Thank you for creating a pleasant and friendly environment in the lab. I give the members of my family my sincerest gratitude for all that they have done for me. I cannot begin to repay my parents for their sacrifices. I am eternally grateful for everything they have done. The support of my sisters and their encouragement gave me the strength to persevere to the end. iii Table of Contents LIST OF TABLES.......................................................................... vii LIST OF FIGURES ........................................................................ xiv ABSTRACT ................................................................................ xvii 1. A BRIEF INTRODUCTION TO GENE EXPRESSION............................. 1 1.1 Central Dogma of Molecular Biology........................................... 1 1.1.1 Basic Transfers .......................................................... 1 1.1.2 Uncommon Transfers ................................................... 3 1.2 Gene Expression ................................................................. 4 1.2.1 Estimating Gene Expression ............................................ 4 1.2.2 DNA Microarrays ...................................................... -

Title: a Yeast Phenomic Model for the Influence of Warburg Metabolism on Genetic
bioRxiv preprint doi: https://doi.org/10.1101/517490; this version posted January 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. 1 Title Page: 2 3 Title: A yeast phenomic model for the influence of Warburg metabolism on genetic 4 buffering of doxorubicin 5 6 Authors: Sean M. Santos1 and John L. Hartman IV1 7 1. University of Alabama at Birmingham, Department of Genetics, Birmingham, AL 8 Email: [email protected], [email protected] 9 Corresponding author: [email protected] 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 1 bioRxiv preprint doi: https://doi.org/10.1101/517490; this version posted January 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. 26 Abstract: 27 Background: 28 Saccharomyces cerevisiae represses respiration in the presence of adequate glucose, 29 mimicking the Warburg effect, termed aerobic glycolysis. We conducted yeast phenomic 30 experiments to characterize differential doxorubicin-gene interaction, in the context of 31 respiration vs. glycolysis. The resulting systems level biology about doxorubicin 32 cytotoxicity, including the influence of the Warburg effect, was integrated with cancer 33 pharmacogenomics data to identify potentially causal correlations between differential 34 gene expression and anti-cancer efficacy.