Evidence for Differential Alternative Splicing in Blood of Young Boys With
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

PARSANA-DISSERTATION-2020.Pdf
DECIPHERING TRANSCRIPTIONAL PATTERNS OF GENE REGULATION: A COMPUTATIONAL APPROACH by Princy Parsana A dissertation submitted to The Johns Hopkins University in conformity with the requirements for the degree of Doctor of Philosophy Baltimore, Maryland July, 2020 © 2020 Princy Parsana All rights reserved Abstract With rapid advancements in sequencing technology, we now have the ability to sequence the entire human genome, and to quantify expression of tens of thousands of genes from hundreds of individuals. This provides an extraordinary opportunity to learn phenotype relevant genomic patterns that can improve our understanding of molecular and cellular processes underlying a trait. The high dimensional nature of genomic data presents a range of computational and statistical challenges. This dissertation presents a compilation of projects that were driven by the motivation to efficiently capture gene regulatory patterns in the human transcriptome, while addressing statistical and computational challenges that accompany this data. We attempt to address two major difficulties in this domain: a) artifacts and noise in transcriptomic data, andb) limited statistical power. First, we present our work on investigating the effect of artifactual variation in gene expression data and its impact on trans-eQTL discovery. Here we performed an in-depth analysis of diverse pre-recorded covariates and latent confounders to understand their contribution to heterogeneity in gene expression measurements. Next, we discovered 673 trans-eQTLs across 16 human tissues using v6 data from the Genotype Tissue Expression (GTEx) project. Finally, we characterized two trait-associated trans-eQTLs; one in Skeletal Muscle and another in Thyroid. Second, we present a principal component based residualization method to correct gene expression measurements prior to reconstruction of co-expression networks. -

Structure and Function of the Fgd Family of Divergent FYVE Domain Proteins
Biochemistry and Cell Biology Structure and Function of the Fgd Family of Divergent FYVE Domain Proteins Journal: Biochemistry and Cell Biology Manuscript ID bcb-2018-0185.R1 Manuscript Type: Mini Review Date Submitted by the 03-Aug-2018 Author: Complete List of Authors: Eitzen, Gary; University of Alberta Faculty of Medicine and Dentistry Smithers, Cameron C.; University of Alberta, Biochemistry Murray, Allan; University of Alberta Faculty of Medicine and Dentistry Overduin, Michael; University of Alberta Faculty of Medicine and Dentistry Draft Fgd, Pleckstrin Homology domain, FYVE domain, Dbl Homology Domain, Keyword: Rho GEF Is the invited manuscript for consideration in a Special CSMB Special Issue Issue? : https://mc06.manuscriptcentral.com/bcb-pubs Page 1 of 37 Biochemistry and Cell Biology Title: Structure and Function of the Fgd Family of Divergent FYVE Domain Proteins Authors: Gary Eitzen1, Cameron C. Smithers2, Allan G Murray3 and Michael Overduin2* Draft 1Department of Cell Biology, 2Department of Biochemistry, 3Department of Medicine, University of Alberta, Edmonton, Alberta, Canada *Corresponding author. Michael Overduin Telephone: +1 780 492 3518 Fax: +1 780 492-0886 E-mail: [email protected] https://mc06.manuscriptcentral.com/bcb-pubs Biochemistry and Cell Biology Page 2 of 37 Abstract FYVE domains are highly conserved protein modules that typically bind phosphatidylinositol 3-phosphate (PI3P) on the surface of early endosomes. Along with pleckstrin homology (PH) and phox homology (PX) domains, FYVE domains are the principal readers of the phosphoinositide (PI) code that mediate specific recognition of eukaryotic organelles. Of all the human FYVE domain-containing proteins, those within the Faciogenital dysplasia (Fgd) subfamily are particularly divergent, and couple with GTPases to exert unique cellular functions. -

Rhoa Promotes Epidermal Stem Cell Proliferation Via PKN1-Cyclin D1 Signaling
RESEARCH ARTICLE RhoA promotes epidermal stem cell proliferation via PKN1-cyclin D1 signaling Fan Wang1, Rixing Zhan2, Liang Chen1, Xia Dai1, Wenping Wang1, Rui Guo1, Xiaoge Li1, Zhe Li1, Liang Wang1, Shupeng Huang1, Jie Shen1, Shirong Li1☯*, Chuan Cao1☯* 1 Department of Plastic and Reconstructive Surgery, Southwestern Hospital, Third Military Medical University, Chongqing, China, 2 School of Nursing, Third Military Medical University, Chongqing, China ☯ These authors contributed equally to this work. * [email protected] (LS); [email protected] (CC) a1111111111 Abstract a1111111111 a1111111111 a1111111111 a1111111111 Objective Epidermal stem cells (ESCs) play a critical role in wound healing, but the mechanism under- lying ESC proliferation is not well defined. Here, we explore the effects of RhoA on ESC pro- liferation and the possible underlying mechanism. OPEN ACCESS Citation: Wang F, Zhan R, Chen L, Dai X, Wang W, Methods Guo R, et al. (2017) RhoA promotes epidermal (+/+) (-/- stem cell proliferation via PKN1-cyclin D1 Human ESCs were enriched by rapid adhesion to collagen IV. RhoA (G14V), RhoA ) signaling. PLoS ONE 12(2): e0172613. (T19N) and pGFP control plasmids were transfected into human ESCs. The effect of RhoA doi:10.1371/journal.pone.0172613 on cell proliferation was detected by cell proliferation and DNA synthesis assays. Induction Editor: Austin John Cooney, University of Texas at of PKN1 activity by RhoA was determined by immunoblot analysis, and the effects of PKN1 Austin Dell Medical School, UNITED STATES on RhoA in terms of inducing cell proliferation and cyclin D1 expression were detected using Received: August 10, 2016 specific siRNA targeting PKN1. The effects of U-46619 (a RhoA agonist) and C3 transferase Accepted: February 6, 2017 (a RhoA antagonist) on ESC proliferation were observed in vivo. -

Prioritization and Evaluation of Depression Candidate Genes by Combining Multidimensional Data Resources
Prioritization and Evaluation of Depression Candidate Genes by Combining Multidimensional Data Resources Chung-Feng Kao1, Yu-Sheng Fang2, Zhongming Zhao3, Po-Hsiu Kuo1,2,4* 1 Department of Public Health and Institute of Epidemiology and Preventive Medicine, College of Public Health, National Taiwan University, Taipei, Taiwan, 2 Institute of Clinical Medicine, School of Medicine, National Cheng-Kung University, Tainan, Taiwan, 3 Departments of Biomedical Informatics and Psychiatry, Vanderbilt University School of Medicine, Nashville, Tennessee, United States of America, 4 Research Center for Genes, Environment and Human Health, National Taiwan University, Taipei, Taiwan Abstract Background: Large scale and individual genetic studies have suggested numerous susceptible genes for depression in the past decade without conclusive results. There is a strong need to review and integrate multi-dimensional data for follow up validation. The present study aimed to apply prioritization procedures to build-up an evidence-based candidate genes dataset for depression. Methods: Depression candidate genes were collected in human and animal studies across various data resources. Each gene was scored according to its magnitude of evidence related to depression and was multiplied by a source-specific weight to form a combined score measure. All genes were evaluated through a prioritization system to obtain an optimal weight matrix to rank their relative importance with depression using the combined scores. The resulting candidate gene list for depression (DEPgenes) was further evaluated by a genome-wide association (GWA) dataset and microarray gene expression in human tissues. Results: A total of 5,055 candidate genes (4,850 genes from human and 387 genes from animal studies with 182 being overlapped) were included from seven data sources. -

Angio-Associated Migratory Cell Protein Interacts with Epidermal
Cellular Signalling 61 (2019) 10–19 Contents lists available at ScienceDirect Cellular Signalling journal homepage: www.elsevier.com/locate/cellsig Angio-associated migratory cell protein interacts with epidermal growth factor receptor and enhances proliferation and drug resistance in human T non-small cell lung cancer cells Shun Yaoa, Feifei Shia, Yingying Wanga,b, Xiaoyang Suna, Wenbo Suna, Yifeng Zhanga, ⁎ ⁎ Xianfang Liuc, Xiangguo Liua,b, , Ling Sua,b, a Shandong Provincial Key Laboratory of Animal Cell and Developmental Biology, School of Life Sciences, Shandong University, Qingdao, China b Shandong Provincial Collaborative Innovation Center of Cell Biology, School of Life Sciences, Shandong Normal University, Jinan, China c The Department of Otolaryngology Head and Neck Surgery, Shandong Provincial Hospital, Affiliated to Shandong University, Jinan, China ARTICLE INFO ABSTRACT Keywords: Angio-associated migratory cell protein (AAMP) is expressed in some human cancer cells. Previous studies have AAMP shown AAMP high expression predicted poor prognosis. But its biological role in non-small cell lung cancer Proliferation (NSCLC) cells is still unknown. In our present study, we attempted to explore the functions of AAMP in NSCLC Tumorigenesis cells. According to our findings, AAMP knockdown inhibited lung cancer cell proliferation and inhibited lung EGFR cancer cell tumorigenesis in the mouse xenograft model. Epidermal growth factor receptor (EGFR) is a primary Icotinib receptor tyrosine kinase (RTK) that promotes proliferation and plays an important role in cancer pathology. We Doxorubicin found AAMP interacted with EGFR and enhanced its dimerization and phosphorylation at tyrosine 1173 which activated ERK1/2 in NSCLC cells. In addition, we showed AAMP conferred the lung cancer cells resistance to chemotherapeutic agents such as icotinib and doxorubicin. -

Analysis of Gene Expression Data for Gene Ontology
ANALYSIS OF GENE EXPRESSION DATA FOR GENE ONTOLOGY BASED PROTEIN FUNCTION PREDICTION A Thesis Presented to The Graduate Faculty of The University of Akron In Partial Fulfillment of the Requirements for the Degree Master of Science Robert Daniel Macholan May 2011 ANALYSIS OF GENE EXPRESSION DATA FOR GENE ONTOLOGY BASED PROTEIN FUNCTION PREDICTION Robert Daniel Macholan Thesis Approved: Accepted: _______________________________ _______________________________ Advisor Department Chair Dr. Zhong-Hui Duan Dr. Chien-Chung Chan _______________________________ _______________________________ Committee Member Dean of the College Dr. Chien-Chung Chan Dr. Chand K. Midha _______________________________ _______________________________ Committee Member Dean of the Graduate School Dr. Yingcai Xiao Dr. George R. Newkome _______________________________ Date ii ABSTRACT A tremendous increase in genomic data has encouraged biologists to turn to bioinformatics in order to assist in its interpretation and processing. One of the present challenges that need to be overcome in order to understand this data more completely is the development of a reliable method to accurately predict the function of a protein from its genomic information. This study focuses on developing an effective algorithm for protein function prediction. The algorithm is based on proteins that have similar expression patterns. The similarity of the expression data is determined using a novel measure, the slope matrix. The slope matrix introduces a normalized method for the comparison of expression levels throughout a proteome. The algorithm is tested using real microarray gene expression data. Their functions are characterized using gene ontology annotations. The results of the case study indicate the protein function prediction algorithm developed is comparable to the prediction algorithms that are based on the annotations of homologous proteins. -

Regulation of Cdc42 and Its Effectors in Epithelial Morphogenesis Franck Pichaud1,2,*, Rhian F
© 2019. Published by The Company of Biologists Ltd | Journal of Cell Science (2019) 132, jcs217869. doi:10.1242/jcs.217869 REVIEW SUBJECT COLLECTION: ADHESION Regulation of Cdc42 and its effectors in epithelial morphogenesis Franck Pichaud1,2,*, Rhian F. Walther1 and Francisca Nunes de Almeida1 ABSTRACT An overview of Cdc42 Cdc42 – a member of the small Rho GTPase family – regulates cell Cdc42 was discovered in yeast and belongs to a large family of small – polarity across organisms from yeast to humans. It is an essential (20 30 kDa) GTP-binding proteins (Adams et al., 1990; Johnson regulator of polarized morphogenesis in epithelial cells, through and Pringle, 1990). It is part of the Ras-homologous Rho subfamily coordination of apical membrane morphogenesis, lumen formation and of GTPases, of which there are 20 members in humans, including junction maturation. In parallel, work in yeast and Caenorhabditis elegans the RhoA and Rac GTPases, (Hall, 2012). Rho, Rac and Cdc42 has provided important clues as to how this molecular switch can homologues are found in all eukaryotes, except for plants, which do generate and regulate polarity through localized activation or inhibition, not have a clear homologue for Cdc42. Together, the function of and cytoskeleton regulation. Recent studies have revealed how Rho GTPases influences most, if not all, cellular processes. important and complex these regulations can be during epithelial In the early 1990s, seminal work from Alan Hall and his morphogenesis. This complexity is mirrored by the fact that Cdc42 can collaborators identified Rho, Rac and Cdc42 as main regulators of exert its function through many effector proteins. -

HOOK3 Is a Scaffold for the Opposite-Polarity Microtubule-Based
bioRxiv preprint doi: https://doi.org/10.1101/508887; this version posted December 31, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. HOOK3 is a scaffold for the opposite-polarity microtubule-based motors cytoplasmic dynein and KIF1C Agnieszka A. Kendrick1, William B. Redwine1,2†, Phuoc Tien Tran1‡, Laura Pontano Vaites2, Monika Dzieciatkowska4, J. Wade Harper2, and Samara L. Reck-Peterson1,3,5 1Department of Cellular and Molecular Medicine, University of California San Diego, La Jolla, CA, 92093. 2 Department of Cell Biology, Harvard Medical School, Boston, MA 02115. 3Section of Cell and Developmental Biology, Division of Biological Sciences, University of California San Diego, La Jolla, CA 92093. 4Department of Biochemistry and Molecular Genetics, University of Colorado Denver, Aurora, CO 80045. 5Howard Hughes Medical Institute †Present address: Stowers Institute for Medical Research, Kansas City, MO 64110 ‡Present address: Department of Molecular and Cellular Biology, Harvard University, Cambridge, MA 02138. *Correspondence to: Samara Reck-Peterson 9500 Gilman Drive, Leichtag 482 La Jolla CA, 92093 [email protected]; https://orcid.org/0000-0002-1553-465X 1 bioRxiv preprint doi: https://doi.org/10.1101/508887; this version posted December 31, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. -
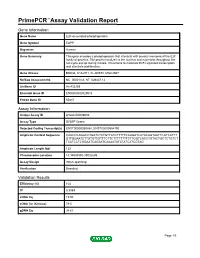
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name E2F-associated phosphoprotein Gene Symbol EAPP Organism Human Gene Summary This gene encodes a phosphoprotein that interacts with several members of the E2F family of proteins. The protein localizes to the nucleus and is present throughout the cell cycle except during mitosis. It functions to modulate E2F-regulated transcription and stimulate proliferation. Gene Aliases BM036, C14orf11, FLJ20578, MGC4957 RefSeq Accession No. NC_000014.8, NT_026437.12 UniGene ID Hs.433269 Ensembl Gene ID ENSG00000129518 Entrez Gene ID 55837 Assay Information Unique Assay ID qHsaCID0008053 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000250454, ENST00000554792 Amplicon Context Sequence CAACCCAGGCCTGATCTCTGTTATCTTTTTCAGGATCATACAGTAATTCGTCATTT GTTGGAATCTTGTGTTGTTTCTTCTTTTTTTTCTTGGTCACCTGTACTGCTCTGTCT TCATCCTCGGAATCAGAATCAAAATATATATCATCGTAG Amplicon Length (bp) 122 Chromosome Location 14:34998580-35002699 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 104 R2 0.9989 cDNA Cq 19.93 cDNA Tm (Celsius) 79.5 gDNA Cq 34.63 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report EAPP, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 3/5 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -

Parental Balanced Chromosomal Rearrangement Leading to Major Genomic Imbalance and an Autosomal Trisomy Resulting in Consecutive Pregnancy Loss: a Case Report
Journal of Genetics (2021)100:54 Ó Indian Academy of Sciences https://doi.org/10.1007/s12041-021-01304-3 (0123456789().,-volV)(0123456789().,-volV) RESEARCH NOTE Parental balanced chromosomal rearrangement leading to major genomic imbalance and an autosomal trisomy resulting in consecutive pregnancy loss: a case report ANUSHKA SHRIVASTAVA1,2, SEEMA THAKUR3, TARA NATH2, ABHIPSA V. F. DEBNATH1 and SONAL R. BAKSHI1* 1Institute of Science, Nirma University, Ahmedabad 382 481, India 2Advanced Genomic Institute and Laboratory Medicine (Labassure), New Delhi 110 003, India 3Department of Clinical Genetics and Fetal Medicine, Fortis Hospital, New Delhi 110 088, India *For correspondence. E-mail: [email protected]. Received 8 November 2020; revised 9 February 2021; accepted 23 March 2021 Abstract. Chromosomal aberrations such as parental balanced translocation contribute to a significant proportion of recurrent pregnancy losses. These have extreme genetic implications on the foetus which can either cause physical and/or mental retardation or early death. In this study, we report a unique clinical case of a couple with three consecutive pregnancy losses and we aim to determine the genetic abnormalities causing the miscarriages. Conventional cytogenetic and molecular genetic analysis were performed on the products of conception as well as for the parents. Chromosomal analysis was performed based on the ISCN 2016 guidelines. This was followed by Chromosomal microarray analysis carried out using ISCA consortium probe set (8X60K). Genetic testing for the 1st product of conception was not performed. However, the 2nd and 3rd products of conception indicated an autosomal trisomy 22 and a 3.7 Mb deletion of 2p (cytoband p25.3) along with 13.6 Mb duplication of 16p (cytoband p13.3p13.12), respectively. -

DHX29 Antibody A
Revision 1 C 0 2 - t DHX29 Antibody a e r o t S Orders: 877-616-CELL (2355) [email protected] Support: 877-678-TECH (8324) 9 5 Web: [email protected] 1 www.cellsignal.com 4 # 3 Trask Lane Danvers Massachusetts 01923 USA For Research Use Only. Not For Use In Diagnostic Procedures. Applications: Reactivity: Sensitivity: MW (kDa): Source: UniProt ID: Entrez-Gene Id: WB, IP H M R Endogenous 155 Rabbit Q7Z478 54505 Product Usage Information Application Dilution Western Blotting 1:1000 Immunoprecipitation 1:50 Storage Supplied in 10 mM sodium HEPES (pH 7.5), 150 mM NaCl, 100 µg/ml BSA and 50% glycerol. Store at –20°C. Do not aliquot the antibody. Specificity / Sensitivity DHX29 Antibody detects endogenous levels of total DHX29 protein. Species Reactivity: Human, Mouse, Rat Source / Purification Polyclonal antibodies are produced by immunizing animals with a synthetic peptide corresponding to residues at the C-terminus of human DHX29. Antibodies were purified by protein A and peptide affinity chromatography. Background DHX29 is an ATP-dependent RNA helicase that belongs to the DEAD-box helicase family (DEAH subfamily). DHX29 contains one central helicase and one helicase at the carboxy-terminal domain (1). Its function has not been fully established but DHX29 was recently shown to facilitate translation initiation on mRNAs with structured 5' untranslated regions (2). DHX29 binds 40S subunits and hydrolyzes ATP, GTP, UTP, and CTP. Hydrolysis of nucleotide triphosphates by DHX29 is strongly stimulated by 43S complexes and is required for DHX29 activity in promoting 48S complex formation (2). 1. de la Cruz, J. -

ELP2 Antibody (C-Term) Purified Rabbit Polyclonal Antibody (Pab) Catalog # Ap2884b
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 ELP2 Antibody (C-term) Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP2884b Specification ELP2 Antibody (C-term) - Product Information Application IF, WB, FC,E Primary Accession Q6IA86 Reactivity Human Host Rabbit Clonality Polyclonal Isotype Rabbit Ig Calculated MW 92500 Antigen Region 737-765 ELP2 Antibody (C-term) - Additional Information Gene ID 55250 Other Names Elongator complex protein 2, ELP2, SHINC-2, STAT3-interacting protein 1, StIP1, Fluorescent confocal image of SY5Y cells ELP2, STATIP1 stained with ELP2 (C-term) antibody. SY5Y cells were fixed with 4% PFA (20 min), Target/Specificity permeabilized with Triton X-100 (0.2%, 30 This ELP2 antibody is generated from min). Cells were then incubated with rabbits immunized with a KLH conjugated AP2884b ELP2 (C-term) primary antibody synthetic peptide between 737-765 amino (1:100, 2 h at room temperature). For acids from the C-terminal region of human secondary antibody, Alexa Fluor® 488 ELP2. conjugated donkey anti-rabbit antibody (green) was used (1:1000, 1h). Nuclei were Dilution counterstained with Hoechst 33342 (blue) (10 IF~~1:100 WB~~1:1000 μg/ml, 5 min). Note the highly specific FC~~1:10~50 localization of the ELP2 immunosignal mainly to the cytoplasm, supported by Human Format Protein Atlas Data (http://www.proteinatlas.or Purified polyclonal antibody supplied in PBS g/ENSG00000134759). with 0.09% (W/V) sodium azide. This antibody is prepared by Saturated Ammonium Sulfate (SAS) precipitation followed by dialysis against PBS. Storage Maintain refrigerated at 2-8°C for up to 2 weeks.