Metabolic Diversity Involved in Biodegradation of 2-Nitroimidazole and 5-Nitroanthranilic Acid
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Downloaded 10/5/2021 9:43:05 AM
Chemical Science View Article Online EDGE ARTICLE View Journal | View Issue Aromatic side-chain flips orchestrate the conformational sampling of functional loops in Cite this: Chem. Sci.,2021,12,9318 † All publication charges for this article human histone deacetylase 8 have been paid for by the Royal Society a a bcd of Chemistry Vaibhav Kumar Shukla, ‡ Lucas Siemons, ‡ Francesco L. Gervasio and D. Flemming Hansen *a Human histone deacetylase 8 (HDAC8) is a key hydrolase in gene regulation and an important drug-target. High-resolution structures of HDAC8 in complex with substrates or inhibitors are available, which have provided insights into the bound state of HDAC8 and its function. Here, using long all-atom unbiased molecular dynamics simulations and Markov state modelling, we show a strong correlation between the conformation of aromatic side chains near the active site and opening and closing of the surrounding functional loops of HDAC8. We also investigated two mutants known to allosterically downregulate the enzymatic activity of HDAC8. Based on experimental data, we hypothesise that I19S-HDAC8 is unable to Received 6th April 2021 Creative Commons Attribution 3.0 Unported Licence. release the product, whereas both product release and substrate binding are impaired in the S39E- Accepted 27th May 2021 HDAC8 mutant. The presented results deliver detailed insights into the functional dynamics of HDAC8 DOI: 10.1039/d1sc01929e and provide a mechanism for the substantial downregulation caused by allosteric mutations, including rsc.li/chemical-science a disease causing one. Introduction II (HDAC-4, -5, -6, -7, -9, and HDAC10), and class III (SIRT1-7) have sequence similarity to yeast Rpd3, Hda1, and Sir2, Acetylation of lysine side chains occurs as a co-translation or respectively, whereas class IV (HDAC11) shares sequence simi- 2 This article is licensed under a post-translational modication of proteins and was rst iden- larity with both class I and II proteins. -

Gentaur Products List
Chapter 2 : Gentaur Products List • Rabbit Anti LAMR1 Polyclonal Antibody Cy5 Conjugated Conjugated • Rabbit Anti Podoplanin gp36 Polyclonal Antibody Cy5 • Rabbit Anti LAMR1 CT Polyclonal Antibody Cy5 • Rabbit Anti phospho NFKB p65 Ser536 Polyclonal Conjugated Conjugated Antibody Cy5 Conjugated • Rabbit Anti CHRNA7 Polyclonal Antibody Cy5 Conjugated • Rat Anti IAA Monoclonal Antibody Cy5 Conjugated • Rabbit Anti EV71 VP1 CT Polyclonal Antibody Cy5 • Rabbit Anti Connexin 40 Polyclonal Antibody Cy5 • Rabbit Anti IAA Indole 3 Acetic Acid Polyclonal Antibody Conjugated Conjugated Cy5 Conjugated • Rabbit Anti LHR CGR Polyclonal Antibody Cy5 Conjugated • Rabbit Anti Integrin beta 7 Polyclonal Antibody Cy5 • Rabbit Anti Natrexone Polyclonal Antibody Cy5 Conjugated • Rabbit Anti MMP 20 Polyclonal Antibody Cy5 Conjugated Conjugated • Rabbit Anti Melamine Polyclonal Antibody Cy5 Conjugated • Rabbit Anti BCHE NT Polyclonal Antibody Cy5 Conjugated • Rabbit Anti NAP1 NAP1L1 Polyclonal Antibody Cy5 • Rabbit Anti Acetyl p53 K382 Polyclonal Antibody Cy5 • Rabbit Anti BCHE CT Polyclonal Antibody Cy5 Conjugated Conjugated Conjugated • Rabbit Anti HPV16 E6 Polyclonal Antibody Cy5 Conjugated • Rabbit Anti CCP Polyclonal Antibody Cy5 Conjugated • Rabbit Anti JAK2 Polyclonal Antibody Cy5 Conjugated • Rabbit Anti HPV18 E6 Polyclonal Antibody Cy5 Conjugated • Rabbit Anti HDC Polyclonal Antibody Cy5 Conjugated • Rabbit Anti Microsporidia protien Polyclonal Antibody Cy5 • Rabbit Anti HPV16 E7 Polyclonal Antibody Cy5 Conjugated • Rabbit Anti Neurocan Polyclonal -

Precursor of Ether Phospholipids Is Synthesized by a Flavoenzyme
Precursor of ether phospholipids is synthesized by a flavoenzyme through covalent catalysis Simone Nencia, Valentina Pianoa, Sara Rosatib, Alessandro Alivertic, Vittorio Pandinic, Marco W. Fraaijed, Albert J. R. Heckb, Dale E. Edmondsone, and Andrea Mattevia,1 aDepartment of Biology and Biotechnology, University of Pavia, 27100 Pavia, Italy; bBiomolecular Mass Spectrometry and Proteomics Group, Bijvoet Center for Biomolecular Research and Utrecht Institute for Pharmaceutical Sciences, Utrecht University and Netherlands Proteomics Centre, 3584 CH Utrecht, The Netherlands; cDepartment of Biosciences, University of Milan, 20133 Milan, Italy; dMolecular Enzymology Group, University of Groningen, 9747 AG Groningen, The Netherlands; and eDepartment of Biochemistry, Emory University, Atlanta, GA 30322 Edited by Emil F. Pai, Ontario Cancer Institute/Princess Margaret Hospital, Toronto, ON, Canada, and accepted by the Editorial Board October 5, 2012 (received for review August 31, 2012) The precursor of the essential ether phospholipids is synthesized ADPS enzymes, whereas type 1 arises from mutations in PEX7, by a peroxisomal enzyme that uses a flavin cofactor to catalyze the protein mediating the peroxisomal import of ADPS (9–12). a reaction that does not alter the redox state of the substrates. Here we report a structural and mechanistic investigation of The enzyme crystal structure reveals a V-shaped active site with mammalian ADPS in WT and mutated forms. The biochemical a narrow constriction in front of the prosthetic group. Mutations hallmark of the enzyme is that it uses a redox cofactor, flavin causing inborn ether phospholipid deficiency, a very severe adenine dinucleotide (FAD), to catalyze a reaction that does not genetic disease, target residues that are part of the catalytic alter the redox state of the substrates (10, 13). -
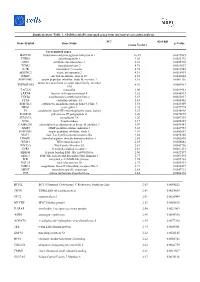
Supplementary Table 1. All Differentially Expressed Genes from Microarray Screening Analysis
Supplementary Table 1. All differentially expressed genes from microarray screening analysis. FCa (Gal-KD Gene Symbol Gene Name p-Value versus Vector) Up-regulated genes HAPLN1 hyaluronan and proteoglycan link protein 1 10.49 0.0027085 THBS1 thrombospondin 1 9.20 0.0022192 ODC1 ornithine decarboxylase 1 6.38 0.0055776 TGM2 transglutaminase 2 4.76 0.0015627 IL7R interleukin 7 receptor 4.75 0.0017245 SERINC2 serine incorporator 2 4.51 0.0014919 ITM2C integral membrane protein 2C 4.32 0.0044644 SERPINB7 serpin peptidase inhibitor, clade B, member 7 4.18 0.0081136 tumor necrosis factor receptor superfamily, member TNFRSF10D 4.01 0.0085561 10d TAGLN transgelin 3.90 0.0099963 LRRN4 leucine rich repeat neuronal 4 3.82 0.0046513 TGFB2 transforming growth factor beta 2 3.51 0.0035017 CPA4 carboxypeptidase A4 3.43 0.0008452 EPB41L3 erythrocyte membrane protein band 4.1-like 3 3.34 0.0025309 NRG1 neuregulin 1 3.28 0.0079724 F3 coagulation factor III (thromboplastin, tissue factor) 3.27 0.0038968 POLR3G polymerase III polypeptide G 3.26 0.0070675 SEMA7A semaphorin 7A 3.20 0.0087335 NT5E 5-nucleotidase 3.17 0.0036353 CAMK2N1 calmodulin-dependent protein kinase II inhibitor 1 3.07 0.0090141 TIMP3 TIMP metallopeptidase inhibitor 3 3.03 0.0047953 SERPINE1 serpin peptidase inhibitor, clade E 2.97 0.0053652 MALL mal, T-cell differentiation protein-like 2.88 0.0078205 DDAH1 dimethylarginine dimethylaminohydrolase 1 2.86 0.0002895 WDR3 WD repeat domain 3 2.85 0.0058842 WNT5A Wnt Family Member 5A 2.81 0.0043796 GPR1 G protein-coupled receptor 1 2.81 0.0021313 -

Phylogenetic Analysis, Subcellular Localization, and Expression
BMC Plant Biology BioMed Central Research article Open Access Phylogenetic analysis, subcellular localization, and expression patterns of RPD3/HDA1 family histone deacetylases in plants Malona V Alinsug, Chun-Wei Yu and Keqiang Wu* Address: Institute of Plant Biology, College of Life Science, National Taiwan University, Taipei, Taiwan Email: Malona V Alinsug - [email protected]; Chun-Wei Yu - [email protected]; Keqiang Wu* - [email protected] * Corresponding author Published: 28 March 2009 Received: 26 November 2008 Accepted: 28 March 2009 BMC Plant Biology 2009, 9:37 doi:10.1186/1471-2229-9-37 This article is available from: http://www.biomedcentral.com/1471-2229/9/37 © 2009 Alinsug et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: Although histone deacetylases from model organisms have been previously identified, there is no clear basis for the classification of histone deacetylases under the RPD3/ HDA1 superfamily, particularly on plants. Thus, this study aims to reconstruct a phylogenetic tree to determine evolutionary relationships between RPD3/HDA1 histone deacetylases from six different plants representing dicots with Arabidopsis thaliana, Populus trichocarpa, and Pinus taeda, monocots with Oryza sativa and Zea mays, and the lower plants with Physcomitrella patens. Results: Sixty two histone deacetylases of RPD3/HDA1 family from the six plant species were phylogenetically analyzed to determine corresponding orthologues. Three clusters were formed separating Class I, Class II, and Class IV. -

2010 Physical Biosciences Research Meeting
2010 Physical Biosciences Research Meeting Sheraton Inner Harbor Hotel Baltimore, MD October 17-20, 2010 Office of Basic Energy Sciences Chemical Sciences, Geosciences & Biosciences Division 2010 Physical Biosciences Research Meeting Program and Abstracts Sheraton Inner Harbor Hotel Baltimore, MD October 17-20, 2010 Chemical Sciences, Geosciences, and Biosciences Division Office of Basic Energy Sciences Office of Science U.S. Department of Energy i Cover art is taken from the public domain and can be found at: http://commons.wikimedia.org/wiki/File:Blue_crab_on_market_in_Piraeus_-_Callinectes_sapidus_Rathbun_20020819- 317.jpg This document was produced under contract number DE-AC05-060R23100 between the U.S. Department of Energy and Oak Ridge Associated Universities. The research grants and contracts described in this document are, unless specifically labeled otherwise, supported by the U.S. DOE Office of Science, Office of Basic Energy Sciences, Chemical Sciences, Geosciences, and Biosciences Division. ii Foreword This volume provides a record of the 2nd biennial meeting of the Principal Investigators (PIs) funded by the Physical Biosciences program, and is sponsored by the Chemical Sciences, Geosciences, and Biosciences Division of the Office of Basic Energy Sciences (BES) in the U.S. Department of Energy (DOE). Within DOE-BES there are two programs that fund basic research in energy-relevant biological sciences, Physical Biosciences and Photosynthetic Systems. These two Biosciences programs, along with a strong program in Solar Photochemistry, comprise the current Photo- and Bio- Chemistry Team. This meeting specifically brings together under one roof all of the PIs funded by the Physical Biosciences program, along with Program Managers and staff not only from DOE-BES, but also other offices within DOE, the national labs, and even other federal funding agencies. -

Biochemical Characterization and Comparison of Aspartylglucosaminidases Secreted in Venom of the Parasitoid Wasps Asobara Tabida and Leptopilina Heterotoma
RESEARCH ARTICLE Biochemical characterization and comparison of aspartylglucosaminidases secreted in venom of the parasitoid wasps Asobara tabida and Leptopilina heterotoma Quentin Coulette1, SeÂverine Lemauf2, Dominique Colinet2, Geneviève PreÂvost1, Caroline Anselme1, Marylène Poirie 2, Jean-Luc Gatti2* a1111111111 1 Unite ªEcologie et Dynamique des Systèmes AnthropiseÂsº (EDYSAN, FRE 3498 CNRS-UPJV), Universite de Picardie Jules Verne, Amiens, France, 2 Universite CoÃte d'Azur, INRA, CNRS, ISA, Sophia Antipolis, a1111111111 France a1111111111 a1111111111 * [email protected] a1111111111 Abstract Aspartylglucosaminidase (AGA) is a low-abundance intracellular enzyme that plays a key OPEN ACCESS role in the last stage of glycoproteins degradation, and whose deficiency leads to human Citation: Coulette Q, Lemauf S, Colinet D, PreÂvost aspartylglucosaminuria, a lysosomal storage disease. Surprisingly, high amounts of AGA- G, Anselme C, Poirie M, et al. (2017) Biochemical characterization and comparison of like proteins are secreted in the venom of two phylogenetically distant hymenopteran para- aspartylglucosaminidases secreted in venom of the sitoid wasp species, Asobara tabida (Braconidae) and Leptopilina heterotoma (Cynipidae). parasitoid wasps Asobara tabida and Leptopilina These venom AGAs have a similar domain organization as mammalian AGAs. They share heterotoma. PLoS ONE 12(7): e0181940. https:// with them key residues for autocatalysis and activity, and the mature - and -subunits also doi.org/10.1371/journal.pone.0181940 α β form an (αβ)2 structure in solution. Interestingly, only one of these AGAs subunits (α for Editor: Erjun Ling, Institute of Plant Physiology and AtAGA and for LhAGA) is glycosylated instead of the two subunits for lysosomal human Ecology Shanghai Institutes for Biological β Sciences, CHINA AGA (hAGA), and these glycosylations are partially resistant to PGNase F treatment. -

Lysine Acetylation: Elucidating the Components of an Emerging Global Signaling Pathway in Trypanosomes
Hindawi Publishing Corporation Journal of Biomedicine and Biotechnology Volume 2012, Article ID 452934, 16 pages doi:10.1155/2012/452934 Review Article Lysine Acetylation: Elucidating the Components of an Emerging Global Signaling Pathway in Trypanosomes Victoria Lucia Alonso1, 2 and Esteban Carlos Serra1, 2 1 Departamento de Microbiolog´ıa, Facultad de Ciencias Bioqu´ımicas y Farmac´euticas, Universidad Nacional de Rosario, Suipacha 531, Rosario 2000, Argentina 2 Instituto de Biolog´ıa Molecular y Celular de Rosario, CONICET-UNR, Suipacha 590, Rosario 2000, Argentina Correspondence should be addressed to Esteban Carlos Serra, [email protected] Received 17 April 2012; Revised 20 July 2012; Accepted 30 July 2012 Academic Editor: Andrea Silvana Ropolo´ Copyright © 2012 V. L. Alonso and E. C. Serra. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. In the past ten years the number of acetylated proteins reported in literature grew exponentially. Several authors have proposed that acetylation might be a key component in most eukaryotic signaling pathways, as important as phosphorylation. The enzymes involved in this process are starting to emerge; acetyltransferases and deacetylases are found inside and outside the nuclear compartment and have different regulatory functions. In trypanosomatids several of these enzymes have been described and are postulated to be novel antiparasitic targets for the rational design of drugs. In this paper we overview the most important known acetylated proteins and the advances made in the identification of new acetylated proteins using high-resolution mass spectrometry. -

Biosynthesis of Natural Products Containing Β-Amino Acids
Natural Product Reports Biosynthesis of natural products containing β -amino acids Journal: Natural Product Reports Manuscript ID: NP-REV-01-2014-000007.R1 Article Type: Review Article Date Submitted by the Author: 21-Apr-2014 Complete List of Authors: Kudo, Fumitaka; Tokyo Institute Of Technology, Department of Chemistry Miyanaga, Akimasa; Tokyo Institute Of Technology, Department of Chemistry Eguchi, T; Tokyo Institute Of Technology, Department of Chemistry and Materials Science Page 1 of 20 Natural Product Reports NPR RSC Publishing REVIEW Biosynthesis of natural products containing βββ- amino acids Cite this: DOI: 10.1039/x0xx00000x Fumitaka Kudo, a Akimasa Miyanaga, a and Tadashi Eguchi *b Received 00th January 2014, We focus here on β-amino acids as components of complex natural products because the presence of β-amino acids Accepted 00th January 2014 produces structural diversity in natural products and provides characteristic architectures beyond that of ordinary DOI: 10.1039/x0xx00000x α-L-amino acids, thus generating significant and unique biological functions in nature. In this review, we first survey the known bioactive β-amino acid-containing natural products including nonribosomal peptides, www.rsc.org/ macrolactam polyketides, and nucleoside-β-amino acid hybrids. Next, the biosynthetic enzymes that form β-amino acids from α-amino acids and de novo synthesis of β-amino acids are summarized. Then, the mechanisms of β- amino acid incorporation into natural products are reviewed. Because it is anticipated that the rational swapping of the β-amino acid moieties with various side chains and stereochemistries by biosynthetic engineering should lead to the creation of novel architectures and bioactive compounds, the accumulation of knowledge regarding β- amino acid-containing natural product biosynthetic machinery could have a significant impact in this field. -

Data-Driven Insights Into Ligands, Proteins, and Genetic Mutations
Data-Driven Insights into Ligands, Proteins, and Genetic Mutations by Jing Lu A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Bioinformatics) in the University of Michigan 2016 Doctoral Committee: Professor Heather A. Carlson, Chair Professor Charles L. Brooks III Assistant Professor Barry Grant Professor David S. Sept Professor Kerby A. Shedden © Jing Lu, 2016 Acknowledgements I would like to thank my advisor, Dr. Heather Carlson, for years of patient guidance, teaching, and support through the course of my PhD. I have learnt how to think critically and be rigorous in every step of research. I also want to express gratitude to my committee: Professor Charles L. Brooks III, Assistant Professor Barry Grant, Professor David S. Sept, Professor Kerby A. Shedden. Their advising is insightful and deepens my understanding of my research projects. I would like to thank Dr. Richard Smith for timely support for both my writing and research. For many Saturdays and Sundays, he promptly responds my requests for proofreading. Much of my work is built on his code in protein and ligand analysis. I would like to thank other members in Dr. Carlson’s lab for helping me with my work. Through the discussion with Dr. Jim Dunbar, I have learnt many critical ideas in Cheminformatics. Also, thank you to Sarah Graham and Jordan Clark for their tremendous friendship and willing to help with my writing. I would also thank previous members in Dr. Carlson’s lab. I would thank Dr. Phani Ghanakota for many late-night discussions and Dr. -

Active Site Tyrosine Is Essential for Amidohydrolase but Not for Esterase
Active site tyrosine is essential for amidohydrolase but not for esterase activity of a class 2 histone deacetylase-like bacterial enzyme Kristin Moreth, Daniel Riester, Christian Hildmann, René Hempel, Dennis Wegener, Andreas Schober, Andreas Schwienhorst To cite this version: Kristin Moreth, Daniel Riester, Christian Hildmann, René Hempel, Dennis Wegener, et al.. Ac- tive site tyrosine is essential for amidohydrolase but not for esterase activity of a class 2 histone deacetylase-like bacterial enzyme. Biochemical Journal, Portland Press, 2006, 401 (3), pp.659-665. 10.1042/BJ20061239. hal-00478649 HAL Id: hal-00478649 https://hal.archives-ouvertes.fr/hal-00478649 Submitted on 30 Apr 2010 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Biochemical Journal Immediate Publication. Published on 12 Oct 2006 as manuscript BJ20061239 Active site tyrosine is essential for amidohydrolase but not for esterase activity of a class 2 histone deacetylase-like bacterial enzyme Kristin Moreth¶, Daniel Riester¶, Christian Hildmann¶, René Hempel¶, Dennis Wegener¶,§,‡, -

Characterization of Aspartylglucosaminidase Activation and Aspartylglucosaminuria Mutations
Arto Pennanen Publications of the National Public Health Institute A 1 / 2004 — INDOOR AIR POLLUTION AND HEALTH RISKS IN FINNISH ICE ARENAS RISKSINFINNISHICE AND HEALTH AIR POLLUTION INDOOR Jani Saarela CHARACTERIZATION OF ASPARTYLGLUCOSAMINIDASE ACTIVATION AND ASPARTYLGLUCOSAMINURIA MUTATIONS ISBN 951-740-485-9 ISSN 0359-3584 ISBN 951-740-486-7 (pdf) Department of Molecular Medicine, ISSN 1458-6290 (pdf) National Public Health Institute, Helsinki, Finland and http://www.ktl.fi /portal/suomi/julkaisut/julkaisusarjat/ Department of Medical Genetics, kansanterveyslaitoksen_julkaisuja_a/ University of Helsinki, Finland Kopijyvä Kuopio 2005 Helsinki 2004 PPennanen_kansi.inddennanen_kansi.indd 1 117.2.20057.2.2005 115:26:195:26:19 CHARACTERIZATION OF ASPARTYLGLUCOSAMINIDASE ACTIVATION AND ASPARTYLGLUCOSAMINURIA MUTATIONS Jani Saarela Department of Molecular Medicine, National Public Health Institute, Helsinki, Finland and Department of Medical Genetics, University of Helsinki, Finland Academic Dissertation To be publicly discussed with the permission of the Medical Faculty of the University of Helsinki, in the lecture room 3 of Biomedicum Helsinki, Haartmaninkatu 8, Helsinki, on January 30th, 2004, at 12 o’clock noon. Helsinki 2004 Supervised by Professor Leena Peltonen-Palotie National Public Health Institute and Department of Medical Genetics University of Helsinki, Helsinki, Finland Reviewed by Professor Ole Kristian Tollersrud and Docent Marc Baumann Department of Medical Biochemistry Protein Chemistry/Proteomics Unit University of Tromsoe and Neuroscience Research Program Tromsoe, Norway University of Helsinki, Helsinki, Finland To be publicly discussed with Professor Marja Makarow Institute of Biotechnology and Department of Applied Biochemistry and Molecular Biology University of Helsinki, Helsinki, Finland Julkaisija-Utgivare-Publisher Kansanterveyslaitos (KTL) Mannerheimintie 166 00300 Helsinki puh. vaihde 09-47441, felefax 09-4744 8408 Folkhälsoinstitutet Mannerheimvägen 166 00300, Helsinki tel.