Systematics of the Hamamelidaceae Based on Morphological and Molecular Evidence Jianhua Li University of New Hampshire, Durham
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Vascular Plants of Massachusetts
The Vascular Plants of Massachusetts: The Vascular Plants of Massachusetts: A County Checklist • First Revision Melissa Dow Cullina, Bryan Connolly, Bruce Sorrie and Paul Somers Somers Bruce Sorrie and Paul Connolly, Bryan Cullina, Melissa Dow Revision • First A County Checklist Plants of Massachusetts: Vascular The A County Checklist First Revision Melissa Dow Cullina, Bryan Connolly, Bruce Sorrie and Paul Somers Massachusetts Natural Heritage & Endangered Species Program Massachusetts Division of Fisheries and Wildlife Natural Heritage & Endangered Species Program The Natural Heritage & Endangered Species Program (NHESP), part of the Massachusetts Division of Fisheries and Wildlife, is one of the programs forming the Natural Heritage network. NHESP is responsible for the conservation and protection of hundreds of species that are not hunted, fished, trapped, or commercially harvested in the state. The Program's highest priority is protecting the 176 species of vertebrate and invertebrate animals and 259 species of native plants that are officially listed as Endangered, Threatened or of Special Concern in Massachusetts. Endangered species conservation in Massachusetts depends on you! A major source of funding for the protection of rare and endangered species comes from voluntary donations on state income tax forms. Contributions go to the Natural Heritage & Endangered Species Fund, which provides a portion of the operating budget for the Natural Heritage & Endangered Species Program. NHESP protects rare species through biological inventory, -

Siebold and Zuccarini's Type Specimens and Original Materials
J. Jpn. Bot. 89: 209–229 (2014) Siebold and Zuccarini’s Type Specimens and Original Materials from Japan, Part 4. Angiosperms. Dicotyledoneae 3 a, b c Shinobu AKIYAMA *, Gerard THIJSSE , Hans-Joachim ESSER d and Hideaki OHBA aDepartment of Botany, National Museum of Nature and Science, 4-1-1, Amakubo, Tsukuba, 305-0005 JAPAN; bNationaal Herbarium Nederland, Naturalis Biodiversity Center, P. O. Box 9514, THE NETHERLANDS; cBotanische Staatssammlung München, Menzinger Straße 67, D-80638 München, GERMANY; dDepartment of Botany, the University Museum, the University of Tokyo, 7-3-1, Hongo, Tokyo, 113-0033 JAPAN *Corresponding author: [email protected] (Accepted on March 1, 2014) Part 4 of the list of type specimens and original materials of the taxa described by Siebold and Zuccarini from Japan concerns the taxa belonging to Piperaceae to Hamamelidaceae, according to the 12th Engler system by Melchior. (Continued from J. Jpn. Bot. 89: 76–102, 2014) Key words: Japanese flora, lectotypification, Siebold collections, Zuccarini. This paper is the part 4 of the list of type Original material: In Japonia legt. Bürger, specimens and original materials of the taxa Herb. Zuccarinii (M0153560) [Category 2]. In described by Siebold and Zuccarini from Japonia legt. Bürger comct. d. Siebold, Herb. Japan (Akiyama et al. 2012, 2013, 2014), and Zuccarinii (M0153559) [Category 2b]. Japonia. treats the taxa belonging to Piperaceae to [Siebold s.n.] (MAK S1676) [Category 2]. Hamamelidaceae according to the 12th Engler system by Melchior. The method, especially the Chloranthaceae evaluation principles of category of the original Chloranthus Sw. materials, is the same as explained in part 1. -

Fossil Evidence of Initial Radiation of Cercidiphyllaceae L
УДК 561.46:551.763 Палеоботаника, 2018, Т. 9, C. 54—75 https://doi.org/10.31111/palaeobotany/2018.9.54 Palaeobotany, 2018, Vol. 9, P. 54—75 FOSSIL EVIDENCE OF INITIAL RADIATION OF CERCIDIPHYLLACEAE L. B. Golovneva, A. A. Zolina Komarov Botanical Institute RAS, St. Petersburg, [email protected] Abstract. Cercidiphyllaceae-like leaves and fruits from the Lower Cretaceous deposits of Northeastern Asia were restudied. In the result one species of Jenkinsella fruits and fi ve species of Trochodendroides leaves were recognized, including Trochodendroides potomacensis (Ward) Bell, T. buorensis Golovneva, T. sittensis Golovneva, sp. nov., T. vachrameeviana (Iljinskaja) Golovneva, comb. nov., and T. denticulata (Budantsev et Kiritchkova) Golovneva, comb. nov. Two new combinations and one new species are published. These plants had very small leaves and probably were shrubs. Fruits of Nyssidium orientale Samylina from the Barremian-Aptian Starosuchan Formation (Primorye, Russia) have no follicular characters as Jenkinsella fruits. Their affi nity, not only to Cercidiphyllum-like plants, but to angiosperms in general, is doubtful. Leaves and fruits of Cercidiphyllum sujfunense Krassilov from the lower-middle Albian Galenki Formation (Primorye) also can not be assigned to Cercidiphyllaceae. Leaves have pinnate, brochidodromous venation and are comparable with those of Asiatifolium elegans Sun, Guo et Zheng, which were recorded from the Frentsevka Formation of the Partizansk coal basin, Primorye, Russia, and from the Chengzihe Formation, Northeastern China. Thus, the fi rst reliable records of the genus Trochodendroides appear in the early-middle Albian. The relationship of these leaves with Cercidiphyllaceae is confi rmed by fi nds of associated fruits Jenkinsella fi la- tovii and by signifi cant diversity of Trochodendroides in the Late Albian-Cenomanian. -

Integrating Palaeontological and Molecular Data Uncovers Multiple
Integrating palaeontological and molecular data uncovers multiple ancient and recent dispersals in the pantropical Hamamelidaceae Xiaoguo Xiang, Kunli Xiang, Rosa del C. Ortiz, Florian Jabbour, Wei Wang To cite this version: Xiaoguo Xiang, Kunli Xiang, Rosa del C. Ortiz, Florian Jabbour, Wei Wang. Integrating palaeontolog- ical and molecular data uncovers multiple ancient and recent dispersals in the pantropical Hamamel- idaceae. Journal of Biogeography, Wiley, 2019, 46 (11), pp.2622-2631. 10.1111/jbi.13690. hal- 02612865 HAL Id: hal-02612865 https://hal.archives-ouvertes.fr/hal-02612865 Submitted on 19 May 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Integrating palaeontological and molecular data uncovers multiple ancient and recent dispersals in the pantropical Hamamelidaceae Xiaoguo Xiang1,2, Kunli Xiang1,3, Rosa Del C. Ortiz4, Florian Jabbour5, Wei Wang1,3 1State Key Laboratory of Systematic and Evolutionary Botany, Institute of Botany, Chinese Academy of Sciences, Beijing, China 2Jiangxi Province Key Laboratory of Watershed Ecosystem -
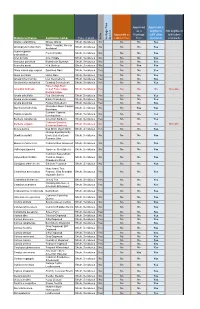
Botanical Name Common Name
Approved Approved & as a eligible to Not eligible to Approved as Frontage fulfill other fulfill other Type of plant a Street Tree Tree standards standards Heritage Tree Tree Heritage Species Botanical Name Common name Native Abelia x grandiflora Glossy Abelia Shrub, Deciduous No No No Yes White Forsytha; Korean Abeliophyllum distichum Shrub, Deciduous No No No Yes Abelialeaf Acanthropanax Fiveleaf Aralia Shrub, Deciduous No No No Yes sieboldianus Acer ginnala Amur Maple Shrub, Deciduous No No No Yes Aesculus parviflora Bottlebrush Buckeye Shrub, Deciduous No No No Yes Aesculus pavia Red Buckeye Shrub, Deciduous No No Yes Yes Alnus incana ssp. rugosa Speckled Alder Shrub, Deciduous Yes No No Yes Alnus serrulata Hazel Alder Shrub, Deciduous Yes No No Yes Amelanchier humilis Low Serviceberry Shrub, Deciduous Yes No No Yes Amelanchier stolonifera Running Serviceberry Shrub, Deciduous Yes No No Yes False Indigo Bush; Amorpha fruticosa Desert False Indigo; Shrub, Deciduous Yes No No No Not eligible Bastard Indigo Aronia arbutifolia Red Chokeberry Shrub, Deciduous Yes No No Yes Aronia melanocarpa Black Chokeberry Shrub, Deciduous Yes No No Yes Aronia prunifolia Purple Chokeberry Shrub, Deciduous Yes No No Yes Groundsel-Bush; Eastern Baccharis halimifolia Shrub, Deciduous No No Yes Yes Baccharis Summer Cypress; Bassia scoparia Shrub, Deciduous No No No Yes Burning-Bush Berberis canadensis American Barberry Shrub, Deciduous Yes No No Yes Common Barberry; Berberis vulgaris Shrub, Deciduous No No No No Not eligible European Barberry Betula pumila -

Number 3, Spring 1998 Director’S Letter
Planning and planting for a better world Friends of the JC Raulston Arboretum Newsletter Number 3, Spring 1998 Director’s Letter Spring greetings from the JC Raulston Arboretum! This garden- ing season is in full swing, and the Arboretum is the place to be. Emergence is the word! Flowers and foliage are emerging every- where. We had a magnificent late winter and early spring. The Cornus mas ‘Spring Glow’ located in the paradise garden was exquisite this year. The bright yellow flowers are bright and persistent, and the Students from a Wake Tech Community College Photography Class find exfoliating bark and attractive habit plenty to photograph on a February day in the Arboretum. make it a winner. It’s no wonder that JC was so excited about this done soon. Make sure you check of themselves than is expected to seedling selection from the field out many of the special gardens in keep things moving forward. I, for nursery. We are looking to propa- the Arboretum. Our volunteer one, am thankful for each and every gate numerous plants this spring in curators are busy planting and one of them. hopes of getting it into the trade. preparing those gardens for The magnolias were looking another season. Many thanks to all Lastly, when you visit the garden I fantastic until we had three days in our volunteers who work so very would challenge you to find the a row of temperatures in the low hard in the garden. It shows! Euscaphis japonicus. We had a twenties. There was plenty of Another reminder — from April to beautiful seven-foot specimen tree damage to open flowers, but the October, on Sunday’s at 2:00 p.m. -

Seeds of Woody Plants
ARNOLDIA A continuation of the BULLETIN OF POPULAR INFORMATION of the Arnold Arboretum, Harvard University VOLUME 133 SEPTEMBER 11, 19533 NuMSERS 7-9 SEEDS OF WOODY PLANTS Collecting - Cleaning - Shipping - Longevity studies of seeds of plants, their collection, care and requirements for THEgermination, has been an intriguing one over the years. Much work has been done, mostly with the seeds of cereals, annuals and trees used for reforestation. Text books on forestry have much detailed information concerning forest tree seeds, but the seeds of many woody ornamental plants have not been studied as thoroughly as many a commercial grower would wish. This bulletin has to deal with such seeds, primarily with suggestions for their collection, care in storage, and shipping before they are sown. This information is taken from many sources. not the least of which are the first hand experiences here at the Arnold Arbore- tum where the propagators have been sowing such seeds for over 75 years. The best reference on the subject with an excellent bibliography, is "Woody Plant Seed Manual" written by the Forest Service, U.S. Department of Agriculture (Misc. Pub. No. 654, issued June 1948). Collecting Seeds should not be collected until they are ripe, for in many cases, seed col- lected prematurely may not germinate. Any collector who has had experience in this field knows that there are various methods of determining the ripeness of the fruit - the shrivelling of the pod or the cord connecting the seed to the pod in the legumes, the color or softness of the pulp of the fruit of Malus, Viburnum and Cotoneaster, and close examination of the cones of coniferous trees sometimes even tested by determining the specific gravity of the unopened cones. -

Gardenergardener®
Theh American A n GARDENERGARDENER® The Magazine of the AAmerican Horticultural Societyy January / February 2016 New Plants for 2016 Broadleaved Evergreens for Small Gardens The Dwarf Tomato Project Grow Your Own Gourmet Mushrooms contents Volume 95, Number 1 . January / February 2016 FEATURES DEPARTMENTS 5 NOTES FROM RIVER FARM 6 MEMBERS’ FORUM 8 NEWS FROM THE AHS 2016 Seed Exchange catalog now available, upcoming travel destinations, registration open for America in Bloom beautifi cation contest, 70th annual Colonial Williamsburg Garden Symposium in April. 11 AHS MEMBERS MAKING A DIFFERENCE Dale Sievert. 40 HOMEGROWN HARVEST Love those leeks! page 400 42 GARDEN SOLUTIONS Understanding mycorrhizal fungi. BOOK REVIEWS page 18 44 The Seed Garden and Rescuing Eden. Special focus: Wild 12 NEW PLANTS FOR 2016 BY CHARLOTTE GERMANE gardening. From annuals and perennials to shrubs, vines, and vegetables, see which of this year’s introductions are worth trying in your garden. 46 GARDENER’S NOTEBOOK Link discovered between soil fungi and monarch 18 THE DWARF TOMATO PROJECT BY CRAIG LEHOULLIER butterfl y health, stinky A worldwide collaborative breeds diminutive plants that produce seeds trick dung beetles into dispersal role, regular-size, fl avorful tomatoes. Mt. Cuba tickseed trial results, researchers unravel how plants can survive extreme drought, grant for nascent public garden in 24 BEST SMALL BROADLEAVED EVERGREENS Delaware, Lady Bird Johnson Wildfl ower BY ANDREW BUNTING Center selects new president and CEO. These small to mid-size selections make a big impact in modest landscapes. 50 GREEN GARAGE Seed-starting products. 30 WEESIE SMITH BY ALLEN BUSH 52 TRAVELER’S GUIDE TO GARDENS Alabama gardener Weesie Smith championed pagepage 3030 Quarryhill Botanical Garden, California. -

Annual Benefit Plant Sale 2012
Annual Benefit Plant Sale 2012 Botanic Gardens COLLEGE OF AGRICULTURE & NATURAL RESOURCES Connect to nature Get inspired by wildflowers, naturalistic gardening and meadows in a whole new way with our seasonal garden tours. Enjoy an art class in the garden or learn about native plant gardening, conservation, and habitats by taking one of our classes. And if you can’t visit us, enroll in our new online distance learning program, Mt. Cuba Center Connect. Visit www.mtcubacenter.org to reserve a tour or sign up for a class. Two-Hour Guided Tours | $5 per person Spring Wildflower Tours April 12th – May 27th Summer Twilight Tours May 30th – July 26th 8th Annual Wildflower Celebration |Free th April 29 , 10am – 4pm Purple pitcherplant (Sarracenia purpurea) Greenville, DE P: 302.239.4244 www.mtcubacenter.org INSPIRATION x EDUCATION x CONSERVATION 2 2012 SPRING PLANT SALE CATALOG WEBSITE: http://ag.udel.edu/udbg/events/annualsale.html WELCOME We welcome you to the twentieth annual UDBG benefit plant sale. In addition to its role as the major source of funding for the UDBG, 2012 BENEFIT PLANT SALE CATALOG we hope it also serves as a major educational event for our members and the public. It presents an opportunity to learn about new plants and consider possibilities. We should always look for ways to expand and improve our knowledge about plants and this catalog offers possibilities to accomplish both. As always, we offer an in- depth look at a particular group of plants, this year the genus Camellia. The selection goes beyond offering various cultivars with differing flower color, to a more extensive exploration of the genus with particular emphasis on hardy selections and new hybrids Camellia ‘Autumn Spirit’. -

International Organisation of Palaeobotany IOP NEWSLETTER
IOP 107 September 2015 INTERNATIONAL UNION OF BIOLOGICAL SCIENCES SECTION FOR PALAEOBOTANY International Organisation of Palaeobotany IOP NEWSLETTER 107 September 2015 CONTENTS FROM THE SECRETARY/TREASURER IPC XIV/IOPC X 2016 MEETING REPORT OBITUARY BOOK REVIEW UPCOMING MEETINGS CALL FOR NEWS and NOTES The views expressed in the newsletter are those of its correspondents, and do not necessarily reflect the policy of IOP. Please send us your contributions for the next edition of our newsletter (December 2015) by November 30th, 2015. President: Johanna Eder-Kovar (Germany) Vice Presidents: Bob Spicer (Great Britain), Harufumi Nishida (Japan), Mihai Popa (Romania) Members at Large: Jun Wang (China), Hans Kerp (Germany), Alexej Herman (Russia) Secretary/Treasurer/Newsletter editor: Mike Dunn (USA) Conference/Congress Chair: Francisco de Assis Ribeiro dos Santos IOP Logo: The evolution of plant architecture (© by A. R. Hemsley) IOP 107 1 September 2015 FROM THE and also, who you want to run your organization. A call for nominees to the SECRETARY/TREASURER Executive Council will go out in the December Newsletter. Dear International Organisation of Palaeobotany Members, Please feel free to contact me with questions, comments, or any information Please accept this July-ish newsletter. you would like passed on to the Membership. I can be reached at: Thanks to everyone who submitted items for the Newsletter. I really appreciate the Mike Dunn support of those who sent items in. The Department of Biological Sciences International Organisation of Palaeobotany Cameron University is a Member-Centric Organization, and this Lawton, Oklahoma 73505 Newsletter is an example of how great and Ph.: 580-581-2287 meaningful we can be when the membership email: [email protected] participates. -

Open As a Single Document
The Witch Hazel Family (Hamamelidaceae) by RICHARD E. WEAVER, JR. The Arnold Arboretum has claimed that there is a tree or shrub in bloom every month of the year on its grounds in Jamaica Plain, Mass. In many years this assertion is true, but only because of a single genus of plants, Hamamelis, the Witch Hazels. As December ar- rives, the last pale yellow flowers begin to fade on the Common Witch Hazel, H. virginiana, a native of the eastern United States. And with the New Year come the fragrant, bronzy flowers of the Vernal Witch Hazel, H. vernalis, closely followed by the bright and beautiful yellow blossoms of the Chinese species, H. mollis. The Witch Hazels belong to the Hamamelidaceae, a family of plants which are mostly neglected by the American horticultural public. Admittedly, the family is insignificant horticulturally com- pared with some others, for example, the Rose Family, as a result of the diversity and sheer number of its genera and species, and the Magnolia Family, because of the universal appeal of some of its members. But a high percentage of the species in the Hamamelida- ceae are first class ornamentals, possessing charm, beautiful and often fragrant flowers, unusual blooming times, and brilliant au- tumnal coloration. Most also are easy to grow if the soil is light and loamy, and they bloom reasonably well in partial shade. In addition, they are not bothered by any serious diseases or insect pests. This article will present a brief discussion of all of the genera which are hardy in the northern half of the United States. -

Endemic Wild Ornamental Plants from Northwestern Yunnan, China
HORTSCIENCE 40(6):1612–1619. 2005. have played an important role in world horti- culture and have been introduced to Western countries where they have been widely cul- Endemic Wild Ornamental Plants tivated. Some of the best known examples include Rhododendron, Primula, Gentiana, from Northwestern Yunnan, China Pedicularis, and Saussurea, which are all im- 1 portant genera in northwestern Yunnan (Chen Xiao-Xian Li and Zhe-Kun Zhou et al., 1989; Feng, 1983; Guan et al., 1998; Hu, Kunming Institute of Botany, Chinese Academy of Sciences, Kunming, P.R. 1990; Shi and Jin, 1999; Yang, 1956;). Many of China 650204 these ornamental species are endemic to small areas of northwestern Yunnan (e.g., Rhododen- Additional index words. horticultural potential dron russatum), therefore, their cultivation not Abstract. Northwestern Yunnan is situated in the southern part of the Hengduan Mountains, only provides for potential sources of income which is a complex and varied natural environment. Consequently, this region supports a generation, but also offers a potential form of great diversity of endemic plants. Using fi eld investigation in combination with analysis conservation management: these plants can of relevant literature and available data, this paper presents a regional ethnobotanical be used directly for their ornamental plant study of this area. Results indicated that northwestern Yunnan has an abundance of wild value or as genetic resources for plant breed- ornamental plants: this study identifi ed 262 endemic species (belonging to 64 genera and ing programs. The aims of current paper are 28 families) with potential ornamental value. The distinguishing features of these wild to describe the unique fl ora of northwestern plants, their characteristics and habitats are analyzed; the ornamental potential of most Yunnan and provide detailed information of plants stems from their wildfl owers, but some species also have ornamental fruits and those resources, in terms of their potential foliage.