PRKAR1A and Thyroid Tumors
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplemental Information to Mammadova-Bach Et Al., “Laminin Α1 Orchestrates VEGFA Functions in the Ecosystem of Colorectal Carcinogenesis”
Supplemental information to Mammadova-Bach et al., “Laminin α1 orchestrates VEGFA functions in the ecosystem of colorectal carcinogenesis” Supplemental material and methods Cloning of the villin-LMα1 vector The plasmid pBS-villin-promoter containing the 3.5 Kb of the murine villin promoter, the first non coding exon, 5.5 kb of the first intron and 15 nucleotides of the second villin exon, was generated by S. Robine (Institut Curie, Paris, France). The EcoRI site in the multi cloning site was destroyed by fill in ligation with T4 polymerase according to the manufacturer`s instructions (New England Biolabs, Ozyme, Saint Quentin en Yvelines, France). Site directed mutagenesis (GeneEditor in vitro Site-Directed Mutagenesis system, Promega, Charbonnières-les-Bains, France) was then used to introduce a BsiWI site before the start codon of the villin coding sequence using the 5’ phosphorylated primer: 5’CCTTCTCCTCTAGGCTCGCGTACGATGACGTCGGACTTGCGG3’. A double strand annealed oligonucleotide, 5’GGCCGGACGCGTGAATTCGTCGACGC3’ and 5’GGCCGCGTCGACGAATTCACGC GTCC3’ containing restriction site for MluI, EcoRI and SalI were inserted in the NotI site (present in the multi cloning site), generating the plasmid pBS-villin-promoter-MES. The SV40 polyA region of the pEGFP plasmid (Clontech, Ozyme, Saint Quentin Yvelines, France) was amplified by PCR using primers 5’GGCGCCTCTAGATCATAATCAGCCATA3’ and 5’GGCGCCCTTAAGATACATTGATGAGTT3’ before subcloning into the pGEMTeasy vector (Promega, Charbonnières-les-Bains, France). After EcoRI digestion, the SV40 polyA fragment was purified with the NucleoSpin Extract II kit (Machery-Nagel, Hoerdt, France) and then subcloned into the EcoRI site of the plasmid pBS-villin-promoter-MES. Site directed mutagenesis was used to introduce a BsiWI site (5’ phosphorylated AGCGCAGGGAGCGGCGGCCGTACGATGCGCGGCAGCGGCACG3’) before the initiation codon and a MluI site (5’ phosphorylated 1 CCCGGGCCTGAGCCCTAAACGCGTGCCAGCCTCTGCCCTTGG3’) after the stop codon in the full length cDNA coding for the mouse LMα1 in the pCIS vector (kindly provided by P. -

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -

Tumour-Agnostic Therapy for Pancreatic Cancer and Biliary Tract Cancer
diagnostics Review Tumour-Agnostic Therapy for Pancreatic Cancer and Biliary Tract Cancer Shunsuke Kato Department of Clinical Oncology, Juntendo University Graduate School of Medicine, 2-1-1, Hongo, Bunkyo-ku, Tokyo 113-8421, Japan; [email protected]; Tel.: +81-3-5802-1543 Abstract: The prognosis of patients with solid tumours has remarkably improved with the develop- ment of molecular-targeted drugs and immune checkpoint inhibitors. However, the improvements in the prognosis of pancreatic cancer and biliary tract cancer is delayed compared to other carcinomas, and the 5-year survival rates of distal-stage disease are approximately 10 and 20%, respectively. How- ever, a comprehensive analysis of tumour cells using The Cancer Genome Atlas (TCGA) project has led to the identification of various driver mutations. Evidently, few mutations exist across organs, and basket trials targeting driver mutations regardless of the primary organ are being actively conducted. Such basket trials not only focus on the gate keeper-type oncogene mutations, such as HER2 and BRAF, but also focus on the caretaker-type tumour suppressor genes, such as BRCA1/2, mismatch repair-related genes, which cause hereditary cancer syndrome. As oncogene panel testing is a vital approach in routine practice, clinicians should devise a strategy for improved understanding of the cancer genome. Here, the gene mutation profiles of pancreatic cancer and biliary tract cancer have been outlined and the current status of tumour-agnostic therapy in these cancers has been reported. Keywords: pancreatic cancer; biliary tract cancer; targeted therapy; solid tumours; driver mutations; agonist therapy Citation: Kato, S. Tumour-Agnostic Therapy for Pancreatic Cancer and 1. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

To View the ESE Recommended Curriculum of Specialisation in Clinical Endocrinology, Diabetes and Metabolism
European Society of Endocrinology Recommended Curriculum of Specialisation in Clinical Endocrinology, Diabetes and Metabolism Version 2, November 2019 Contents Endorsement ........................................................................................................................................ 2 Introduction .......................................................................................................................................... 3 1. Diabetes mellitus .............................................................................................................................. 4 2. Lipid disorders ................................................................................................................................... 5 3. Obesity and bariatric endocrinology ................................................................................................. 5 4. Pituitary ............................................................................................................................................ 5 5. Thyroid .............................................................................................................................................. 6 6. Parathyroid, calcium and bone ......................................................................................................... 7 7. Adrenal ............................................................................................................................................. 8 8. Reproductive endocrinology and sexual function -

RD-Action Matchmaker – Summary of Disease Expertise Recorded Under
Summary of disease expertise recorded via RD-ACTION Matchmaker under each Thematic Grouping and EURORDIS Members’ Thematic Grouping Thematic Reported expertise of those completing the EURORDIS Member perspectives on Grouping matchmaker under each heading Grouping RD Thematically Rare Bone Achondroplasia/Hypochondroplasia Achondroplasia Amelia skeletal dysplasia’s including Achondroplasia/Growth hormone cleidocranial dysostosis, arthrogryposis deficiency/MPS/Turner Brachydactyly chondrodysplasia punctate Fibrous dysplasia of bone Collagenopathy and oncologic disease such as Fibrodysplasia ossificans progressive Li-Fraumeni syndrome Osteogenesis imperfecta Congenital hand and fore-foot conditions Sterno Costo Clavicular Hyperostosis Disorders of Sex Development Duchenne Muscular Dystrophy Ehlers –Danlos syndrome Fibrodysplasia Ossificans Progressiva Growth disorders Hypoparathyroidism Hypophosphatemic rickets & Nutritional Rickets Hypophosphatasia Jeune’s syndrome Limb reduction defects Madelung disease Metabolic Osteoporosis Multiple Hereditary Exostoses Osteogenesis imperfecta Osteoporosis Paediatric Osteoporosis Paget’s disease Phocomelia Pseudohypoparathyroidism Radial dysplasia Skeletal dysplasia Thanatophoric dwarfism Ulna dysplasia Rare Cancer and Adrenocortical tumours Acute monoblastic leukaemia Tumours Carcinoid tumours Brain tumour Craniopharyngioma Colon cancer, familial nonpolyposis Embryonal tumours of CNS Craniopharyngioma Ependymoma Desmoid disease Epithelial thymic tumours in -

Multiple Orbital Neurofibromas, Painful Peripheral Nerve Tumors, Distinctive
European Journal of Human Genetics (2012) 20, 618–625 & 2012 Macmillan Publishers Limited All rights reserved 1018-4813/12 www.nature.com/ejhg ARTICLE Multiple orbital neurofibromas, painful peripheral nerve tumors, distinctive face and marfanoid habitus: a new syndrome D Babovic-Vuksanovic*,1, Ludwine Messiaen2, Christoph Nagel3, Hilde Brems4, Bernd Scheithauer5, Ellen Denayer4, Rong Mao6, Raf Sciot7, Karen M Janowski2, Martin U Schuhmann3, Kathleen Claes8, Eline Beert4, James A Garrity9, Robert J Spinner10, Anat Stemmer-Rachamimov11, Ralitza Gavrilova1, Frank Van Calenbergh12, Victor Mautner13 and Eric Legius4 Four unrelated patients having an unusual clinical phenotype, including multiple peripheral nerve sheath tumors, are reported. Their clinical features were not typical of any known familial tumor syndrome. The patients had multiple painful neurofibromas, including bilateral orbital plexiform neurofibromas, and spinal as well as mucosal neurofibromas. In addition, they exhibited a marfanoid habitus, shared similar facial features, and had enlarged corneal nerves as well as neuronal migration defects. Comprehensive NF1, NF2 and SMARCB1 mutation analyses revealed no mutation in blood lymphocytes and in schwann cells cultured from plexiform neurofibromas. Furthermore, no mutations in RET, PRKAR1A, PTEN and other RAS-pathway genes were found in blood leukocytes. Collectively, the clinical and pathological findings in these four cases fit no known syndrome and likely represent a new disorder. European Journal of Human Genetics (2012) 20, 618–625; doi:10.1038/ejhg.2011.275; published online 18 January 2012 Keywords: orbital neurofibroma; plexiform neurofibroma; enlarged corneal nerves; marfanoid habitus INTRODUCTION Ruvalcaba syndrome, a part of the PHTS spectrum, develop cafe´- Peripheral nerve sheath tumors, specifically neurofibromas and au-lait spots and macules of the glans penis. -

DNAJB1–PRKACA Fusion Kinase Interacts with Β-Catenin and the Liver
DNAJB1–PRKACA fusion kinase interacts with INAUGURAL ARTICLE β-catenin and the liver regenerative response to drive fibrolamellar hepatocellular carcinoma Edward R. Kastenhubera,b, Gadi Lalazarc, Shauna L. Houlihana, Darjus F. Tschaharganehd,e, Timour Baslana, Chi-Chao Chena, David Requenac, Sha Tiana, Benedikt Bosbachf, John E. Wilkinsong, Sanford M. Simonc, and Scott W. Lowea,h,1 aDepartment of Cancer Biology and Genetics, Memorial Sloan Kettering Cancer Center, New York, NY 10065; bLouis V. Gerstner Jr. Graduate School of Biomedical Sciences, Memorial Sloan Kettering Cancer Center, New York, NY 10065; cLaboratory of Cellular Biophysics, The Rockefeller University, New York, NY 10065; dHelmholtz University Group “Cell Plasticity and Epigenetic Remodeling,” German Cancer Research Center (DKFZ), 69120 Heidelberg, Germany; eInstitute of Pathology, University Hospital, 69120 Heidelberg, Germany; fOncology Target Discovery Program, Pfizer Inc., Pearl River, NY 10965; gDepartment of Pathology, University of Michigan School of Medicine, Ann Arbor, MI 48109; and hHoward Hughes Medical Institute, New York, NY 10065 This contribution is part of the special series of Inaugural Articles by members of the National Academy of Sciences elected in 2017. Contributed by Scott W. Lowe, October 26, 2017 (sent for review September 22, 2017; reviewed by Nabeel M. Bardeesy and David A. Largaespada) A segmental deletion resulting in DNAJB1–PRKACA gene fusion is any known etiological risk factors such as alcoholism, chronic hep- now recognized as the signature genetic event of fibrolamellar hepa- atitis infection, or liver flukes (8). tocellular carcinoma (FL-HCC), a rare but lethal liver cancer that pri- Currently, FL-HCC is diagnosed on the basis of histological marily affects adolescents and young adults. -
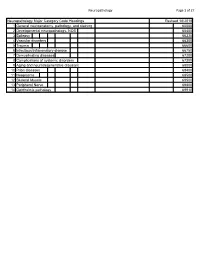
Neuropathology Category Code List
Neuropathology Page 1 of 27 Neuropathology Major Category Code Headings Revised 10/2018 1 General neuroanatomy, pathology, and staining 65000 2 Developmental neuropathology, NOS 65400 3 Epilepsy 66230 4 Vascular disorders 66300 5 Trauma 66600 6 Infectious/inflammatory disease 66750 7 Demyelinating diseases 67200 8 Complications of systemic disorders 67300 9 Aging and neurodegenerative diseases 68000 10 Prion diseases 68400 11 Neoplasms 68500 12 Skeletal Muscle 69500 13 Peripheral Nerve 69800 14 Ophthalmic pathology 69910 Neuropathology Page 2 of 27 Neuropathology 1 General neuroanatomy, pathology, and staining 65000 A Neuroanatomy, NOS 65010 1 Neocortex 65011 2 White matter 65012 3 Entorhinal cortex/hippocampus 65013 4 Deep (basal) nuclei 65014 5 Brain stem 65015 6 Cerebellum 65016 7 Spinal cord 65017 8 Pituitary 65018 9 Pineal 65019 10 Tracts 65020 11 Vascular supply 65021 12 Notochord 65022 B Cell types 65030 1 Neurons 65031 2 Astrocytes 65032 3 Oligodendroglia 65033 4 Ependyma 65034 5 Microglia and mononuclear cells 65035 6 Choroid plexus 65036 7 Meninges 65037 8 Blood vessels 65038 C Cerebrospinal fluid 65045 D Pathologic responses in neurons and axons 65050 1 Axonal degeneration/spheroid/reaction 65051 2 Central chromatolysis 65052 3 Tract degeneration 65053 4 Swollen/ballooned neurons 65054 5 Trans-synaptic neuronal degeneration 65055 6 Olivary hypertrophy 65056 7 Acute ischemic (hypoxic) cell change 65057 8 Apoptosis 65058 9 Protein aggregation 65059 10 Protein degradation/ubiquitin pathway 65060 E Neuronal nuclear inclusions 65100 -

PRKACA Mediates Resistance to HER2-Targeted Therapy in Breast Cancer Cells and Restores Anti-Apoptotic Signaling
Oncogene (2015) 34, 2061–2071 © 2015 Macmillan Publishers Limited All rights reserved 0950-9232/15 www.nature.com/onc ORIGINAL ARTICLE PRKACA mediates resistance to HER2-targeted therapy in breast cancer cells and restores anti-apoptotic signaling SE Moody1,2,3, AC Schinzel1, S Singh1, F Izzo1, MR Strickland1, L Luo1,2, SR Thomas3, JS Boehm3, SY Kim4, ZC Wang5,6 and WC Hahn1,2,3 Targeting HER2 with antibodies or small molecule inhibitors in HER2-positive breast cancer leads to improved survival, but resistance is a common clinical problem. To uncover novel mechanisms of resistance to anti-HER2 therapy in breast cancer, we performed a kinase open reading frame screen to identify genes that rescue HER2-amplified breast cancer cells from HER2 inhibition or suppression. In addition to multiple members of the MAPK (mitogen-activated protein kinase) and PI3K (phosphoinositide 3-kinase) signaling pathways, we discovered that expression of the survival kinases PRKACA and PIM1 rescued cells from anti-HER2 therapy. Furthermore, we observed elevated PRKACA expression in trastuzumab-resistant breast cancer samples, indicating that this pathway is activated in breast cancers that are clinically resistant to trastuzumab-containing therapy. We found that neither PRKACA nor PIM1 restored MAPK or PI3K activation after lapatinib or trastuzumab treatment, but rather inactivated the pro-apoptotic protein BAD, the BCl-2-associated death promoter, thereby permitting survival signaling through BCL- XL. Pharmacological blockade of BCL-XL/BCL-2 partially abrogated the rescue effects conferred by PRKACA and PIM1, and sensitized cells to lapatinib treatment. These observations suggest that combined targeting of HER2 and the BCL-XL/BCL-2 anti-apoptotic pathway may increase responses to anti-HER2 therapy in breast cancer and decrease the emergence of resistant disease. -

Identification of HRAS Mutations and Absence of GNAQ Or GNA11
Modern Pathology (2013) 26, 1320–1328 1320 & 2013 USCAP, Inc All rights reserved 0893-3952/13 $32.00 Identification of HRAS mutations and absence of GNAQ or GNA11 mutations in deep penetrating nevi Ryan P Bender1, Matthew J McGinniss2, Paula Esmay1, Elsa F Velazquez3,4 and Julie DR Reimann3,4 1Caris Life Sciences, Phoenix, AZ, USA; 2Genoptix Medical Laboratory, Carlsbad, CA, USA; 3Dermatopathology Division, Miraca Life Sciences Research Institute, Newton, MA, USA and 4Department of Dermatology, Tufts Medical Center, Boston, MA, USA HRAS is mutated in B15% of Spitz nevi, and GNAQ or GNA11 is mutated in blue nevi (46–83% and B7% respectively). Epithelioid blue nevi and deep penetrating nevi show features of both blue nevi (intradermal location, pigmentation) and Spitz nevi (epithelioid morphology). Epithelioid blue nevi and deep penetrating nevi can also show overlapping features with melanoma, posing a diagnostic challenge. Although epithelioid blue nevi are considered blue nevic variants, no GNAQ or GNA11 mutations have been reported. Classification of deep penetrating nevi as blue nevic variants has also been proposed, however, no GNAQ or GNA11 mutations have been reported and none have been tested for HRAS mutations. To better characterize these tumors, we performed mutational analysis for GNAQ, GNA11, and HRAS, with blue nevi and Spitz nevi as controls. Within deep penetrating nevi, none demonstrated GNAQ or GNA11 mutations (0/38). However, 6% revealed HRAS mutation (2/32). Twenty percent of epithelioid blue nevi contained a GNAQ mutation (2/10), while none displayed GNA11 or HRAS mutation. Eighty-seven percent of blue nevi contained a GNAQ mutation (26/30), 4% a GNA11 mutation (1/28), and none an HRAS mutation. -

Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association Between BMI and Adult-Onset Non- Atopic
Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association between BMI and Adult-Onset Non- Atopic Asthma Ayoung Jeong 1,2, Medea Imboden 1,2, Akram Ghantous 3, Alexei Novoloaca 3, Anne-Elie Carsin 4,5,6, Manolis Kogevinas 4,5,6, Christian Schindler 1,2, Gianfranco Lovison 7, Zdenko Herceg 3, Cyrille Cuenin 3, Roel Vermeulen 8, Deborah Jarvis 9, André F. S. Amaral 9, Florian Kronenberg 10, Paolo Vineis 11,12 and Nicole Probst-Hensch 1,2,* 1 Swiss Tropical and Public Health Institute, 4051 Basel, Switzerland; [email protected] (A.J.); [email protected] (M.I.); [email protected] (C.S.) 2 Department of Public Health, University of Basel, 4001 Basel, Switzerland 3 International Agency for Research on Cancer, 69372 Lyon, France; [email protected] (A.G.); [email protected] (A.N.); [email protected] (Z.H.); [email protected] (C.C.) 4 ISGlobal, Barcelona Institute for Global Health, 08003 Barcelona, Spain; [email protected] (A.-E.C.); [email protected] (M.K.) 5 Universitat Pompeu Fabra (UPF), 08002 Barcelona, Spain 6 CIBER Epidemiología y Salud Pública (CIBERESP), 08005 Barcelona, Spain 7 Department of Economics, Business and Statistics, University of Palermo, 90128 Palermo, Italy; [email protected] 8 Environmental Epidemiology Division, Utrecht University, Institute for Risk Assessment Sciences, 3584CM Utrecht, Netherlands; [email protected] 9 Population Health and Occupational Disease, National Heart and Lung Institute, Imperial College, SW3 6LR London, UK; [email protected] (D.J.); [email protected] (A.F.S.A.) 10 Division of Genetic Epidemiology, Medical University of Innsbruck, 6020 Innsbruck, Austria; [email protected] 11 MRC-PHE Centre for Environment and Health, School of Public Health, Imperial College London, W2 1PG London, UK; [email protected] 12 Italian Institute for Genomic Medicine (IIGM), 10126 Turin, Italy * Correspondence: [email protected]; Tel.: +41-61-284-8378 Int.