Prevalence of Mutations in TSHR, GNAS, PRKAR1A and RAS Genes
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Hras Intracellular Trafficking and Signal Transduction Jodi Ho-Jung Mckay Iowa State University
Iowa State University Capstones, Theses and Retrospective Theses and Dissertations Dissertations 2007 HRas intracellular trafficking and signal transduction Jodi Ho-Jung McKay Iowa State University Follow this and additional works at: https://lib.dr.iastate.edu/rtd Part of the Biological Phenomena, Cell Phenomena, and Immunity Commons, Cancer Biology Commons, Cell Biology Commons, Genetics and Genomics Commons, and the Medical Cell Biology Commons Recommended Citation McKay, Jodi Ho-Jung, "HRas intracellular trafficking and signal transduction" (2007). Retrospective Theses and Dissertations. 13946. https://lib.dr.iastate.edu/rtd/13946 This Dissertation is brought to you for free and open access by the Iowa State University Capstones, Theses and Dissertations at Iowa State University Digital Repository. It has been accepted for inclusion in Retrospective Theses and Dissertations by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. HRas intracellular trafficking and signal transduction by Jodi Ho-Jung McKay A dissertation submitted to the graduate faculty in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY Major: Genetics Program of Study Committee: Janice E. Buss, Co-major Professor Linda Ambrosio, Co-major Professor Diane Bassham Drena Dobbs Ted Huiatt Iowa State University Ames, Iowa 2007 Copyright © Jodi Ho-Jung McKay, 2007. All rights reserved. UMI Number: 3274881 Copyright 2007 by McKay, Jodi Ho-Jung All rights reserved. UMI Microform 3274881 Copyright 2008 by ProQuest Information and Learning Company. All rights reserved. This microform edition is protected against unauthorized copying under Title 17, United States Code. ProQuest Information and Learning Company 300 North Zeeb Road P.O. -

Myopia Genetics Report
Special Issue IMI – Myopia Genetics Report Milly S. Tedja,1,2 Annechien E. G. Haarman,1,2 Magda A. Meester-Smoor,1,2 Jaakko Kaprio,3,4 David A. Mackey,5–7 Jeremy A. Guggenheim,8 Christopher J. Hammond,9 Virginie J. M. Verhoeven,1,2,10 and Caroline C. W. Klaver1,2,11; for the CREAM Consortium 1Department of Ophthalmology, Erasmus Medical Center, Rotterdam, the Netherlands 2Department of Epidemiology, Erasmus Medical Center, Rotterdam, the Netherlands 3Institute for Molecular Medicine, University of Helsinki, Helsinki, Finland 4Department of Public Health, University of Helsinki, Helsinki, Finland 5Centre for Eye Research Australia, Ophthalmology, Department of Surgery, University of Melbourne, Royal Victorian Eye and Ear Hospital, Melbourne, Victoria, Australia 6Department of Ophthalmology, Menzies Institute of Medical Research, University of Tasmania, Hobart, Tasmania, Australia 7Centre for Ophthalmology and Visual Science, Lions Eye Institute, University of Western Australia, Perth, Western Australia, Australia 8School of Optometry and Vision Sciences, Cardiff University, Cardiff, United Kingdom 9Section of Academic Ophthalmology, School of Life Course Sciences, King’s College London, London, United Kingdom 10Department of Clinical Genetics, Erasmus Medical Center, Rotterdam, the Netherlands 11Department of Ophthalmology, Radboud University Medical Center, Nijmegen, the Netherlands Correspondence: Caroline C. W. The knowledge on the genetic background of refractive error and myopia has expanded Klaver, Erasmus Medical Center, dramatically in the past few years. This white paper aims to provide a concise summary of Room Na-2808, P.O. Box 2040, 3000 current genetic findings and defines the direction where development is needed. CA, Rotterdam, the Netherlands; [email protected]. We performed an extensive literature search and conducted informal discussions with key MST and AEGH contributed equally to stakeholders. -

Identification of HRAS Mutations and Absence of GNAQ Or GNA11
Modern Pathology (2013) 26, 1320–1328 1320 & 2013 USCAP, Inc All rights reserved 0893-3952/13 $32.00 Identification of HRAS mutations and absence of GNAQ or GNA11 mutations in deep penetrating nevi Ryan P Bender1, Matthew J McGinniss2, Paula Esmay1, Elsa F Velazquez3,4 and Julie DR Reimann3,4 1Caris Life Sciences, Phoenix, AZ, USA; 2Genoptix Medical Laboratory, Carlsbad, CA, USA; 3Dermatopathology Division, Miraca Life Sciences Research Institute, Newton, MA, USA and 4Department of Dermatology, Tufts Medical Center, Boston, MA, USA HRAS is mutated in B15% of Spitz nevi, and GNAQ or GNA11 is mutated in blue nevi (46–83% and B7% respectively). Epithelioid blue nevi and deep penetrating nevi show features of both blue nevi (intradermal location, pigmentation) and Spitz nevi (epithelioid morphology). Epithelioid blue nevi and deep penetrating nevi can also show overlapping features with melanoma, posing a diagnostic challenge. Although epithelioid blue nevi are considered blue nevic variants, no GNAQ or GNA11 mutations have been reported. Classification of deep penetrating nevi as blue nevic variants has also been proposed, however, no GNAQ or GNA11 mutations have been reported and none have been tested for HRAS mutations. To better characterize these tumors, we performed mutational analysis for GNAQ, GNA11, and HRAS, with blue nevi and Spitz nevi as controls. Within deep penetrating nevi, none demonstrated GNAQ or GNA11 mutations (0/38). However, 6% revealed HRAS mutation (2/32). Twenty percent of epithelioid blue nevi contained a GNAQ mutation (2/10), while none displayed GNA11 or HRAS mutation. Eighty-seven percent of blue nevi contained a GNAQ mutation (26/30), 4% a GNA11 mutation (1/28), and none an HRAS mutation. -

Role of Phosphodiesterases in the Pathophysiology of Neurodevelopmental Disorders
Molecular Psychiatry https://doi.org/10.1038/s41380-020-00997-9 REVIEW ARTICLE Role of phosphodiesterases in the pathophysiology of neurodevelopmental disorders 1 2 Sébastien Delhaye ● Barbara Bardoni Received: 4 July 2020 / Revised: 3 December 2020 / Accepted: 9 December 2020 © The Author(s), under exclusive licence to Springer Nature Limited part of Springer Nature 2021. This article is published with open access Abstract Phosphodiesterases (PDEs) are enzymes involved in the homeostasis of both cAMP and cGMP. They are members of a family of proteins that includes 11 subfamilies with different substrate specificities. Their main function is to catalyze the hydrolysis of cAMP, cGMP, or both. cAMP and cGMP are two key second messengers that modulate a wide array of intracellular processes and neurobehavioral functions, including memory and cognition. Even if these enzymes are present in all tissues, we focused on those PDEs that are expressed in the brain. We took into consideration genetic variants in patients affected by neurodevelopmental disorders, phenotypes of animal models, and pharmacological effects of PDE inhibitors, a class of drugs in rapid evolution and increasing application to brain disorders. Collectively, these data indicate the potential 1234567890();,: 1234567890();,: of PDE modulators to treat neurodevelopmental diseases characterized by learning and memory impairment, alteration of behaviors associated with depression, and deficits in social interaction. Indeed, clinical trials are in progress to treat patients with Alzheimer’s disease, schizophrenia, depression, and autism spectrum disorders. Among the most recent results, the application of some PDE inhibitors (PDE2A, PDE3, PDE4/4D, and PDE10A) to treat neurodevelopmental diseases, including autism spectrum disorders and intellectual disability, is a significant advance, since no specific therapies are available for these disorders that have a large prevalence. -

Genomic Alterations Associated with Recurrence and TNBC Subtype in High
Author Manuscript Published OnlineFirst on August 31, 2018; DOI: 10.1158/1541-7786.MCR-18-0619 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. Genomic Alterations Associated with Recurrence and TNBC Subtype in High- risk Early Breast Cancers Timothy R. Wilson1*, Akshata R. Udyavar2*, Ching-Wei Chang3, Jill M. Spoerke1, Junko Aimi1, Heidi M. Savage1, Anneleen Daemen2, Joyce A. O’Shaughnessy4,5,6, Richard Bourgon2 and Mark R. Lackner1 Departments of 1Oncology Biomarker Development, 2Bioinformatics and Computational Biology, 3Biostatistics, Genentech Inc., 1 DNA Way, South San Francisco, CA, USA.4US Oncology, 5Baylor University Medical Center, 6Texas Oncology, Dallas, TX, USA. *These authors contributed equally to this work. Correspondence and request for materials should be addressed to T.R.W. (email: [email protected]) or M.R.L. (email: [email protected]). Running Title: Genomic Profiling of High-risk Early Breast Cancers Keywords: early breast cancer, next generation sequencing, risk of recurrence, triple negative breast cancer, tumor mutational burden Conflicts of interest: TRW, ARY, CWC, JMS, JA, HMS, AD, RB and MRL are employed by Genentech and have stocks in Roche. JOS has received speaking fees from Blue Earth Diagnostics and Novartis, consulting fees from AstraZeneca, Celgene, Eli Lilly, LaRoche, Lilly USA, Merck, Novartis and Pfizer, Honoraria from Seattle Genetics and Downloaded from mcr.aacrjournals.org on September 27, 2021. © 2018 American Association for Cancer Research. Author Manuscript Published OnlineFirst on August 31, 2018; DOI: 10.1158/1541-7786.MCR-18-0619 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. -

Mosaic Activating Mutations in GNA11 and GNAQ Are Associated with Phakomatosis Pigmentovascularis and Extensive Dermal Melanocytosis Anna C
ORIGINAL ARTICLE Mosaic Activating Mutations in GNA11 and GNAQ Are Associated with Phakomatosis Pigmentovascularis and Extensive Dermal Melanocytosis Anna C. Thomas1,18, Zhiqiang Zeng2,18, Jean-Baptiste Rivie`re3,18, Ryan O’Shaughnessy4, Lara Al-Olabi1, Judith St.-Onge3, David J. Atherton5,He´le`ne Aubert6, Lorea Bagazgoitia7, Se´bastien Barbarot6, Emmanuelle Bourrat8,9, Christine Chiaverini10, W. Kling Chong11, Yannis Duffourd3, Mary Glover5, Leopold Groesser12, Smail Hadj-Rabia13, Henning Hamm14, Rudolf Happle15, Imran Mushtaq16, Jean-Philippe Lacour10, Regula Waelchli5, Marion Wobser14, Pierre Vabres3,17,19, E. Elizabeth Patton2,19 and Veronica A. Kinsler1,5,19 Common birthmarks can be an indicator of underlying genetic disease but are often overlooked. Mongolian blue spots (dermal melanocytosis) are usually localized and transient, but they can be extensive, permanent, and associated with extracutaneous abnormalities. Co-occurrence with vascular birthmarks defines a subtype of phakomatosis pigmentovascularis, a group of syndromes associated with neurovascular, ophthalmological, overgrowth, and malignant complications. Here, we discover that extensive dermal melanocytosis and pha- komatosis pigmentovascularis are associated with activating mutations in GNA11 and GNAQ, genes that encode Ga subunits of heterotrimeric G proteins. The mutations were detected at very low levels in affected tissues but were undetectable in the blood, indicating that these conditions are postzygotic mosaic disorders. R183C Q209L In vitro expression of mutant GNA11 and GNA11 in human cell lines demonstrated activation of the downstream p38 MAPK signaling pathway and the p38, JNK, and ERK pathways, respectively. Transgenic R183C mosaic zebrafish models expressing mutant GNA11 under promoter mitfa developed extensive dermal melanocytosis recapitulating the human phenotype. Phakomatosis pigmentovascularis and extensive dermal melanocytosis are therefore diagnoses in the group of mosaic heterotrimeric G-protein disorders, joining McCune-Albright and Sturge-Weber syndromes. -

Investigating the Mechanism of Hepatocellular Carcinoma Progression by Constructing Genetic and Epigenetic Networks Using NGS Data Identification and Big Database Mining Method
www.impactjournals.com/oncotarget/ Oncotarget, Vol. 7, No. 48 Research Paper Investigating the mechanism of hepatocellular carcinoma progression by constructing genetic and epigenetic networks using NGS data identification and big database mining method Cheng-Wei Li1, Ping-Yao Chang1, Bor-Sen Chen1 1Laboratory of Control and Systems Biology, National Tsing Hua University, Hsinchu, Taiwan Correspondence to: Bor-Sen Chen, email: [email protected] Keywords: DNA methylation, multiple potential drugs, hepatocarcinogenesis, miRNAs, principal network projection Received: February 19, 2016 Accepted: October 26, 2016 Published: November 04, 2016 ABSTRACT The mechanisms leading to the development and progression of hepatocellular carcinoma (HCC) are complicated and regulated genetically and epigenetically. The recent advancement in high-throughput sequencing has facilitated investigations into the role of genetic and epigenetic regulations in hepatocarcinogenesis. Therefore, we used systems biology and big database mining to construct genetic and epigenetic networks (GENs) using the information about mRNA, miRNA, and methylation profiles of HCC patients. Our approach involves analyzing gene regulatory networks (GRNs), protein-protein networks (PPINs), and epigenetic networks at different stages of hepatocarcinogenesis. The core GENs, influencing each stage of HCC, were extracted via principal network projection (PNP). The pathways during different stages of HCC were compared. We observed that extracellular signals were further transduced to -

FARE2021WINNERS Sorted by Institute
FARE2021WINNERS Sorted By Institute Swati Shah Postdoctoral Fellow CC Radiology/Imaging/PET and Neuroimaging Characterization of CNS involvement in Ebola-Infected Macaques using Magnetic Resonance Imaging, 18F-FDG PET and Immunohistology The Ebola (EBOV) virus outbreak in Western Africa resulted in residual neurologic abnormalities in survivors. Many case studies detected EBOV in the CSF, suggesting that the neurologic sequelae in survivors is related to viral presence. In the periphery, EBOV infects endothelial cells and triggers a “cytokine stormâ€. However, it is unclear whether a similar process occurs in the brain, with secondary neuroinflammation, neuronal loss and blood-brain barrier (BBB) compromise, eventually leading to lasting neurological damage. We have used in vivo imaging and post-necropsy immunostaining to elucidate the CNS pathophysiology in Rhesus macaques infected with EBOV (Makona). Whole brain MRI with T1 relaxometry (pre- and post-contrast) and FDG-PET were performed to monitor the progression of disease in two cohorts of EBOV infected macaques from baseline to terminal endpoint (day 5-6). Post-necropsy, multiplex fluorescence immunohistochemical (MF-IHC) staining for various cellular markers in the thalamus and brainstem was performed. Serial blood and CSF samples were collected to assess disease progression. The linear mixed effect model was used for statistical analysis. Post-infection, we first detected EBOV in the serum (day 3) and CSF (day 4) with dramatic increases until euthanasia. The standard uptake values of FDG-PET relative to whole brain uptake (SUVr) in the midbrain, pons, and thalamus increased significantly over time (p<0.01) and positively correlated with blood viremia (p≤0.01). -
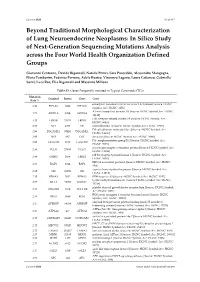
Beyond Traditional Morphological Characterization of Lung
Cancers 2020 S1 of S15 Beyond Traditional Morphological Characterization of Lung Neuroendocrine Neoplasms: In Silico Study of Next-Generation Sequencing Mutations Analysis across the Four World Health Organization Defined Groups Giovanni Centonze, Davide Biganzoli, Natalie Prinzi, Sara Pusceddu, Alessandro Mangogna, Elena Tamborini, Federica Perrone, Adele Busico, Vincenzo Lagano, Laura Cattaneo, Gabriella Sozzi, Luca Roz, Elia Biganzoli and Massimo Milione Table S1. Genes Frequently mutated in Typical Carcinoids (TCs). Mutation Original Entrez Gene Gene Rate % eukaryotic translation initiation factor 1A X-linked [Source: HGNC 4.84 EIF1AX 1964 EIF1AX Symbol; Acc: HGNC: 3250] AT-rich interaction domain 1A [Source: HGNC Symbol;Acc: HGNC: 4.71 ARID1A 8289 ARID1A 11110] LDL receptor related protein 1B [Source: HGNC Symbol; Acc: 4.35 LRP1B 53353 LRP1B HGNC: 6693] 3.53 NF1 4763 NF1 neurofibromin 1 [Source: HGNC Symbol;Acc: HGNC: 7765] DS cell adhesion molecule like 1 [Source: HGNC Symbol; Acc: 2.90 DSCAML1 57453 DSCAML1 HGNC: 14656] 2.90 DST 667 DST dystonin [Source: HGNC Symbol;Acc: HGNC: 1090] FA complementation group D2 [Source: HGNC Symbol; Acc: 2.90 FANCD2 2177 FANCD2 HGNC: 3585] piccolo presynaptic cytomatrix protein [Source: HGNC Symbol; Acc: 2.90 PCLO 27445 PCLO HGNC: 13406] erb-b2 receptor tyrosine kinase 2 [Source: HGNC Symbol; Acc: 2.44 ERBB2 2064 ERBB2 HGNC: 3430] BRCA1 associated protein 1 [Source: HGNC Symbol; Acc: HGNC: 2.35 BAP1 8314 BAP1 950] capicua transcriptional repressor [Source: HGNC Symbol; Acc: 2.35 CIC 23152 CIC HGNC: -

Identification of Novel GNAS Mutations in Intramuscular Myxoma Using Next- Generation Sequencing with Single-Molecule Tagged Molecular Inversion Probes Elise M
Bekers et al. Diagnostic Pathology (2019) 14:15 https://doi.org/10.1186/s13000-019-0787-3 RESEARCH Open Access Identification of novel GNAS mutations in intramuscular myxoma using next- generation sequencing with single-molecule tagged molecular inversion probes Elise M. Bekers1,2* , Astrid Eijkelenboom1, Paul Rombout1, Peter van Zwam3, Suzanne Mol4, Emiel Ruijter5, Blanca Scheijen1 and Uta Flucke1 Abstract Background: Intramuscular myxoma (IM) is a hypocellular benign soft tissue neoplasm characterized by abundant myxoid stroma and occasional hypercellular areas. These tumors can, especially on biopsy material, be difficult to distinguish from low-grade fibromyxoid sarcoma or low-grade myxofibrosarcoma. GNAS mutations are frequently involved in IM, in contrast to these other malignant tumors. Therefore, sensitive molecular techniques for detection of GNAS aberrations in IM, which frequently yield low amounts of DNA due to poor cellularity, will be beneficial for differential diagnosis. Methods: In our study, a total of 34 IM samples from 33 patients were analyzed for the presence of GNAS mutations, of which 29 samples were analyzed using a gene-specific TaqMan genotyping assay for the detection of GNAS hotspot mutations c.601C > T and c602G > A in IM, and 32 samples using a novel next generation sequencing (NGS)-based approach employing single-molecule tagged molecular inversion probes (smMIP) to identify mutations in exon 8 and 9 of GNAS. Results between the two assays were compared for their ability to detect GNAS mutations with high confidence. Results: In total, 23 of 34 samples were successfully analyzed with both techniques showing GNAS mutations in 12 out of 23 (52%) samples. -

Original Article Lncrna UCHL1-AS1 Prevents Cell Mobility of Hepatocellular Carcinoma: a Study Based on in Vitro and Bioinformatics
Int J Clin Exp Pathol 2018;11(5):2270-2280 www.ijcep.com /ISSN:1936-2625/IJCEP0073243 Original Article LncRNA UCHL1-AS1 prevents cell mobility of hepatocellular carcinoma: a study based on in vitro and bioinformatics Rui Zhang*, Yichen Wei*, Li’ou Zhu, Lanshan Huang, Yan Wei, Gang Chen, Yiwu Dang, Zhenbo Feng Department of Pathology, The First Affiliated Hospital of Guangxi Medical University, Nanning 530021, Guangxi Zhuang Autonomous Region, China. *Equal contributors. Received January 22, 2018; Accepted March 8, 2018; Epub May 1, 2018; Published May 15, 2018 Abstract: We set out to investigate biological functions and potential molecular mechanisms of long non-coding RNA (lncRNA) in hepatocellular carcinoma (HCC). HCC cell line Bel-7404 was cultured and transfected with antisense to the ubiquitin carboxyl-terminal hydrolase L1 (UCHL1-AS1). Viability and mobility were detected by MTT and wound healing assays. Additionally, enrichment analysis and functional networks of UCHL1-AS1 related genes in HCC were performed. Results showed that high level UCHL1-AS1 could effectively inhibit HCC cell migration. However, there was no significant correlation between overexpressed UCHL1-AS1 and HCC proliferation. Meanwhile, BMP4, CALM3, and HRAS were selected from 204 genes that related to UCHL1-AS1. All of these hub genes play critical roles in HCC occurrence and development. Thus, underlying molecular mechanisms among hub genes and UCHL1-AS1 in HCC might be valuable for prognosis and treatment. Keywords: Long non-coding RNAs, UCHL1-AS1, hepatocellular carcinoma, mobility, bioinformatics Introduction which play pivotal roles in different processes of cell biology and pathology. LncRNAs have As the most common liver cancer worldwide, been broadly examined in tumors and some hepatocellular carcinoma (HCC) ranks highly in new functions of lncRNAs have updated our overall cancer-related deaths [1, 2].