Site-Specific Processing of Ras and Rap1 Switch I by a MARTX Toxin
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Hras Intracellular Trafficking and Signal Transduction Jodi Ho-Jung Mckay Iowa State University
Iowa State University Capstones, Theses and Retrospective Theses and Dissertations Dissertations 2007 HRas intracellular trafficking and signal transduction Jodi Ho-Jung McKay Iowa State University Follow this and additional works at: https://lib.dr.iastate.edu/rtd Part of the Biological Phenomena, Cell Phenomena, and Immunity Commons, Cancer Biology Commons, Cell Biology Commons, Genetics and Genomics Commons, and the Medical Cell Biology Commons Recommended Citation McKay, Jodi Ho-Jung, "HRas intracellular trafficking and signal transduction" (2007). Retrospective Theses and Dissertations. 13946. https://lib.dr.iastate.edu/rtd/13946 This Dissertation is brought to you for free and open access by the Iowa State University Capstones, Theses and Dissertations at Iowa State University Digital Repository. It has been accepted for inclusion in Retrospective Theses and Dissertations by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. HRas intracellular trafficking and signal transduction by Jodi Ho-Jung McKay A dissertation submitted to the graduate faculty in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY Major: Genetics Program of Study Committee: Janice E. Buss, Co-major Professor Linda Ambrosio, Co-major Professor Diane Bassham Drena Dobbs Ted Huiatt Iowa State University Ames, Iowa 2007 Copyright © Jodi Ho-Jung McKay, 2007. All rights reserved. UMI Number: 3274881 Copyright 2007 by McKay, Jodi Ho-Jung All rights reserved. UMI Microform 3274881 Copyright 2008 by ProQuest Information and Learning Company. All rights reserved. This microform edition is protected against unauthorized copying under Title 17, United States Code. ProQuest Information and Learning Company 300 North Zeeb Road P.O. -

Myopia Genetics Report
Special Issue IMI – Myopia Genetics Report Milly S. Tedja,1,2 Annechien E. G. Haarman,1,2 Magda A. Meester-Smoor,1,2 Jaakko Kaprio,3,4 David A. Mackey,5–7 Jeremy A. Guggenheim,8 Christopher J. Hammond,9 Virginie J. M. Verhoeven,1,2,10 and Caroline C. W. Klaver1,2,11; for the CREAM Consortium 1Department of Ophthalmology, Erasmus Medical Center, Rotterdam, the Netherlands 2Department of Epidemiology, Erasmus Medical Center, Rotterdam, the Netherlands 3Institute for Molecular Medicine, University of Helsinki, Helsinki, Finland 4Department of Public Health, University of Helsinki, Helsinki, Finland 5Centre for Eye Research Australia, Ophthalmology, Department of Surgery, University of Melbourne, Royal Victorian Eye and Ear Hospital, Melbourne, Victoria, Australia 6Department of Ophthalmology, Menzies Institute of Medical Research, University of Tasmania, Hobart, Tasmania, Australia 7Centre for Ophthalmology and Visual Science, Lions Eye Institute, University of Western Australia, Perth, Western Australia, Australia 8School of Optometry and Vision Sciences, Cardiff University, Cardiff, United Kingdom 9Section of Academic Ophthalmology, School of Life Course Sciences, King’s College London, London, United Kingdom 10Department of Clinical Genetics, Erasmus Medical Center, Rotterdam, the Netherlands 11Department of Ophthalmology, Radboud University Medical Center, Nijmegen, the Netherlands Correspondence: Caroline C. W. The knowledge on the genetic background of refractive error and myopia has expanded Klaver, Erasmus Medical Center, dramatically in the past few years. This white paper aims to provide a concise summary of Room Na-2808, P.O. Box 2040, 3000 current genetic findings and defines the direction where development is needed. CA, Rotterdam, the Netherlands; [email protected]. We performed an extensive literature search and conducted informal discussions with key MST and AEGH contributed equally to stakeholders. -

Supplementary Materials
Supplementary Materials COMPARATIVE ANALYSIS OF THE TRANSCRIPTOME, PROTEOME AND miRNA PROFILE OF KUPFFER CELLS AND MONOCYTES Andrey Elchaninov1,3*, Anastasiya Lokhonina1,3, Maria Nikitina2, Polina Vishnyakova1,3, Andrey Makarov1, Irina Arutyunyan1, Anastasiya Poltavets1, Evgeniya Kananykhina2, Sergey Kovalchuk4, Evgeny Karpulevich5,6, Galina Bolshakova2, Gennady Sukhikh1, Timur Fatkhudinov2,3 1 Laboratory of Regenerative Medicine, National Medical Research Center for Obstetrics, Gynecology and Perinatology Named after Academician V.I. Kulakov of Ministry of Healthcare of Russian Federation, Moscow, Russia 2 Laboratory of Growth and Development, Scientific Research Institute of Human Morphology, Moscow, Russia 3 Histology Department, Medical Institute, Peoples' Friendship University of Russia, Moscow, Russia 4 Laboratory of Bioinformatic methods for Combinatorial Chemistry and Biology, Shemyakin-Ovchinnikov Institute of Bioorganic Chemistry of the Russian Academy of Sciences, Moscow, Russia 5 Information Systems Department, Ivannikov Institute for System Programming of the Russian Academy of Sciences, Moscow, Russia 6 Genome Engineering Laboratory, Moscow Institute of Physics and Technology, Dolgoprudny, Moscow Region, Russia Figure S1. Flow cytometry analysis of unsorted blood sample. Representative forward, side scattering and histogram are shown. The proportions of negative cells were determined in relation to the isotype controls. The percentages of positive cells are indicated. The blue curve corresponds to the isotype control. Figure S2. Flow cytometry analysis of unsorted liver stromal cells. Representative forward, side scattering and histogram are shown. The proportions of negative cells were determined in relation to the isotype controls. The percentages of positive cells are indicated. The blue curve corresponds to the isotype control. Figure S3. MiRNAs expression analysis in monocytes and Kupffer cells. Full-length of heatmaps are presented. -

Rap1 Acts Via Multiple Mechanisms to Position Canoe/Afadin and Adherens Junctions and Mediate Apical-Basal Polarity Establishment
bioRxiv preprint doi: https://doi.org/10.1101/170977; this version posted July 31, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Rap1 acts via multiple mechanisms to position Canoe/Afadin and adherens junctions and mediate apical-basal polarity establishment Teresa T. Bonello1, Kia Z. Perez-Vale2, Kaelyn D. Sumigray3, and Mark Peifer1,2,3* 1 Department of Biology, University of North Carolina at Chapel Hill, CB#3280, Chapel Hill, NC 27599-3280, USA 2 Curriculum in Genetics and Molecular Biology, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599, USA 3 Lineberger Comprehensive Cancer Center, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599, USA Running Title Active Rap1 positions Canoe and AJs 6950 words * To whom correspondence should be addressed Email: [email protected] Phone: (919) 962-2272 Abbreviations used: α-cat, alpha-catenin; β-cat, beta-catenin; AJ, adherens junction; Arm, Armadillo; Baz, BazooKa; CA, constitutively active; Cno, Canoe; DE-cad, Drosophila E-cadherin; Dzy, Dizzy; GAP, GTPase activating protein; GDP, guanosine diphosphate; GEF, guanine nucleotide exchange factor; GFP, green fluorescent protein; GTP, guanosine triphosphate; IF, immunofluorescence; MIP, maximum intensity projection; RA, Ras-associated; RFP, red fluorescent protein; SAJ, spot adherens junction; shRNA, short hairpin RNA; TCJ, tricellular junction; WT, wildtype 1 bioRxiv preprint doi: https://doi.org/10.1101/170977; this version posted July 31, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. -

Identification of HRAS Mutations and Absence of GNAQ Or GNA11
Modern Pathology (2013) 26, 1320–1328 1320 & 2013 USCAP, Inc All rights reserved 0893-3952/13 $32.00 Identification of HRAS mutations and absence of GNAQ or GNA11 mutations in deep penetrating nevi Ryan P Bender1, Matthew J McGinniss2, Paula Esmay1, Elsa F Velazquez3,4 and Julie DR Reimann3,4 1Caris Life Sciences, Phoenix, AZ, USA; 2Genoptix Medical Laboratory, Carlsbad, CA, USA; 3Dermatopathology Division, Miraca Life Sciences Research Institute, Newton, MA, USA and 4Department of Dermatology, Tufts Medical Center, Boston, MA, USA HRAS is mutated in B15% of Spitz nevi, and GNAQ or GNA11 is mutated in blue nevi (46–83% and B7% respectively). Epithelioid blue nevi and deep penetrating nevi show features of both blue nevi (intradermal location, pigmentation) and Spitz nevi (epithelioid morphology). Epithelioid blue nevi and deep penetrating nevi can also show overlapping features with melanoma, posing a diagnostic challenge. Although epithelioid blue nevi are considered blue nevic variants, no GNAQ or GNA11 mutations have been reported. Classification of deep penetrating nevi as blue nevic variants has also been proposed, however, no GNAQ or GNA11 mutations have been reported and none have been tested for HRAS mutations. To better characterize these tumors, we performed mutational analysis for GNAQ, GNA11, and HRAS, with blue nevi and Spitz nevi as controls. Within deep penetrating nevi, none demonstrated GNAQ or GNA11 mutations (0/38). However, 6% revealed HRAS mutation (2/32). Twenty percent of epithelioid blue nevi contained a GNAQ mutation (2/10), while none displayed GNA11 or HRAS mutation. Eighty-seven percent of blue nevi contained a GNAQ mutation (26/30), 4% a GNA11 mutation (1/28), and none an HRAS mutation. -

Prevalence of Mutations in TSHR, GNAS, PRKAR1A and RAS Genes
European Journal of Endocrinology (2008) 159 623–631 ISSN 0804-4643 CLINICAL STUDY Prevalence of mutations in TSHR, GNAS, PRKAR1A and RAS genes in a large series of toxic thyroid adenomas from Galicia, an iodine-deficient area in NW Spain F Palos-Paz1, O Perez-Guerra1, J Cameselle-Teijeiro3,CRueda-Chimeno5, F Barreiro-Morandeira4, J Lado-Abeal1,2 and the Galician Group for the Study of Toxic Multinodular Goitre: D Araujo Vilar1,2, R Argueso7, O Barca1, MBotana7, J M Cabezas-Agrı´cola2, P Catalina6, L Dominguez Gerpe1, T Fernandez9, A Mato8, A Nun˜o11,MPenin10 and B Victoria1 1Unidade de Enfermedades Tiroideas e Metabo´licas (UETeM), 2Endocrinology Section, Department of Medicine, 3Pathology Department and 4Surgery Department, Complexo Hospitalario Universitary de Santiago (CHUS), University of Santiago de Compostela, Santiago de Compostela, 15705, Spain, 5General Surgery Section and 6Endocrinology Section, Complexo Hospitalario de Pontevedra, Pontevedra, Spain, 7Endocrinology Section, Complexo Hospitalario Xeral-Calde, Lugo, Spain, 8Endocrinology Section, Complexo Hospitalario de Ourense, Ourense, Spain, 9Endocrinology Section, Complexo Hospitalario Universitario Juan-Canalejo, A Corun˜a, Spain, 10Endocrinology Section, Hospital Arquitecto Marcide, Ferrol, Spain and 11General Surgery Section, Hospital do Meixoeiro, Complexo Hospitalario Universitario de Vigo, Vigo, Spain (Correspondence should be addressed to J Lado-Abeal who is now at UETeM, Department of Medicine, School of Medicine, University of Santiago de Compostela, C/San Francisco sn 15705, Santiago de Compostela, Spain; Email: [email protected]) Abstract Objective: Toxic thyroid adenoma (TA) is a common cause of hyperthyroidism. Mutations in the TSH receptor (TSHR) gene, and less frequently in the adenylate cyclase-stimulating G alpha protein (GNAS) gene, are well established causes of TA in Europe. -

Mosaic Activating Mutations in GNA11 and GNAQ Are Associated with Phakomatosis Pigmentovascularis and Extensive Dermal Melanocytosis Anna C
ORIGINAL ARTICLE Mosaic Activating Mutations in GNA11 and GNAQ Are Associated with Phakomatosis Pigmentovascularis and Extensive Dermal Melanocytosis Anna C. Thomas1,18, Zhiqiang Zeng2,18, Jean-Baptiste Rivie`re3,18, Ryan O’Shaughnessy4, Lara Al-Olabi1, Judith St.-Onge3, David J. Atherton5,He´le`ne Aubert6, Lorea Bagazgoitia7, Se´bastien Barbarot6, Emmanuelle Bourrat8,9, Christine Chiaverini10, W. Kling Chong11, Yannis Duffourd3, Mary Glover5, Leopold Groesser12, Smail Hadj-Rabia13, Henning Hamm14, Rudolf Happle15, Imran Mushtaq16, Jean-Philippe Lacour10, Regula Waelchli5, Marion Wobser14, Pierre Vabres3,17,19, E. Elizabeth Patton2,19 and Veronica A. Kinsler1,5,19 Common birthmarks can be an indicator of underlying genetic disease but are often overlooked. Mongolian blue spots (dermal melanocytosis) are usually localized and transient, but they can be extensive, permanent, and associated with extracutaneous abnormalities. Co-occurrence with vascular birthmarks defines a subtype of phakomatosis pigmentovascularis, a group of syndromes associated with neurovascular, ophthalmological, overgrowth, and malignant complications. Here, we discover that extensive dermal melanocytosis and pha- komatosis pigmentovascularis are associated with activating mutations in GNA11 and GNAQ, genes that encode Ga subunits of heterotrimeric G proteins. The mutations were detected at very low levels in affected tissues but were undetectable in the blood, indicating that these conditions are postzygotic mosaic disorders. R183C Q209L In vitro expression of mutant GNA11 and GNA11 in human cell lines demonstrated activation of the downstream p38 MAPK signaling pathway and the p38, JNK, and ERK pathways, respectively. Transgenic R183C mosaic zebrafish models expressing mutant GNA11 under promoter mitfa developed extensive dermal melanocytosis recapitulating the human phenotype. Phakomatosis pigmentovascularis and extensive dermal melanocytosis are therefore diagnoses in the group of mosaic heterotrimeric G-protein disorders, joining McCune-Albright and Sturge-Weber syndromes. -

Investigating the Mechanism of Hepatocellular Carcinoma Progression by Constructing Genetic and Epigenetic Networks Using NGS Data Identification and Big Database Mining Method
www.impactjournals.com/oncotarget/ Oncotarget, Vol. 7, No. 48 Research Paper Investigating the mechanism of hepatocellular carcinoma progression by constructing genetic and epigenetic networks using NGS data identification and big database mining method Cheng-Wei Li1, Ping-Yao Chang1, Bor-Sen Chen1 1Laboratory of Control and Systems Biology, National Tsing Hua University, Hsinchu, Taiwan Correspondence to: Bor-Sen Chen, email: [email protected] Keywords: DNA methylation, multiple potential drugs, hepatocarcinogenesis, miRNAs, principal network projection Received: February 19, 2016 Accepted: October 26, 2016 Published: November 04, 2016 ABSTRACT The mechanisms leading to the development and progression of hepatocellular carcinoma (HCC) are complicated and regulated genetically and epigenetically. The recent advancement in high-throughput sequencing has facilitated investigations into the role of genetic and epigenetic regulations in hepatocarcinogenesis. Therefore, we used systems biology and big database mining to construct genetic and epigenetic networks (GENs) using the information about mRNA, miRNA, and methylation profiles of HCC patients. Our approach involves analyzing gene regulatory networks (GRNs), protein-protein networks (PPINs), and epigenetic networks at different stages of hepatocarcinogenesis. The core GENs, influencing each stage of HCC, were extracted via principal network projection (PNP). The pathways during different stages of HCC were compared. We observed that extracellular signals were further transduced to -

Rap1 Regulates Hepatic Stellate Cell Migration Through the Modulation of Rhoa Activity in Response to TGF‑Β1
INTERNATIONAL JOURNAL OF MOleCular meDICine 44: 491-502, 2019 Rap1 regulates hepatic stellate cell migration through the modulation of RhoA activity in response to TGF‑β1 MI-YOUNG MOON1, HEE-JUN KIM2, MO-JONG KIM2, SUNHO UHM1, JI‑WON PARK1, KI-TAE SUK3, JAE‑BONG PARK4, DONG-JUN KIM3 and SUNG-EUN KIM1 1Department of Internal Medicine, Hallym University Sacred Heart Hospital, College of Medicine, Hallym University, Anyang, Gyeonggi 14068; 2Ilsong Institute of Life Science, Hallym University, Anyang, Gyeonggi 14066; 3Department of Internal Medicine, Hallym University Chuncheon Sacred Heart Hospital, College of Medicine, Hallym University, Chuncheon, Gangwon 24253; 4Department of Biochemistry, College of Medicine, Hallym University, Chuncheon, Gangwon 24252, Republic of Korea Received November 1, 2018; Accepted May 28, 2019 DOI: 10.3892/ijmm.2019.4215 Abstract. Although the migration of hepatic stellate cells activation of RhoA in TGF‑β1-stimulated HSC‑T6 cells. These (HSCs) is important for hepatic fibrosis, the regulation of this findings suggest that TGF‑β1 regulates Rap1, resulting in the migration is poorly understood. Notably, transforming growth suppression of RhoA, activation of and formation of F‑actin factor (TGF)-β1 induces monocyte migration to sites of injury during the migration of HSCs. or inflammation during the early phase, but inhibits cell migra- tion during the late phase. In the present study, the role of Introduction transforming protein RhoA signaling in TGF-β1-induced HSC migration was investigated. TGF‑β1 was found to increase Hepatic fibrosis is characterized by the excessive deposition the protein and mRNA levels of smooth muscle actin and of extracellular matrix (ECM) mediated by activated hepatic collagen type I in HSC‑T6 cells. -

FARE2021WINNERS Sorted by Institute
FARE2021WINNERS Sorted By Institute Swati Shah Postdoctoral Fellow CC Radiology/Imaging/PET and Neuroimaging Characterization of CNS involvement in Ebola-Infected Macaques using Magnetic Resonance Imaging, 18F-FDG PET and Immunohistology The Ebola (EBOV) virus outbreak in Western Africa resulted in residual neurologic abnormalities in survivors. Many case studies detected EBOV in the CSF, suggesting that the neurologic sequelae in survivors is related to viral presence. In the periphery, EBOV infects endothelial cells and triggers a “cytokine stormâ€. However, it is unclear whether a similar process occurs in the brain, with secondary neuroinflammation, neuronal loss and blood-brain barrier (BBB) compromise, eventually leading to lasting neurological damage. We have used in vivo imaging and post-necropsy immunostaining to elucidate the CNS pathophysiology in Rhesus macaques infected with EBOV (Makona). Whole brain MRI with T1 relaxometry (pre- and post-contrast) and FDG-PET were performed to monitor the progression of disease in two cohorts of EBOV infected macaques from baseline to terminal endpoint (day 5-6). Post-necropsy, multiplex fluorescence immunohistochemical (MF-IHC) staining for various cellular markers in the thalamus and brainstem was performed. Serial blood and CSF samples were collected to assess disease progression. The linear mixed effect model was used for statistical analysis. Post-infection, we first detected EBOV in the serum (day 3) and CSF (day 4) with dramatic increases until euthanasia. The standard uptake values of FDG-PET relative to whole brain uptake (SUVr) in the midbrain, pons, and thalamus increased significantly over time (p<0.01) and positively correlated with blood viremia (p≤0.01). -
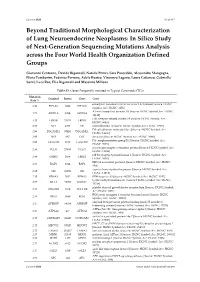
Beyond Traditional Morphological Characterization of Lung
Cancers 2020 S1 of S15 Beyond Traditional Morphological Characterization of Lung Neuroendocrine Neoplasms: In Silico Study of Next-Generation Sequencing Mutations Analysis across the Four World Health Organization Defined Groups Giovanni Centonze, Davide Biganzoli, Natalie Prinzi, Sara Pusceddu, Alessandro Mangogna, Elena Tamborini, Federica Perrone, Adele Busico, Vincenzo Lagano, Laura Cattaneo, Gabriella Sozzi, Luca Roz, Elia Biganzoli and Massimo Milione Table S1. Genes Frequently mutated in Typical Carcinoids (TCs). Mutation Original Entrez Gene Gene Rate % eukaryotic translation initiation factor 1A X-linked [Source: HGNC 4.84 EIF1AX 1964 EIF1AX Symbol; Acc: HGNC: 3250] AT-rich interaction domain 1A [Source: HGNC Symbol;Acc: HGNC: 4.71 ARID1A 8289 ARID1A 11110] LDL receptor related protein 1B [Source: HGNC Symbol; Acc: 4.35 LRP1B 53353 LRP1B HGNC: 6693] 3.53 NF1 4763 NF1 neurofibromin 1 [Source: HGNC Symbol;Acc: HGNC: 7765] DS cell adhesion molecule like 1 [Source: HGNC Symbol; Acc: 2.90 DSCAML1 57453 DSCAML1 HGNC: 14656] 2.90 DST 667 DST dystonin [Source: HGNC Symbol;Acc: HGNC: 1090] FA complementation group D2 [Source: HGNC Symbol; Acc: 2.90 FANCD2 2177 FANCD2 HGNC: 3585] piccolo presynaptic cytomatrix protein [Source: HGNC Symbol; Acc: 2.90 PCLO 27445 PCLO HGNC: 13406] erb-b2 receptor tyrosine kinase 2 [Source: HGNC Symbol; Acc: 2.44 ERBB2 2064 ERBB2 HGNC: 3430] BRCA1 associated protein 1 [Source: HGNC Symbol; Acc: HGNC: 2.35 BAP1 8314 BAP1 950] capicua transcriptional repressor [Source: HGNC Symbol; Acc: 2.35 CIC 23152 CIC HGNC: -

Rap1a and Rap1b Ras-Family Proteins Are Prominently Expressed in the Nucleus of Squamous Carcinomas: Nuclear Translocation of GTP-Bound Active Form
Oncogene (2003) 22, 6243–6256 & 2003 Nature Publishing Group All rights reserved 0950-9232/03 $25.00 www.nature.com/onc Rap1A and rap1B ras-family proteins are prominently expressed in the nucleus of squamous carcinomas: nuclear translocation of GTP-bound active form Raj S Mitra1,4, Zhaocheng Zhang1,4, Bradley S Henson1, David M Kurnit2, Thomas ECarey 1,3, Nisha J D’Silva*,1 1Department of Oral Medicine, Pathology and Oncology, University of Michigan School of Dentistry, Ann Arbor, MI 48109-1078, USA; 2Departments of Pediatrics and Human Genetics, University of Michigan Medical School, Ann Arbor, MI 481091, USA; 3Department of Otolaryngology, Laboratory of Head and Neck Cancer Biology, The University of Michigan Medical School and the Comprehensive Cancer Center, Ann Arbor, MI 48109-0506, USA We recently showed that rap1 regulates growth and exclusively in eukaryotes (Takai et al., 2001). Those proliferation in normal keratinocytes, which provoked us SMGs that have been identified (currently 4100) have to investigate its expression and regulation in malignant homology to ras, and belong to one of the five cells. Rap1 is variably expressed in whole cell lysates of subfamilies namely ras, rho, rab, sar1/arf and ran squamous cell carcinoma (SCC) cell lines. Immunoblot (Takai et al., 2001). These ras-like proteins shuttle analysis of nuclear and cytosolic fractions and immuno- between inactive GDP- and active GTP-bound states. histochemistry revealed that in addition to cytoplasmic Ras subfamily proteins, including ras and rap1, are key expression, SCC cells also exhibit prominent punctate players in receptor-linked signaling pathways that rap1 expression in the nucleus.