The Application of a Characterized Pre-Clinical
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Options for the Treatment of Gemcitabine-Resistant Advanced Pancreatic Cancer
JOP. J Pancreas (Online) 2010 Mar 5; 11(2):113-123. REVIEW Options for the Treatment of Gemcitabine-Resistant Advanced Pancreatic Cancer Ioannis Gounaris, Kamarul Zaki, Pippa Corrie Oncology Centre, Cambridge University Hospitals NHS Trust. Cambridge, United Kingdom Summary Context Pancreatic cancer is noteworthy in that the number of patients dying from the disease is roughly equal to the number diagnosed. For more than a decade, gemcitabine has constituted the standard of care for the palliative treatment of the majority of patients who present with metastatic or relapsed disease, although the survival gains are limited. Despite a median survival of less than 6 months, there is a significant proportion of advanced pancreatic cancer patients who progress on gemcitabine that remains fit and these patients are candidates for second-line treatment. Methods The OVID MEDLINE database was searched from 1950 to present using the MeSH terms “pancreatic neoplasms”, “drug treatment” and “gemcitabine”. After excluding non-relevant results, 31 published studies were identified. These results were supplemented by searching the last three (2007-2009) American Society of Clinical Oncology (ASCO) Proceedings of Annual Meetings for studies published only in abstract form and reviewing reference lists of published articles. Results and discussion The evidence for second line treatments of metastatic pancreatic cancer consists mostly of single arm, small phase II studies. Oxaliplatin-fluoropyrimidine combinations appear promising and have shown increased survival compared to best supportive care. As the molecular pathways governing pancreatic cancer are unravelled, novel targeted therapies may offer the greatest promise for this disease either given alone, combined with one another, or with cytotoxic agents. -

Study Protocol and Statistical Analysis Plan
Confidential Clinical study protocol number: J1228 Page 1 Version Date: May 7, 2018 IRB study Number: NA_00067315 A Trial of maintenance Rituximab with mTor inhibition after High-dose Consolidative Therapy in CD20+, B-cell Lymphomas, Gray Zone Lymphoma, and Hodgkin’s Lymphoma Principal Investigator: Douglas E. Gladstone, MD The Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins 1650 Orleans Street, CRBI-287 Baltimore, MD 21287 Phone: 410-955-8781 Fax: 410-614-1005 Email: [email protected] IRB Protocol Number: NA_00067315 Study Number: J1228 IND Number: EXEMPT Novartis Protocol Number: CRAD001NUS157T Version: May 7, 2018 Co-Investigators: Jonathan Powell 1650 Orleans Street, CRBI-443 Phone: 410-502-7887 Fax: 443-287-4653 Email: [email protected] Richard Jones 1650 Orleans Street, CRBI-244 Phone: 410-955-2006 Fax: 410-614-7279 Email: [email protected] Confidential Clinical study protocol number: J1228 Page 2 Version Date: May 7, 2018 IRB study Number: NA_00067315 Satish Shanbhag Johns Hopkins Bayview Medical Center 301 Building, Suite 4500 4940 Eastern Ave Phone: 410-550-4061 Fax: 410-550-5445 Email: [email protected] Statisticians: Gary Rosner Phone: 410-955-4884 Email: [email protected] Marianna Zahurak Phone: 410-955-4219 Email: [email protected] Confidential Clinical study protocol number: J1228 Page 3 Version Date: May 7, 2018 IRB study Number: NA_00067315 Table of contents Table of contents ......................................................................................................................... 3 List of abbreviations -

LEUKEMIA CHEMOTHERAPY REGIMENS (Part 1 of 2) the Selection, Dosing, and Administration of Anti-Cancer Agents and the Management of Associated Toxicities Are Complex
LEUKEMIA CHEMOTHERAPY REGIMENS (Part 1 of 2) The selection, dosing, and administration of anti-cancer agents and the management of associated toxicities are complex. Drug dose modifications and schedule and initiation of supportive care interventions are often necessary because of expected toxicities and because of individual patient variability, prior treatment, and comorbidities. Thus, the optimal delivery of anti-cancer agents requires a healthcare delivery team experienced in the use of such agents and the management of associated toxicities in patients with cancer. The chemotherapy regimens below may include both FDA-approved and unapproved uses/regimens and are provided as references only to the latest treatment strategies. Clinicians must choose and verify treatment options based on the individual patient. REGIMEN DOSING Acute Myeloid Leukemia (AML) Induction Therapy Cytarabine (Cytosar-U; ARA-C) + Days 1–3: An anthracycline (eg, daunorubicin at least 60mg/m2/day IV, an anthracycline idarubicin 10–12mg/m2/day IV, or mitoxantrone 10–12mg/m2/day IV), plus (daunorubicin [Cerubidine], Days 1–7: Cytarabine 100–200mg/m2/day continuous IV infusion. idarubicin [Idamycin], OR mitoxantrone [Novantrone])1, 2 Days 1–3: An anthracycline (eg, daunorubicin 45mg/m2/day IV, idarubicin 12mg/m2/day IV, or mitoxantrone 12mg/m2/day IV), plus Days 1–7: Cytarabine 100mg/m2/day continuous IV infusion. Intermediate-dose cytarabine3 Cycle 1 Days 1–7: Cytarabine 200mg/m2/day continuous IV infusion, plus Days 5–6: Idarubicin 12mg/m2/day IV. Cycle 2 Days 1–6: Cytarabine 1,000mg/m2 continuous IV infusion for 3 hrs twice daily, plus Days 3, 5 and 7: Amsacrine 120mg/m2/day. -

LEUSTATIN (Cladribine) Injection Should Be Administered Under the Supervision of a Qualified Physician Experienced in the Use of Antineoplastic Therapy
LEUSTATIN (cladribine) Injection For Intravenous Infusion Only WARNING LEUSTATIN (cladribine) Injection should be administered under the supervision of a qualified physician experienced in the use of antineoplastic therapy. Suppression of bone marrow function should be anticipated. This is usually reversible and appears to be dose dependent. Serious neurological toxicity (including irreversible paraparesis and quadraparesis) has been reported in patients who received LEUSTATIN Injection by continuous infusion at high doses (4 to 9 times the recommended dose for Hairy Cell Leukemia). Neurologic toxicity appears to demonstrate a dose relationship; however, severe neurological toxicity has been reported rarely following treatment with standard cladribine dosing regimens. Acute nephrotoxicity has been observed with high doses of LEUSTATIN (4 to 9 times the recommended dose for Hairy Cell Leukemia), especially when given concomitantly with other nephrotoxic agents/therapies. DESCRIPTION LEUSTATIN (cladribine) Injection (also commonly known as 2-chloro-2΄-deoxy- β -D-adenosine) is a synthetic antineoplastic agent for continuous intravenous infusion. It is a clear, colorless, sterile, preservative-free, isotonic solution. LEUSTATIN Injection is available in single-use vials containing 10 mg (1 mg/mL) of cladribine, a chlorinated purine nucleoside analog. Each milliliter of LEUSTATIN Injection contains 1 mg of the active ingredient and 9 mg (0.15 mEq) of sodium chloride as an inactive ingredient. The solution has a pH range of 5.5 to 8.0. Phosphoric -

Synthesis of Ribavirin, Tecadenoson, and Cladribine by Enzymatic Transglycosylation
catalysts Article Synthesis of Ribavirin, Tecadenoson, and Cladribine by Enzymatic Transglycosylation 1, 2 2,3 2 Marco Rabuffetti y, Teodora Bavaro , Riccardo Semproli , Giulia Cattaneo , Michela Massone 1, Carlo F. Morelli 1 , Giovanna Speranza 1,* and Daniela Ubiali 2,* 1 Department of Chemistry, University of Milan, via Golgi 19, I-20133 Milano, Italy; marco.rabuff[email protected] (M.R.); [email protected] (M.M.); [email protected] (C.F.M.) 2 Department of Drug Sciences, University of Pavia, viale Taramelli 12, I-27100 Pavia, Italy; [email protected] (T.B.); [email protected] (R.S.); [email protected] (G.C.) 3 Consorzio Italbiotec, via Fantoli 15/16, c/o Polo Multimedica, I-20138 Milano, Italy * Correspondence: [email protected] (G.S.); [email protected] (D.U.); Tel.: +39-02-50314097 (G.S.); +39-0382-987889 (D.U.) Present address: Department of Food, Environmental and Nutritional Sciences, University of Milan, y via Mangiagalli 25, I-20133 Milano, Italy. Received: 7 March 2019; Accepted: 8 April 2019; Published: 12 April 2019 Abstract: Despite the impressive progress in nucleoside chemistry to date, the synthesis of nucleoside analogues is still a challenge. Chemoenzymatic synthesis has been proven to overcome most of the constraints of conventional nucleoside chemistry. A purine nucleoside phosphorylase from Aeromonas hydrophila (AhPNP) has been used herein to catalyze the synthesis of Ribavirin, Tecadenoson, and Cladribine, by a “one-pot, one-enzyme” transglycosylation, which is the transfer of the carbohydrate moiety from a nucleoside donor to a heterocyclic base. As the sugar donor, 7-methylguanosine iodide and its 20-deoxy counterpart were synthesized and incubated either with the “purine-like” base or the modified purine of the three selected APIs. -

(DAC) Followed by Clofarabine, Idarubicin, and Cytarabine (CIA) in Acute Leukemia 2012-1064
2012-1064 September 02, 2014 Page 1 Protocol Page Phase I/II Study of Decitabine (DAC) followed by Clofarabine, Idarubicin, and Cytarabine (CIA) in Acute Leukemia 2012-1064 Core Protocol Information Short Title Decitabine followed by Clofarabine, Idarubicin, and Cytarabine in Acute Leukemia Study Chair: Nitin Jain Additional Contact: Allison Pike Jeannice Y. Theriot Leukemia Protocol Review Group Department: Leukemia Phone: 713-745-6080 Unit: 428 Full Title: Phase I/II Study of Decitabine (DAC) followed by Clofarabine, Idarubicin, and Cytarabine (CIA) in Acute Leukemia Protocol Type: Standard Protocol Protocol Phase: Phase I/Phase II Version Status: Terminated 01/12/2018 Version: 12 Submitted by: Jeannice Y. Theriot--4/26/2017 2:13:38 PM OPR Action: Accepted by: Melinda E. Gordon -- 5/1/2017 7:55:15 AM Which Committee will review this protocol? The Clinical Research Committee - (CRC) 2012-1064 September 02, 2014 Page 2 Protocol Body Phase I/II Study of Decitabine (DAC) followed by Clofarabine, Idarubicin, and Cytarabine (CIA) in Acute Leukemia 1. OBJECTIVES Phase I Primary: To determine the maximal tolerated dose (MTD) of clofarabine to be used in portion II of the study Phase II Primary: To determine the response rate of the DAC-CIA regimen Secondary: A) To determine the toxicity of the combination regimen B) To determine the disease-free survival (DFS) and overall survival (OS) rates 2. RATIONALE 2.1 Acute Myelogenous Leukemia Acute myelogenous leukemia (AML) is the most common acute leukemia in adults. It is estimated that 13,780 men and women will be diagnosed with and 10,200 men and women will die of acute myeloid leukemia in the year 2012.1 AML is a disease with a poor prognosis with a 5-year survival of only around 30%.2,3 Certain subgroups of AML have a particularly worse Page 1 of 34 outcome such as patients with relapsed and/or refractory AML and AML arising from antecedent myelodysplastic syndrome (MDS) or myeloproliferative neoplasms (MPNs). -

Drug Name Plate Number Well Location % Inhibition, Screen Axitinib 1 1 20 Gefitinib (ZD1839) 1 2 70 Sorafenib Tosylate 1 3 21 Cr
Drug Name Plate Number Well Location % Inhibition, Screen Axitinib 1 1 20 Gefitinib (ZD1839) 1 2 70 Sorafenib Tosylate 1 3 21 Crizotinib (PF-02341066) 1 4 55 Docetaxel 1 5 98 Anastrozole 1 6 25 Cladribine 1 7 23 Methotrexate 1 8 -187 Letrozole 1 9 65 Entecavir Hydrate 1 10 48 Roxadustat (FG-4592) 1 11 19 Imatinib Mesylate (STI571) 1 12 0 Sunitinib Malate 1 13 34 Vismodegib (GDC-0449) 1 14 64 Paclitaxel 1 15 89 Aprepitant 1 16 94 Decitabine 1 17 -79 Bendamustine HCl 1 18 19 Temozolomide 1 19 -111 Nepafenac 1 20 24 Nintedanib (BIBF 1120) 1 21 -43 Lapatinib (GW-572016) Ditosylate 1 22 88 Temsirolimus (CCI-779, NSC 683864) 1 23 96 Belinostat (PXD101) 1 24 46 Capecitabine 1 25 19 Bicalutamide 1 26 83 Dutasteride 1 27 68 Epirubicin HCl 1 28 -59 Tamoxifen 1 29 30 Rufinamide 1 30 96 Afatinib (BIBW2992) 1 31 -54 Lenalidomide (CC-5013) 1 32 19 Vorinostat (SAHA, MK0683) 1 33 38 Rucaparib (AG-014699,PF-01367338) phosphate1 34 14 Lenvatinib (E7080) 1 35 80 Fulvestrant 1 36 76 Melatonin 1 37 15 Etoposide 1 38 -69 Vincristine sulfate 1 39 61 Posaconazole 1 40 97 Bortezomib (PS-341) 1 41 71 Panobinostat (LBH589) 1 42 41 Entinostat (MS-275) 1 43 26 Cabozantinib (XL184, BMS-907351) 1 44 79 Valproic acid sodium salt (Sodium valproate) 1 45 7 Raltitrexed 1 46 39 Bisoprolol fumarate 1 47 -23 Raloxifene HCl 1 48 97 Agomelatine 1 49 35 Prasugrel 1 50 -24 Bosutinib (SKI-606) 1 51 85 Nilotinib (AMN-107) 1 52 99 Enzastaurin (LY317615) 1 53 -12 Everolimus (RAD001) 1 54 94 Regorafenib (BAY 73-4506) 1 55 24 Thalidomide 1 56 40 Tivozanib (AV-951) 1 57 86 Fludarabine -

COMPARISON of the WHO ATC CLASSIFICATION & Ephmra/Intellus Worldwide ANATOMICAL CLASSIFICATION
COMPARISON OF THE WHO ATC CLASSIFICATION & EphMRA/Intellus Worldwide ANATOMICAL CLASSIFICATION: VERSION June 2019 2 Comparison of the WHO ATC Classification and EphMRA / Intellus Worldwide Anatomical Classification The following booklet is designed to improve the understanding of the two classification systems. The development of the two systems had previously taken place separately. EphMRA and WHO are now working together to ensure that there is a convergence of the 2 systems rather than a divergence. In order to better understand the two classification systems, we should pay attention to the way in which substances/products are classified. WHO mainly classifies substances according to the therapeutic or pharmaceutical aspects and in one class only (particular formulations or strengths can be given separate codes, e.g. clonidine in C02A as antihypertensive agent, N02C as anti-migraine product and S01E as ophthalmic product). EphMRA classifies products, mainly according to their indications and use. Therefore, it is possible to find the same compound in several classes, depending on the product, e.g., NAPROXEN tablets can be classified in M1A (antirheumatic), N2B (analgesic) and G2C if indicated for gynaecological conditions only. The purposes of classification are also different: The main purpose of the WHO classification is for international drug utilisation research and for adverse drug reaction monitoring. This classification is recommended by the WHO for use in international drug utilisation research. The EphMRA/Intellus Worldwide classification has a primary objective to satisfy the marketing needs of the pharmaceutical companies. Therefore, a direct comparison is sometimes difficult due to the different nature and purpose of the two systems. -

Recent Topics on the Mechanisms of Immunosuppressive Therapy-Related Neurotoxicities
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 29 May 2019 doi:10.20944/preprints201905.0358.v1 Peer-reviewed version available at Int. J. Mol. Sci. 2019, 20, 3210; doi:10.3390/ijms20133210 1 of 39 1 Review 2 Recent topics on the mechanisms of 3 immunosuppressive therapy-related neurotoxicities 4 Wei Zhang 1, Nobuaki Egashira 1,2,* and Satohiro Masuda 1,2 5 1 Department of Clinical Pharmacology and Biopharmaceutics, Graduate School of Pharmaceutical Sciences, 6 Kyushu University, Fukuoka 812-8582, Japan 7 2 Department of Pharmacy, Kyushu University Hospital, Fukuoka 812-8582, Japan 8 * Correspondence: [email protected], Tel.: 81-92-642-5920 9 Abstract: Although transplantation procedures have been developed for patients with end-stagec 10 hepatic insufficiency or other diseases, allograft rejection still threatens patient health and lifespan. 11 Over the last few decades, the emergence of immunosuppressive agents, such as calcineurin 12 inhibitors (CNIs) and mammalian target of rapamycin (mTOR) inhibitors, have strikingly 13 increased graft survival. Unfortunately, immunosuppressive agent-related neurotoxicity is 14 commonly occurred in clinical situations, with the majority of neurotoxicity cases caused by CNIs. 15 The possible mechanisms whereby CNIs cause neurotoxicity include: increasing the permeability 16 or injury of the blood-brain barrier, alterations of mitochondrial function, and alterations in 17 electrophysiological state. Other immunosuppressants can also induce neuropsychiatric 18 complications. For example, mTOR inhibitors induce seizures; mycophenolate mofetil induces 19 depression and headache; methotrexate affects the central nervous system; mouse monoclonal 20 immunoglobulin G2 antibody against cluster of differentiation 3 also induces headache; and 21 patients using corticosteroids usually experience cognitive alteration. -

Corporate Overview May 2017
Corporate Overview May 2017 DEVELOPING PRECISION MEDICINES TO TREAT CANCER Forward Looking Statements This presentation contains forward-looking statements. Such statements include, but are not limited to, statements regarding our research, pre-clinical and clinical development activities, plans and projected timelines for tipifarnib, KO- 947 and KO-539, plans regarding regulatory filings, our expectations regarding the relative benefits of our product candidates versus competitive therapies, and our expectations regarding the therapeutic and commercial potential of our product candidates. The words “believe,” “may,” “will,” “estimate,” “promise,” “plan”, “continue,” “anticipate,” “intend,” “expect,” “potential” and similar expressions (including the negative thereof), are intended to identify forward-looking statements. Because such statements are subject to risks and uncertainties, actual results may differ materially from those expressed or implied by such forward-looking statements. Risks that contribute to the uncertain nature of the forward-looking statements include: our future preclinical studies and clinical trials may not be successful; the U.S. Food and Drug Administration (FDA) may not agree with our interpretation of the data from clinical trials of our product candidates; we may decide, or the FDA may require us, to conduct additional clinical trials or to modify our ongoing clinical trials; we may experience delays in the commencement, enrollment, completion or analysis of clinical testing for our product candidates, or significant issues regarding the adequacy of our clinical trial designs or the execution of our clinical trials may arise, which could result in increased costs and delays, or limit our ability to obtain regulatory approval; our product candidates may not receive regulatory approval or be successfully commercialized; unexpected adverse side effects or inadequate therapeutic efficacy of our product candidates could delay or prevent regulatory approval or commercialization; we may not be able to obtain additional financing. -

(12) United States Patent (10) Patent No.: US 8,911,786 B2 Desai Et Al
US00891. 1786B2 (12) United States Patent (10) Patent No.: US 8,911,786 B2 Desai et al. (45) Date of Patent: Dec. 16, 2014 (54) NANOPARTICLE COMPRISING RAPAMYCIN A61K 45/06 (2013.01); A61K 47/42 (2013.01); AND ALBUMINAS ANTICANCERAGENT A61N 5/10 (2013.01); A61N 7700 (2013.01) (75) Inventors: Neil P. Desai, Los Angeles, CA (US); USPC ............ 424/491; 424/489: 424/490; 424/500 Patrick Soon-Shiong, Los Angeles, CA (58) Field of Classification Search (US); Vuong Trieu, Calabasas, CA (US) USPC .......... 424/465-489, 490, 491, 500: 514/19.3 See application file for complete search history. (73) Assignee: Abraxis Bioscience, LLC, Los Angeles, CA (US) (56) References Cited (*) Notice: Subject to any disclaimer, the term of this patent is extended or adjusted under 35 U.S. PATENT DOCUMENTS U.S.C. 154(b) by 344 days. 5,206,018 A * 4/1993 Sehgal et al. ................. 424,122 5,362.478 A 11/1994 Desai et al. (21) Appl. No.: 12/530,188 5.439,686 A 8, 1995 Desai et al. 5,498.421 A 3, 1996 Grinstaffet al. (22) PCT Filed: Mar. 7, 2008 5,505,932 A 4/1996 Grinstaffet al. 5,508,021 A 4/1996 Grinstaffet al. (86). PCT No.: PCT/US2O08/OO3O96 5,512,268 A 4/1996 Grinstaffet al. 5,540,931 A 7/1996 Hewitt et al. S371 (c)(1), 5,560,933 A 10/1996 Soon-Shiong et al. (2), (4) Date: Mar. 4, 2010 5,635,207 A 6/1997 Grinstaffet al. 5,639,473 A 6/1997 Grinstaffet al. -
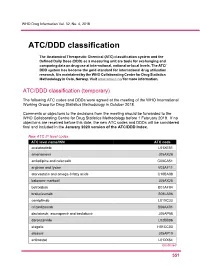
ATC/DDD Classification
WHO Drug Information Vol. 32, No. 4, 2018 ATC/DDD classification The Anatomical Therapeutic Chemical (ATC) classification system and the Defined Daily Dose (DDD) as a measuring unit are tools for exchanging and comparing data on drug use at international, national or local levels. The ATC/ DDD system has become the gold standard for international drug utilization research. It is maintained by the WHO Collaborating Centre for Drug Statistics Methodology in Oslo, Norway. Visit www.whocc.no/ for more information. ATC/DDD classification (temporary) The following ATC codes and DDDs were agreed at the meeting of the WHO International Working Group for Drug Statistics Methodology in October 2018. Comments or objections to the decisions from the meeting should be forwarded to the WHO Collaborating Centre for Drug Statistics Methodology before 1 February 2019. If no objections are received before this date, the new ATC codes and DDDs will be considered final and included in the January 2020 version of the ATC/DDD Index. New ATC 5th level codes ATC level name/INN ATC code acalabrutinib L01XE51 amenamevir J05AX26 amlodipine and celecoxib C08CA51 arginine and lysine V03AF11 atorvastatin and omega-3 fatty acids C10BA08 baloxavir marboxil J05AX25 betrixaban B01AF04 brolucizumab S01LA06 cemiplimab L01XC33 crizanlizumab B06AX01 daclatasvir, asunaprevir and beclabuvir J05AP58 darolutamide L02BB06 elagolix H01CC03 elbasvir J05AP10 entinostat L01XX64 Continued 5 51 ATC/DDD classification WHO Drug Information Vol. 32, No. 4, 2018 New ATC 5th level codes (continued)