Cdna Microarray Analysis and Influx Transporter OATP1B1 in Liver Cells After Exposure to Gadoxetate Disodium, a Gadolinium-Based
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Mitochondrial Dynamics During Spermatogenesis Grigor Varuzhanyan and David C
© 2020. Published by The Company of Biologists Ltd | Journal of Cell Science (2020) 133, jcs235937. doi:10.1242/jcs.235937 REVIEW SUBJECT COLLECTION: MITOCHONDRIA Mitochondrial dynamics during spermatogenesis Grigor Varuzhanyan and David C. Chan* ABSTRACT Mitophagy is an additional layer of quality control that utilizes Mitochondrial fusion and fission (mitochondrial dynamics) are autophagy (Mizushima, 2007; Mizushima et al., 1998; Tsukada and homeostatic processes that safeguard normal cellular function. This Ohsumi, 1993) to remove excessive mitochondria or damaged relationship is especially strong in tissues with constitutively high energy mitochondria that are beyond repair (Pickles et al., 2018). During demands, such as brain, heart and skeletal muscle. Less is known about mitophagy, microtubule-associated protein 1A/1B light chain 3 the role of mitochondrial dynamics in developmental systems that involve (collectively LC3; also known as MAP1LC3A, MAP1LC3B and changes in metabolic function. One such system is spermatogenesis. MAP1LC3C) is recruited to the autophagosomal membrane and binds The first mitochondrial dynamics gene, Fuzzy onions (Fzo), was to mitochondria that selectively express mitophagy receptors on their discovered in 1997 to mediate mitochondrial fusion during Drosophila outer membrane. This molecular recognition designates damaged spermatogenesis. In mammals, however, the role of mitochondrial fusion mitochondria as cargo for autophagosomes, and the resulting during spermatogenesis remained unknown for nearly two decades after mitophagosomes subsequently fuse with lysosomes for degradation discovery of Fzo. Mammalian spermatogenesis is one of the most and recycling of the engulfed organelles. Much of the molecular complex and lengthy differentiation processes in biology, transforming workings of mitophagy have been parsed in cultured cells, and spermatogonial stem cells into highly specialized sperm cells over a recently developed mouse models are enabling the analysis of 5-week period. -

Small Cell Carcinoma of the Ovary, Hypercalcemic Type (SCCOHT) Beyond SMARCA4 Mutations: a Comprehensive Genomic Analysis
cells Article Small Cell Carcinoma of the Ovary, Hypercalcemic Type (SCCOHT) beyond SMARCA4 Mutations: A Comprehensive Genomic Analysis Aurélie Auguste 1,Félix Blanc-Durand 2 , Marc Deloger 3 , Audrey Le Formal 1, Rohan Bareja 4,5, David C. Wilkes 4 , Catherine Richon 6,Béatrice Brunn 2, Olivier Caron 6, Mojgan Devouassoux-Shisheboran 7,Sébastien Gouy 2, Philippe Morice 2, Enrica Bentivegna 2, Andrea Sboner 4,5,8, Olivier Elemento 4,8, Mark A. Rubin 9 , Patricia Pautier 2, Catherine Genestie 10, Joanna Cyrta 4,9,11 and Alexandra Leary 1,2,* 1 Medical Oncologist, Gynecology Unit, Lead Translational Research Team, INSERM U981, Gustave Roussy, 94805 Villejuif, France; [email protected] (A.A.); [email protected] (A.L.F.) 2 Gynecological Unit, Department of Medicine, Gustave Roussy, 94805 Villejuif, France; [email protected] (F.B.-D.); [email protected] (B.B.); [email protected] (S.G.); [email protected] (P.M.); [email protected] (E.B.); [email protected] (P.P.) 3 Bioinformatics Core Facility, Gustave Roussy Cancer Center, UMS CNRS 3655/INSERM 23 AMMICA, 94805 Villejuif, France; [email protected] 4 Caryl and Israel Englander Institute for Precision Medicine, Weill Cornell Medicine, New York, NY 10001, USA; [email protected] (R.B.); [email protected] (D.C.W.); [email protected] (A.S.); [email protected] (O.E.); [email protected] (J.C.) 5 Institute for Computational Biomedicine, Weill Cornell -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -
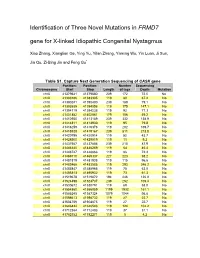
Identification of Three Novel Mutations in FRMD7 Gene for X-Linked Idiopathic Congenital Nystagmus
Identification of Three Novel Mutations in FRMD7 gene for X-linked Idiopathic Congenital Nystagmus Xiao Zhang, Xianglian Ge, Ying Yu, Yilan Zhang, Yaming Wu, Yin Luan, Ji Sun, Jia Qu, Zi-Bing Jin and Feng Gu* Table S1. Capture Next Generation Sequencing of CASK gene Position: Position: Number Sequencing Chromosome Start Stop Length of tags Depth Mutation chrX 41379641 41379880 239 172 72.0 No chrX 41383186 41383305 119 80 67.2 No chrX 41390241 41390480 239 189 79.1 No chrX 41393939 41394058 119 175 147.1 No chrX 41394119 41394238 119 92 77.3 No chrX 41401882 41402061 179 106 59.2 No chrX 41412950 41413189 239 332 138.9 No chrX 41414811 41414930 119 95 79.8 No chrX 41416259 41416378 119 202 169.7 No chrX 41418928 41419167 239 511 213.8 No chrX 41420795 41420914 119 52 43.7 No chrX 41428900 41429019 119 11 9.2 No chrX 41437567 41437806 239 210 87.9 No chrX 41446140 41446259 119 54 45.4 No chrX 41448747 41448866 119 86 72.3 No chrX 41469110 41469337 227 223 98.2 No chrX 41481819 41481938 119 115 96.6 No chrX 41483466 41483585 119 293 246.2 No chrX 41485847 41485966 119 75 63.0 No chrX 41495813 41495932 119 73 61.3 No chrX 41519678 41519872 194 246 126.8 No chrX 41524498 41524737 239 252 105.4 No chrX 41530672 41530791 119 69 58.0 No chrX 41554860 41556059 1199 1932 161.1 No chrX 41586245 41587324 1079 1044 96.8 No chrX 41598613 41598732 119 27 22.7 No chrX 41604756 41604875 119 27 22.7 No chrX 41646424 41646543 119 124 104.2 No chrX 41712364 41712483 119 37 31.1 No chrX 41782152 41782271 119 5 4.2 No Table S2. -

Genome-Wide Analysis of DNA Methylation, Copy Number Variation, and Gene Expression in Monozygotic Twins Discordant for Primary Biliary Cirrhosis
UC Davis UC Davis Previously Published Works Title Genome-wide analysis of DNA methylation, copy number variation, and gene expression in monozygotic twins discordant for primary biliary cirrhosis. Permalink https://escholarship.org/uc/item/34d4m5nk Journal Frontiers in immunology, 5(MAR) ISSN 1664-3224 Authors Selmi, Carlo Cavaciocchi, Francesca Lleo, Ana et al. Publication Date 2014 DOI 10.3389/fimmu.2014.00128 Peer reviewed eScholarship.org Powered by the California Digital Library University of California ORIGINAL RESEARCH ARTICLE published: 28 March 2014 doi: 10.3389/fimmu.2014.00128 Genome-wide analysis of DNA methylation, copy number variation, and gene expression in monozygotic twins discordant for primary biliary cirrhosis Carlo Selmi 1,2*, Francesca Cavaciocchi 1,3, Ana Lleo4, Cristina Cheroni 5, Raffaele De Francesco5, Simone A. Lombardi 1, Maria De Santis 1,3, Francesca Meda1, Maria Gabriella Raimondo1, Chiara Crotti 1, Marco Folci 1, Luca Zammataro1, Marlyn J. Mayo6, Nancy Bach7, Shinji Shimoda8, Stuart C. Gordon9, Monica Miozzo10,11, Pietro Invernizzi 4, Mauro Podda1, Rossana Scavelli 5, Michelle R. Martin12, Michael F. Seldin13,14, Janine M. LaSalle 12 and M. Eric Gershwin2 1 Division of Rheumatology and Clinical Immunology, Humanitas Clinical and Research Center, Milan, Italy 2 Division of Rheumatology, Allergy, and Clinical Immunology, University of California at Davis, Davis, CA, USA 3 BIOMETRA Department, University of Milan, Milan, Italy 4 Liver Unit and Center for Autoimmune Liver Diseases, Humanitas Clinical and Research Center, Milan, Italy 5 National Institute of Molecular Genetics (INGM), Milan, Italy 6 University of Texas Southwestern, Dallas, TX, USA 7 Mt. Sinai University, NewYork, NY, USA 8 Clinical Research Center, National Nagasaki Medical Center, Nagasaki, Japan 9 Henry Ford Hospital, Detroit, MI, USA 10 Department of Pathophysiology and Transplantation, University of Milan, Milan, Italy 11 Division of Pathology, Fondazione IRCCS Cà Granda Ospedale Maggiore Policlinico, Milan, Italy 12 Genome Center and M.I.N.D. -

Genome-Wide Analysis of Cancer/Testis Gene Expression
Genome-wide analysis of cancer/testis gene expression Oliver Hofmanna,b,1, Otavia L. Caballeroc, Brian J. Stevensond,e, Yao-Tseng Chenf, Tzeela Cohenc, Ramon Chuac, Christopher A. Maherb, Sumir Panjib, Ulf Schaeferb, Adele Krugerb, Minna Lehvaslaihob, Piero Carnincig,h, Yoshihide Hayashizakig,h, C. Victor Jongeneeld,e, Andrew J. G. Simpsonc, Lloyd J. Oldc,1, and Winston Hidea,b aDepartment of Biostatistics, Harvard School of Public Health, 655 Huntington Avenue, SPH2, 4th Floor, Boston, MA 02115; bSouth African National Bioinformatics Institute, University of the Western Cape, Private Bag X17, Bellville 7535, South Africa; cLudwig Institute for Cancer Research, New York Branch at Memorial Sloan-Kettering Cancer Center, 1275 York Avenue, New York, NY 10021; dLudwig Institute for Cancer Research, Lausanne Branch, 1015 Lausanne, Switzerland; eSwiss Institute of Bioinformatics, 1015 Lausanne, Switzerland; fWeill Medical College of Cornell University, 1300 York Avenue, New York, NY 10021; gGenome Exploration Research Group (Genome Network Project Core Group), RIKEN Genomic Sciences Center (GSC), RIKEN Yokohama Institute, 1-7-22 Suehiro-cho, Tsurumi-ku, Yokohama, Kanagawa, 230-0045, Japan; and hGenome Science Laboratory, Discovery Research Institute, RIKEN Wako Institute, 2-1 Hirosawa, Wako, Saitama, 3510198, Japan Contributed by Lloyd J. Old, October 28, 2008 (sent for review June 6, 2008) Cancer/Testis (CT) genes, normally expressed in germ line cells but expression profile information frequently limited to the original also activated in a wide range of cancer types, often encode defining articles. In some cases, e.g., ACRBP, the original antigens that are immunogenic in cancer patients, and present CT-restricted expression in normal tissues could not be con- potential for use as biomarkers and targets for immunotherapy. -

Functional Annotation of the Human Retinal Pigment Epithelium
BMC Genomics BioMed Central Research article Open Access Functional annotation of the human retinal pigment epithelium transcriptome Judith C Booij1, Simone van Soest1, Sigrid MA Swagemakers2,3, Anke HW Essing1, Annemieke JMH Verkerk2, Peter J van der Spek2, Theo GMF Gorgels1 and Arthur AB Bergen*1,4 Address: 1Department of Molecular Ophthalmogenetics, Netherlands Institute for Neuroscience (NIN), an institute of the Royal Netherlands Academy of Arts and Sciences (KNAW), Meibergdreef 47, 1105 BA Amsterdam, the Netherlands (NL), 2Department of Bioinformatics, Erasmus Medical Center, 3015 GE Rotterdam, the Netherlands, 3Department of Genetics, Erasmus Medical Center, 3015 GE Rotterdam, the Netherlands and 4Department of Clinical Genetics, Academic Medical Centre Amsterdam, the Netherlands Email: Judith C Booij - [email protected]; Simone van Soest - [email protected]; Sigrid MA Swagemakers - [email protected]; Anke HW Essing - [email protected]; Annemieke JMH Verkerk - [email protected]; Peter J van der Spek - [email protected]; Theo GMF Gorgels - [email protected]; Arthur AB Bergen* - [email protected] * Corresponding author Published: 20 April 2009 Received: 10 July 2008 Accepted: 20 April 2009 BMC Genomics 2009, 10:164 doi:10.1186/1471-2164-10-164 This article is available from: http://www.biomedcentral.com/1471-2164/10/164 © 2009 Booij et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. -

(51) International Patent Classification: A61P 35/00 (2006.01) G01N 33/50
( (51) International Patent Classification: Published: A61P 35/00 (2006.01) G01N 33/50 (2006.01) — with international search report (Art. 21(3)) G16H 50/20 (2018.01) A61K 39/00 (2006.01) G16H 50/30 (2018.01) (21) International Application Number: PCT/US20 19/058578 (22) International Filing Date: 29 October 2019 (29. 10.2019) (25) Filing Language: English (26) Publication Language: English (30) Priority Data: 62/752,288 29 October 2018 (29. 10.2018) US 62/788,3 13 04 January 2019 (04.01.2019) US 62/855,332 31 May 2019 (3 1.05.2019) US (71) Applicant: GENOCEA BIOSCIENCES, INC. [US/US]; Cambridge Discovery Park, 100 Acorn Park Drive, 5th Floor, Cambridge, Massachusetts 02140 (US). (72) Inventors: FLECHTNER, Jessica Baker; 1 Briant Drive, Sudbury, Massachusetts 01776 (US). DOBSON, Jason R.; 44 Pole Plain Road, Sharon, Massachusetts 02067 (US). (74) Agent: DAUKSS, Dana M. et al.; Choate, Hall & Ste¬ wart LLP, Two International Place, Boston, Massachusetts 021 10 (US). (81) Designated States (unless otherwise indicated, for every kind of national protection available) : AE, AG, AL, AM, AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IR, IS, JO, JP, KE, KG, KH, KN, KP, KR, KW, KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. -

A Comprehensive Analysis of the MAGE Family As Prognostic and Diagnostic Markers for Hepatocellular Carcinoma
A Comprehensive Analysis of the MAGE Family as Prognostic and Diagnostic Markers for Hepatocellular Carcinoma Rong Li Guangdong Provincial Key Laboratory of Liver Disease Research, Guangdong province engineering laboratory for transplantation medicine, Third Aliated Hospital of Sun Yat-sen University Jiao Gong Department of Laboratory Medicine, Third Aliated Hospital of Sun Yat-sen University Cuicui Xiao Department of Anesthesiology, Cell-Gene therapy Translational Medicine Research Center, Third Aliated Hospital of Sun Yat-sen University Shuguang Zhu Department of Hepatic Surgery and Liver Transplantation Center, The Third Aliated Hospital of Sun Yat-sen University Zhongying Hu Guangdong provincial Key Laboratory of Liver Disease Research, Third Aliated Hospital of Sun Yat- sen University Jinliang Liang Guangdong Provincial Key Laboratory of Liver Disease Research, Third Aliated Hospital of Sun Yat- sen University Xuejiao Li Guangdong Provincial Key Laboratory of Liver Disease Research, Third Aliated Hospital of Sun Yat- sen University Xijing Yan Department of Hepatic Surgery and Liver Transplantation Center, Third Aliated Hospital of Sun Yat-sen University Xijian Zhang Department of Hepatic Surgery and Liver Transplantation Center, Third Aliated Hospital of Sun Yat-sen University Danyang Li Department of Infectious Diseases, Third Aliated Hospital of Sun Yat-sen University Wei Liu Guangdong Provincial Key Laboratory of Liver Disease Research, Guangdong province engineering laboratory for transplantation medicine, Third Aliated Hospital -

Genome-Wide Gene Expression Profiling of Randall's Plaques In
CLINICAL RESEARCH www.jasn.org Genome-Wide Gene Expression Profiling of Randall’s Plaques in Calcium Oxalate Stone Formers † † Kazumi Taguchi,* Shuzo Hamamoto,* Atsushi Okada,* Rei Unno,* Hideyuki Kamisawa,* Taku Naiki,* Ryosuke Ando,* Kentaro Mizuno,* Noriyasu Kawai,* Keiichi Tozawa,* Kenjiro Kohri,* and Takahiro Yasui* *Department of Nephro-urology, Nagoya City University Graduate School of Medical Sciences, Nagoya, Japan; and †Department of Urology, Social Medical Corporation Kojunkai Daido Hospital, Daido Clinic, Nagoya, Japan ABSTRACT Randall plaques (RPs) can contribute to the formation of idiopathic calcium oxalate (CaOx) kidney stones; however, genes related to RP formation have not been identified. We previously reported the potential therapeutic role of osteopontin (OPN) and macrophages in CaOx kidney stone formation, discovered using genome-recombined mice and genome-wide analyses. Here, to characterize the genetic patho- genesis of RPs, we used microarrays and immunohistology to compare gene expression among renal papillary RP and non-RP tissues of 23 CaOx stone formers (SFs) (age- and sex-matched) and normal papillary tissue of seven controls. Transmission electron microscopy showed OPN and collagen expression inside and around RPs, respectively. Cluster analysis revealed that the papillary gene expression of CaOx SFs differed significantly from that of controls. Disease and function analysis of gene expression revealed activation of cellular hyperpolarization, reproductive development, and molecular transport in papillary tissue from RPs and non-RP regions of CaOx SFs. Compared with non-RP tissue, RP tissue showed upregulation (˃2-fold) of LCN2, IL11, PTGS1, GPX3,andMMD and downregulation (0.5-fold) of SLC12A1 and NALCN (P,0.01). In network and toxicity analyses, these genes associated with activated mitogen- activated protein kinase, the Akt/phosphatidylinositol 3-kinase pathway, and proinflammatory cytokines that cause renal injury and oxidative stress. -

Table S1. 103 Ferroptosis-Related Genes Retrieved from the Genecards
Table S1. 103 ferroptosis-related genes retrieved from the GeneCards. Gene Symbol Description Category GPX4 Glutathione Peroxidase 4 Protein Coding AIFM2 Apoptosis Inducing Factor Mitochondria Associated 2 Protein Coding TP53 Tumor Protein P53 Protein Coding ACSL4 Acyl-CoA Synthetase Long Chain Family Member 4 Protein Coding SLC7A11 Solute Carrier Family 7 Member 11 Protein Coding VDAC2 Voltage Dependent Anion Channel 2 Protein Coding VDAC3 Voltage Dependent Anion Channel 3 Protein Coding ATG5 Autophagy Related 5 Protein Coding ATG7 Autophagy Related 7 Protein Coding NCOA4 Nuclear Receptor Coactivator 4 Protein Coding HMOX1 Heme Oxygenase 1 Protein Coding SLC3A2 Solute Carrier Family 3 Member 2 Protein Coding ALOX15 Arachidonate 15-Lipoxygenase Protein Coding BECN1 Beclin 1 Protein Coding PRKAA1 Protein Kinase AMP-Activated Catalytic Subunit Alpha 1 Protein Coding SAT1 Spermidine/Spermine N1-Acetyltransferase 1 Protein Coding NF2 Neurofibromin 2 Protein Coding YAP1 Yes1 Associated Transcriptional Regulator Protein Coding FTH1 Ferritin Heavy Chain 1 Protein Coding TF Transferrin Protein Coding TFRC Transferrin Receptor Protein Coding FTL Ferritin Light Chain Protein Coding CYBB Cytochrome B-245 Beta Chain Protein Coding GSS Glutathione Synthetase Protein Coding CP Ceruloplasmin Protein Coding PRNP Prion Protein Protein Coding SLC11A2 Solute Carrier Family 11 Member 2 Protein Coding SLC40A1 Solute Carrier Family 40 Member 1 Protein Coding STEAP3 STEAP3 Metalloreductase Protein Coding ACSL1 Acyl-CoA Synthetase Long Chain Family Member 1 Protein -
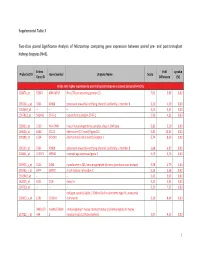
Supplemental Table 3 Two-Class Paired Significance Analysis of Microarrays Comparing Gene Expression Between Paired
Supplemental Table 3 Two‐class paired Significance Analysis of Microarrays comparing gene expression between paired pre‐ and post‐transplant kidneys biopsies (N=8). Entrez Fold q‐value Probe Set ID Gene Symbol Unigene Name Score Gene ID Difference (%) Probe sets higher expressed in post‐transplant biopsies in paired analysis (N=1871) 218870_at 55843 ARHGAP15 Rho GTPase activating protein 15 7,01 3,99 0,00 205304_s_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 6,30 4,50 0,00 1563649_at ‐‐ ‐‐ ‐‐ 6,24 3,51 0,00 1567913_at 541466 CT45‐1 cancer/testis antigen CT45‐1 5,90 4,21 0,00 203932_at 3109 HLA‐DMB major histocompatibility complex, class II, DM beta 5,83 3,20 0,00 204606_at 6366 CCL21 chemokine (C‐C motif) ligand 21 5,82 10,42 0,00 205898_at 1524 CX3CR1 chemokine (C‐X3‐C motif) receptor 1 5,74 8,50 0,00 205303_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 5,68 6,87 0,00 226841_at 219972 MPEG1 macrophage expressed gene 1 5,59 3,76 0,00 203923_s_at 1536 CYBB cytochrome b‐245, beta polypeptide (chronic granulomatous disease) 5,58 4,70 0,00 210135_s_at 6474 SHOX2 short stature homeobox 2 5,53 5,58 0,00 1562642_at ‐‐ ‐‐ ‐‐ 5,42 5,03 0,00 242605_at 1634 DCN decorin 5,23 3,92 0,00 228750_at ‐‐ ‐‐ ‐‐ 5,21 7,22 0,00 collagen, type III, alpha 1 (Ehlers‐Danlos syndrome type IV, autosomal 201852_x_at 1281 COL3A1 dominant) 5,10 8,46 0,00 3493///3 IGHA1///IGHA immunoglobulin heavy constant alpha 1///immunoglobulin heavy 217022_s_at 494 2 constant alpha 2 (A2m marker) 5,07 9,53 0,00 1 202311_s_at