Isolation and Characterization of Multi-Protein Complexes Enriched in the K-Cl Co-Transporter 2 from Brain Plasma Membranes
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

1 Evidence for Gliadin Antibodies As Causative Agents in Schizophrenia
1 Evidence for gliadin antibodies as causative agents in schizophrenia. C.J.Carter PolygenicPathways, 20 Upper Maze Hill, Saint-Leonard’s on Sea, East Sussex, TN37 0LG [email protected] Tel: 0044 (0)1424 422201 I have no fax Abstract Antibodies to gliadin, a component of gluten, have frequently been reported in schizophrenia patients, and in some cases remission has been noted following the instigation of a gluten free diet. Gliadin is a highly immunogenic protein, and B cell epitopes along its entire immunogenic length are homologous to the products of numerous proteins relevant to schizophrenia (p = 0.012 to 3e-25). These include members of the DISC1 interactome, of glutamate, dopamine and neuregulin signalling networks, and of pathways involved in plasticity, dendritic growth or myelination. Antibodies to gliadin are likely to cross react with these key proteins, as has already been observed with synapsin 1 and calreticulin. Gliadin may thus be a causative agent in schizophrenia, under certain genetic and immunological conditions, producing its effects via antibody mediated knockdown of multiple proteins relevant to the disease process. Because of such homology, an autoimmune response may be sustained by the human antigens that resemble gliadin itself, a scenario supported by many reports of immune activation both in the brain and in lymphocytes in schizophrenia. Gluten free diets and removal of such antibodies may be of therapeutic benefit in certain cases of schizophrenia. 2 Introduction A number of studies from China, Norway, and the USA have reported the presence of gliadin antibodies in schizophrenia 1-5. Gliadin is a component of gluten, intolerance to which is implicated in coeliac disease 6. -

Supplementary Data
Supplemental Figure 1: In vitro viability data A C HDQ-P1 basal-like cells HDQ-P1 untreated HDQ-P1 40 0.04 uM MEKi 35 30 25 MCF7 20 15 CAL85-1 %BrdU 10 Incorporation 5 MDA-MB-231 0 MDA-MB-468 0 6E-05 3E-04 0.002 0.008 0.04 0.2 1 5 25 MEKi (ȝM) T47D luminal cells 50 T-47D untreated T-47D 40 0.2uM MEKi 30 20 PD0325901 (ȝM) %BrdU 10 Incorporation 0 0 6E-05 3E-04 0.002 0.008 0.04 0.2 1 5 25 MEKi (ȝM) B D HDQ-P1 Vehicle Treated 1PM MEKi 100 G1: 45.2% 1500 G1: 62.5% MCF7 1000 S: 4.2% S: 3.4% 75 BT474 800 G2/M: 34.9% 1000 G2/M: 27.4% MDA-MB-231 34.9 27.4 50 600 HDQ-P1 # Cells 4.17 #Cells 3.36 Cell viability (%) 400 CAL85-1 45.2 500 62.5 25 0.001 0.01 0.1 1 10 200 MEK inhibitor 2 (PM) 0 0 0 200 400 600 800 1000 0 200 400 600 800 1000 Propidium Iodide Propidium Iodide EGFR B A WEGFR Levels 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 --- MDA-MB-134VI Basal-like HDQ-P1 BT549 Basal-like Her2 DU4475 CAL51 + HCC1806 Supplemental Figure2:PTENandEGFRinBreastLines - MDA-MB-435s CAL85-1 + HCC70 CAL120 -- MX1 CAL148 CAL-120 + CAL85-1 HCC70 -- SW527 HCC1143 CAL-51 HCC1395 + MDA-MB-231 HCC1937 + HCC1143 + BT20 MDA-MB-231 + Hs578T MDA-MB-436 + MDA-MB-468 MDA-MB-468 - HCC1395 MFM223 + BT-549 - CAL-148 AU565 + HCC1937 BT474 + HCC38 EFM192A + MDA-MB-436 + HCC1954 HCC1419 + AU565 ------ HCC1569 BT474 Her2 orLuminal EFM-192A HCC1954 HCC1419 HCC2218 HCC1569 HCC202 MDA-MB-361 HCC2218 MDA-MB-453 + JIM-T --- SKBR3 KPL4 MDA-MB-361 UACC812 MDA-MB-453 UACC893 + SKBR3 ZR75-30 ---------- UACC-812 -

Defining Functional Interactions During Biogenesis of Epithelial Junctions
ARTICLE Received 11 Dec 2015 | Accepted 13 Oct 2016 | Published 6 Dec 2016 | Updated 5 Jan 2017 DOI: 10.1038/ncomms13542 OPEN Defining functional interactions during biogenesis of epithelial junctions J.C. Erasmus1,*, S. Bruche1,*,w, L. Pizarro1,2,*, N. Maimari1,3,*, T. Poggioli1,w, C. Tomlinson4,J.Lees5, I. Zalivina1,w, A. Wheeler1,w, A. Alberts6, A. Russo2 & V.M.M. Braga1 In spite of extensive recent progress, a comprehensive understanding of how actin cytoskeleton remodelling supports stable junctions remains to be established. Here we design a platform that integrates actin functions with optimized phenotypic clustering and identify new cytoskeletal proteins, their functional hierarchy and pathways that modulate E-cadherin adhesion. Depletion of EEF1A, an actin bundling protein, increases E-cadherin levels at junctions without a corresponding reinforcement of cell–cell contacts. This unexpected result reflects a more dynamic and mobile junctional actin in EEF1A-depleted cells. A partner for EEF1A in cadherin contact maintenance is the formin DIAPH2, which interacts with EEF1A. In contrast, depletion of either the endocytic regulator TRIP10 or the Rho GTPase activator VAV2 reduces E-cadherin levels at junctions. TRIP10 binds to and requires VAV2 function for its junctional localization. Overall, we present new conceptual insights on junction stabilization, which integrate known and novel pathways with impact for epithelial morphogenesis, homeostasis and diseases. 1 National Heart and Lung Institute, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. 2 Computing Department, Imperial College London, London SW7 2AZ, UK. 3 Bioengineering Department, Faculty of Engineering, Imperial College London, London SW7 2AZ, UK. 4 Department of Surgery & Cancer, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -
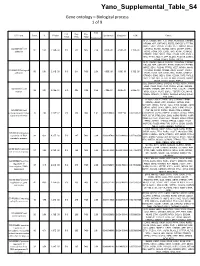
Supplemental Tables4.Pdf
Yano_Supplemental_Table_S4 Gene ontology – Biological process 1 of 9 Fold List Pop Pop GO Term Count % PValue Bonferroni Benjamini FDR Genes Total Hits Total Enrichment DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0007155~cell CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0022610~biological CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DCC, ENAH, PLXNA2, CAPZA2, ATP5B, ASTN1, PAX6, ZEB2, CDH2, CDH4, GLI3, CD24A, EPHB1, NRCAM, GO:0006928~cell CTTNBP2, EDNRB, APP, PTK2, ETV1, CLASP2, STRBP, 36 4.04 3.46E-07 510 205 7436 2.56 7.28E-04 3.64E-04 5.98E-04 motion NRG1, DCLK1, PLAT, SGPL1, TGFBR1, EVL, MYH9, YWHAE, NCKAP1, CTNNA2, SEMA6A, EPHA4, NDEL1, FYN, LRP6 PLXNA2, ADCY5, PAX6, GLI3, CTNNB1, LPHN2, EDNRB, LPHN3, APP, CSNK2A1, GPR45, NRG1, RAPGEF1, WWOX, SGPL1, TLE4, SPEN, NCAM1, DDR1, GRB10, GRM3, GNAQ, HIPK1, GNB1, HIPK2, PYGO1, GO:0007166~cell RNF138, ROR2, CNTN1, -

Serum Albumin OS=Homo Sapiens
Protein Name Cluster of Glial fibrillary acidic protein OS=Homo sapiens GN=GFAP PE=1 SV=1 (P14136) Serum albumin OS=Homo sapiens GN=ALB PE=1 SV=2 Cluster of Isoform 3 of Plectin OS=Homo sapiens GN=PLEC (Q15149-3) Cluster of Hemoglobin subunit beta OS=Homo sapiens GN=HBB PE=1 SV=2 (P68871) Vimentin OS=Homo sapiens GN=VIM PE=1 SV=4 Cluster of Tubulin beta-3 chain OS=Homo sapiens GN=TUBB3 PE=1 SV=2 (Q13509) Cluster of Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 (P60709) Cluster of Tubulin alpha-1B chain OS=Homo sapiens GN=TUBA1B PE=1 SV=1 (P68363) Cluster of Isoform 2 of Spectrin alpha chain, non-erythrocytic 1 OS=Homo sapiens GN=SPTAN1 (Q13813-2) Hemoglobin subunit alpha OS=Homo sapiens GN=HBA1 PE=1 SV=2 Cluster of Spectrin beta chain, non-erythrocytic 1 OS=Homo sapiens GN=SPTBN1 PE=1 SV=2 (Q01082) Cluster of Pyruvate kinase isozymes M1/M2 OS=Homo sapiens GN=PKM PE=1 SV=4 (P14618) Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 Clathrin heavy chain 1 OS=Homo sapiens GN=CLTC PE=1 SV=5 Filamin-A OS=Homo sapiens GN=FLNA PE=1 SV=4 Cytoplasmic dynein 1 heavy chain 1 OS=Homo sapiens GN=DYNC1H1 PE=1 SV=5 Cluster of ATPase, Na+/K+ transporting, alpha 2 (+) polypeptide OS=Homo sapiens GN=ATP1A2 PE=3 SV=1 (B1AKY9) Fibrinogen beta chain OS=Homo sapiens GN=FGB PE=1 SV=2 Fibrinogen alpha chain OS=Homo sapiens GN=FGA PE=1 SV=2 Dihydropyrimidinase-related protein 2 OS=Homo sapiens GN=DPYSL2 PE=1 SV=1 Cluster of Alpha-actinin-1 OS=Homo sapiens GN=ACTN1 PE=1 SV=2 (P12814) 60 kDa heat shock protein, mitochondrial OS=Homo -

Spectrin Binding Motifs Control Scribble Cortical Dynamics And
1 2 3 Spectrin binding motifs control Scribble cortical 4 dynamics and polarity function 5 6 Batiste Boëda and Sandrine Etienne Manneville 7 8 Institut Pasteur (CNRS URA 3691-INSERM), France 9 Correspondance to Sandrine Etienne-Manneville. Cell Polarity, Migration and Cancer Unit, 10 Institut Pasteur, 25 rue du Dr Roux, 75724 Paris cedex 15, France; Phone: +33 1 4438 9591; 11 FAX: +33 1 4568 8548. 12 e-mail: [email protected] 13 14 Competing interests statement: 15 The authors declare that no competing interests exist. 16 17 1 18 Abstract 19 The tumor suppressor protein Scribble (SCRIB) plays an evolutionary conserved role in 20 cell polarity. Despite being central for its function, the molecular basis of SCRIB 21 recruitment and stabilization at the cell cortex is poorly understood. Here we show that 22 SCRIB binds directly to the CH1 domain of spectrins, a molecular scaffold that 23 contributes to the cortical actin cytoskeleton and connects it to the plasma membrane. 24 We have identified a short evolutionary conserved peptide motif named SADH motif 25 (SCRIB ABLIMs DMTN Homology) which is necessary and sufficient to mediate protein 26 interaction with spectrins. The SADH domains contribute to SCRIB dynamics at the 27 cell cortex and SCRIB polarity function. Furthermore, mutations in SCRIB SADH 28 domains associated with spina bifida and cancer impact the stability of SCRIB at the 29 plasma membrane, suggesting that SADH domain alterations may participate in human 30 pathology. 31 32 33 34 35 36 37 2 38 Introduction 39 The protein SCRIB has been implicated in a staggering array of cellular processes 40 including polarity, migration, proliferation, differentiation, apoptosis, stemcell 41 maintenance, and vesicle trafficking [1]. -

Myopathy Genes (HGNC) Neuropathy (HGNC) Neuromuscular Disease
Myopathy Genes Neuropathy Neuromuscular Disease (HGNC) (HGNC) (HGNC) ABHD5 ABCA1 ADCK3 ACTG2 ACO2 AGRN AGK AGXT ALS2 ALDOA AIFM1 ANG AMER1 ALAD AP4B1 ANO5 AMACR AP4E1 AR AP1S1 AP4M1 AUH APTX AP4S1 B4GALT1 AR AP5Z1 CACNA1S ATL3 ATM CASQ1 B4GALNT1 ATXN10 CCDC78 BAG3 ATXN7 CHCHD10 BRP44L BEAN1 CHRNA1 C12orf65 C9orf72 CHRNB1 C19orf12 CACNB4 CHRND C1NH CAPN3 CHRNE CECR1 CHAT CLPB CISD2 CHKB COL6A1 CLCF1 CHMP2B COL6A2 CLCN2 CHRNG COL6A3 CLP1 CLCN1 COLQ CMT2G COL9A3 CTNS CMT2H COQ2 DGUOK CMTDIA COQ6 DNA2 CMTX2 COQ9 DNAJB6 CMTX3 COX15 DNAJC19 COASY CPT1A DNM2 COX6A1 CYP7B1 DPM2 CPOX DAG1 DYSF CYP27A1 DDHD2 EMD CYP2U1 DOK7 EPG5 DARS2 DPAGT1 FAM111B DCAF8 DPM3 FBXL4 DDHD1 DUX4 FKBP14 DFNX5 ECEL1 FKRP DHTKD1 ERBB3 FLH1 DIAPH3 ERLIN2 FLNC DNAJB2 FA2H HNRNPA1 DNAJC3 FKTN HNRNPDL ELOVL5 FUS HNRPA2B1 ERCC8 G6PC KLHL40 FAH GFPT1 KLHL41 FAM126A GLE1 LAMA2 FBN1 GYS2 LDB3 FMR1 HSPD1 LMOD3 FXN IFRD1 MEGF10 GALC INF2 MGME1 GBE1 ISPD MTAP GJC2 ITGA7 MTMR14 GP1BA ITPR1 MYF6 HADHA KCNA1 MYH14 HADHB KCNC3 MYLK2 HFE KCNE3 NARS2 HINT1 KCNJ18 NEB HK1 KCNJ2 ORAI1 HMBS KIAA0196 PRKAG2 HSD17B4 KIF21A PTEN HSN1B L1CAM RBCK1 IARS2 LAMB2 RET IGHMBP2 LARGE RMND1 KCNJ10 MCCC2 SCN4A KIF5A MRE11A SERAC1 LRSAM1 MRPL3 SGCA LYST MTO1 SIL1 MANBA MTPAP SPEG MARS MTTP STAC3 MTATP6 MUSK STIM1 MYH14 MYBPC3 SYNE1 MYOT MYH3 SYNE2 NAMSD MYH8 TAZ NF2 NF1 TIA1 NGLY1 NIPA1 TMEM43 NMSR NOP56 TNPO3 NOTCH3 OPTN TNXB OPA1 PDSS2 TPM2 OPA3 PDYN TRPV4 OTOF PFN1 UBA1 PDK3 PHKA2 VCP PDSS1 PHKG2 XDH PEX10 PHOX2A ACADS PEX2 PIP5K1C ACADVL PMM2 PLEC ACTA1 PNPLA6 PLP1 AGL PPOX POMGNT1 AMPD1 PRICKLE1 -

Supplementary Materials
Supplementary materials Supplementary Table S1: MGNC compound library Ingredien Molecule Caco- Mol ID MW AlogP OB (%) BBB DL FASA- HL t Name Name 2 shengdi MOL012254 campesterol 400.8 7.63 37.58 1.34 0.98 0.7 0.21 20.2 shengdi MOL000519 coniferin 314.4 3.16 31.11 0.42 -0.2 0.3 0.27 74.6 beta- shengdi MOL000359 414.8 8.08 36.91 1.32 0.99 0.8 0.23 20.2 sitosterol pachymic shengdi MOL000289 528.9 6.54 33.63 0.1 -0.6 0.8 0 9.27 acid Poricoic acid shengdi MOL000291 484.7 5.64 30.52 -0.08 -0.9 0.8 0 8.67 B Chrysanthem shengdi MOL004492 585 8.24 38.72 0.51 -1 0.6 0.3 17.5 axanthin 20- shengdi MOL011455 Hexadecano 418.6 1.91 32.7 -0.24 -0.4 0.7 0.29 104 ylingenol huanglian MOL001454 berberine 336.4 3.45 36.86 1.24 0.57 0.8 0.19 6.57 huanglian MOL013352 Obacunone 454.6 2.68 43.29 0.01 -0.4 0.8 0.31 -13 huanglian MOL002894 berberrubine 322.4 3.2 35.74 1.07 0.17 0.7 0.24 6.46 huanglian MOL002897 epiberberine 336.4 3.45 43.09 1.17 0.4 0.8 0.19 6.1 huanglian MOL002903 (R)-Canadine 339.4 3.4 55.37 1.04 0.57 0.8 0.2 6.41 huanglian MOL002904 Berlambine 351.4 2.49 36.68 0.97 0.17 0.8 0.28 7.33 Corchorosid huanglian MOL002907 404.6 1.34 105 -0.91 -1.3 0.8 0.29 6.68 e A_qt Magnogrand huanglian MOL000622 266.4 1.18 63.71 0.02 -0.2 0.2 0.3 3.17 iolide huanglian MOL000762 Palmidin A 510.5 4.52 35.36 -0.38 -1.5 0.7 0.39 33.2 huanglian MOL000785 palmatine 352.4 3.65 64.6 1.33 0.37 0.7 0.13 2.25 huanglian MOL000098 quercetin 302.3 1.5 46.43 0.05 -0.8 0.3 0.38 14.4 huanglian MOL001458 coptisine 320.3 3.25 30.67 1.21 0.32 0.9 0.26 9.33 huanglian MOL002668 Worenine -

Cldn19 Clic2 Clmp Cln3
NewbornDx™ Advanced Sequencing Evaluation When time to diagnosis matters, the NewbornDx™ Advanced Sequencing Evaluation from Athena Diagnostics delivers rapid, 5- to 7-day results on a targeted 1,722-genes. A2ML1 ALAD ATM CAV1 CLDN19 CTNS DOCK7 ETFB FOXC2 GLUL HOXC13 JAK3 AAAS ALAS2 ATP1A2 CBL CLIC2 CTRC DOCK8 ETFDH FOXE1 GLYCTK HOXD13 JUP AARS2 ALDH18A1 ATP1A3 CBS CLMP CTSA DOK7 ETHE1 FOXE3 GM2A HPD KANK1 AASS ALDH1A2 ATP2B3 CC2D2A CLN3 CTSD DOLK EVC FOXF1 GMPPA HPGD K ANSL1 ABAT ALDH3A2 ATP5A1 CCDC103 CLN5 CTSK DPAGT1 EVC2 FOXG1 GMPPB HPRT1 KAT6B ABCA12 ALDH4A1 ATP5E CCDC114 CLN6 CUBN DPM1 EXOC4 FOXH1 GNA11 HPSE2 KCNA2 ABCA3 ALDH5A1 ATP6AP2 CCDC151 CLN8 CUL4B DPM2 EXOSC3 FOXI1 GNAI3 HRAS KCNB1 ABCA4 ALDH7A1 ATP6V0A2 CCDC22 CLP1 CUL7 DPM3 EXPH5 FOXL2 GNAO1 HSD17B10 KCND2 ABCB11 ALDOA ATP6V1B1 CCDC39 CLPB CXCR4 DPP6 EYA1 FOXP1 GNAS HSD17B4 KCNE1 ABCB4 ALDOB ATP7A CCDC40 CLPP CYB5R3 DPYD EZH2 FOXP2 GNE HSD3B2 KCNE2 ABCB6 ALG1 ATP8A2 CCDC65 CNNM2 CYC1 DPYS F10 FOXP3 GNMT HSD3B7 KCNH2 ABCB7 ALG11 ATP8B1 CCDC78 CNTN1 CYP11B1 DRC1 F11 FOXRED1 GNPAT HSPD1 KCNH5 ABCC2 ALG12 ATPAF2 CCDC8 CNTNAP1 CYP11B2 DSC2 F13A1 FRAS1 GNPTAB HSPG2 KCNJ10 ABCC8 ALG13 ATR CCDC88C CNTNAP2 CYP17A1 DSG1 F13B FREM1 GNPTG HUWE1 KCNJ11 ABCC9 ALG14 ATRX CCND2 COA5 CYP1B1 DSP F2 FREM2 GNS HYDIN KCNJ13 ABCD3 ALG2 AUH CCNO COG1 CYP24A1 DST F5 FRMD7 GORAB HYLS1 KCNJ2 ABCD4 ALG3 B3GALNT2 CCS COG4 CYP26C1 DSTYK F7 FTCD GP1BA IBA57 KCNJ5 ABHD5 ALG6 B3GAT3 CCT5 COG5 CYP27A1 DTNA F8 FTO GP1BB ICK KCNJ8 ACAD8 ALG8 B3GLCT CD151 COG6 CYP27B1 DUOX2 F9 FUCA1 GP6 ICOS KCNK3 ACAD9 ALG9 -

MYO5A Rabbit Pab
Leader in Biomolecular Solutions for Life Science MYO5A Rabbit pAb Catalog No.: A9830 Basic Information Background Catalog No. This gene is one of three myosin V heavy-chain genes, belonging to the myosin gene A9830 superfamily. Myosin V is a class of actin-based motor proteins involved in cytoplasmic vesicle transport and anchorage, spindle-pole alignment and mRNA translocation. The Observed MW protein encoded by this gene is abundant in melanocytes and nerve cells. Mutations in 240kDa this gene cause Griscelli syndrome type-1 (GS1), Griscelli syndrome type-3 (GS3) and neuroectodermal melanolysosomal disease, or Elejalde disease. Multiple alternatively Calculated MW spliced transcript variants encoding different isoforms have been reported, but the full- 212kDa/215kDa/218kDa length nature of some variants has not been determined. Category Primary antibody Applications WB,IHC,IF Cross-Reactivity Human, Mouse, Rat Recommended Dilutions Immunogen Information WB 1:500 - 1:2000 Gene ID Swiss Prot 4644 Q9Y4I1 IHC 1:50 - 1:100 Immunogen 1:50 - 1:100 IF Recombinant fusion protein containing a sequence corresponding to amino acids 950-1150 of human MYO5A (NP_000250.3). Synonyms MYO5A;GS1;MYH12;MYO5;MYR12;myosin VA Contact Product Information www.abclonal.com Source Isotype Purification Rabbit IgG Affinity purification Storage Store at -20℃. Avoid freeze / thaw cycles. Buffer: PBS with 0.02% sodium azide,50% glycerol,pH7.3. Validation Data Western blot analysis of extracts of various cell lines, using MYO5A antibody (A9830) at 1:1000 dilution. Secondary antibody: HRP Goat Anti-Rabbit IgG (H+L) (AS014) at 1:10000 dilution. Lysates/proteins: 25ug per lane. Blocking buffer: 3% nonfat dry milk in TBST. -
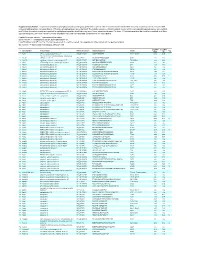
Supplementary Table 1
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding