Strong Correlation Between Air-Liquid Interface Cultures and in Vivo Transcriptomics of Nasal Brush Biopsy
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Identification and Developmental Expression of the Full Complement Of
Goldstone et al. BMC Genomics 2010, 11:643 http://www.biomedcentral.com/1471-2164/11/643 RESEARCH ARTICLE Open Access Identification and developmental expression of the full complement of Cytochrome P450 genes in Zebrafish Jared V Goldstone1, Andrew G McArthur2, Akira Kubota1, Juliano Zanette1,3, Thiago Parente1,4, Maria E Jönsson1,5, David R Nelson6, John J Stegeman1* Abstract Background: Increasing use of zebrafish in drug discovery and mechanistic toxicology demands knowledge of cytochrome P450 (CYP) gene regulation and function. CYP enzymes catalyze oxidative transformation leading to activation or inactivation of many endogenous and exogenous chemicals, with consequences for normal physiology and disease processes. Many CYPs potentially have roles in developmental specification, and many chemicals that cause developmental abnormalities are substrates for CYPs. Here we identify and annotate the full suite of CYP genes in zebrafish, compare these to the human CYP gene complement, and determine the expression of CYP genes during normal development. Results: Zebrafish have a total of 94 CYP genes, distributed among 18 gene families found also in mammals. There are 32 genes in CYP families 5 to 51, most of which are direct orthologs of human CYPs that are involved in endogenous functions including synthesis or inactivation of regulatory molecules. The high degree of sequence similarity suggests conservation of enzyme activities for these CYPs, confirmed in reports for some steroidogenic enzymes (e.g. CYP19, aromatase; CYP11A, P450scc; CYP17, steroid 17a-hydroxylase), and the CYP26 retinoic acid hydroxylases. Complexity is much greater in gene families 1, 2, and 3, which include CYPs prominent in metabolism of drugs and pollutants, as well as of endogenous substrates. -

Defining Functional Interactions During Biogenesis of Epithelial Junctions
ARTICLE Received 11 Dec 2015 | Accepted 13 Oct 2016 | Published 6 Dec 2016 | Updated 5 Jan 2017 DOI: 10.1038/ncomms13542 OPEN Defining functional interactions during biogenesis of epithelial junctions J.C. Erasmus1,*, S. Bruche1,*,w, L. Pizarro1,2,*, N. Maimari1,3,*, T. Poggioli1,w, C. Tomlinson4,J.Lees5, I. Zalivina1,w, A. Wheeler1,w, A. Alberts6, A. Russo2 & V.M.M. Braga1 In spite of extensive recent progress, a comprehensive understanding of how actin cytoskeleton remodelling supports stable junctions remains to be established. Here we design a platform that integrates actin functions with optimized phenotypic clustering and identify new cytoskeletal proteins, their functional hierarchy and pathways that modulate E-cadherin adhesion. Depletion of EEF1A, an actin bundling protein, increases E-cadherin levels at junctions without a corresponding reinforcement of cell–cell contacts. This unexpected result reflects a more dynamic and mobile junctional actin in EEF1A-depleted cells. A partner for EEF1A in cadherin contact maintenance is the formin DIAPH2, which interacts with EEF1A. In contrast, depletion of either the endocytic regulator TRIP10 or the Rho GTPase activator VAV2 reduces E-cadherin levels at junctions. TRIP10 binds to and requires VAV2 function for its junctional localization. Overall, we present new conceptual insights on junction stabilization, which integrate known and novel pathways with impact for epithelial morphogenesis, homeostasis and diseases. 1 National Heart and Lung Institute, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. 2 Computing Department, Imperial College London, London SW7 2AZ, UK. 3 Bioengineering Department, Faculty of Engineering, Imperial College London, London SW7 2AZ, UK. 4 Department of Surgery & Cancer, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. -

Analysis of the Human Serum Proteome
Cedarville University DigitalCommons@Cedarville Pharmaceutical Sciences Faculty Publications Department of Pharmaceutical Sciences 6-2004 Analysis of the Human Serum Proteome King C. Chan David A. Lucas Denise Hise Carl F. Schaefer Zhen Xiao See next page for additional authors Follow this and additional works at: https://digitalcommons.cedarville.edu/ pharmaceutical_sciences_publications Part of the Pharmacy and Pharmaceutical Sciences Commons This Article is brought to you for free and open access by DigitalCommons@Cedarville, a service of the Centennial Library. It has been accepted for inclusion in Pharmaceutical Sciences Faculty Publications by an authorized administrator of DigitalCommons@Cedarville. For more information, please contact [email protected]. Authors King C. Chan, David A. Lucas, Denise Hise, Carl F. Schaefer, Zhen Xiao, George M. Janini, Kenneth H. Buetow, Haleem J. Issaq, Timothy D. Veenstra, and Thomas P. Conrads Clinical Proteomics Journal Copyright ©Humana Press Inc. All rights of any nature whatsoever are reserved. ISSN 1542-6416/04/01:101–225/$25.00 Serum/Plasma Proteome Analysis of the Human Serum Proteome King C. Chan,1,† David A. Lucas,1,† Denise Hise,2 Carl F. Schaefer,2 Zhen Xiao,1 George M. Janini,1 Kenneth H. Buetow,2 Haleem J. Issaq,1 Timothy D.Veenstra,1 and Thomas P. Conrads1,* 1Laboratory of Proteomics and Analytical Technologies, National Cancer Institute at Frederick, SAIC-Frederick, Inc, PO Box B, Frederick, MD 21702 2Center for Bioinformatics, National Cancer Institute, Bethesda, MD 20892 †These authors contributed equally to this work. each of which was analyzed by microcapillary Abstract reversed-phase liquid chromatography coupled Changes in serum proteins that signal online with MS/MS analysis. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Synonymous Single Nucleotide Polymorphisms in Human Cytochrome
DMD Fast Forward. Published on February 9, 2009 as doi:10.1124/dmd.108.026047 DMD #26047 TITLE PAGE: A BIOINFORMATICS APPROACH FOR THE PHENOTYPE PREDICTION OF NON- SYNONYMOUS SINGLE NUCLEOTIDE POLYMORPHISMS IN HUMAN CYTOCHROME P450S LIN-LIN WANG, YONG LI, SHU-FENG ZHOU Department of Nutrition and Food Hygiene, School of Public Health, Peking University, Beijing 100191, P. R. China (LL Wang & Y Li) Discipline of Chinese Medicine, School of Health Sciences, RMIT University, Bundoora, Victoria 3083, Australia (LL Wang & SF Zhou). 1 Copyright 2009 by the American Society for Pharmacology and Experimental Therapeutics. DMD #26047 RUNNING TITLE PAGE: a) Running title: Prediction of phenotype of human CYPs. b) Author for correspondence: A/Prof. Shu-Feng Zhou, MD, PhD Discipline of Chinese Medicine, School of Health Sciences, RMIT University, WHO Collaborating Center for Traditional Medicine, Bundoora, Victoria 3083, Australia. Tel: + 61 3 9925 7794; fax: +61 3 9925 7178. Email: [email protected] c) Number of text pages: 21 Number of tables: 10 Number of figures: 2 Number of references: 40 Number of words in Abstract: 249 Number of words in Introduction: 749 Number of words in Discussion: 1459 d) Non-standard abbreviations: CYP, cytochrome P450; nsSNP, non-synonymous single nucleotide polymorphism. 2 DMD #26047 ABSTRACT Non-synonymous single nucleotide polymorphisms (nsSNPs) in coding regions that can lead to amino acid changes may cause alteration of protein function and account for susceptivity to disease. Identification of deleterious nsSNPs from tolerant nsSNPs is important for characterizing the genetic basis of human disease, assessing individual susceptibility to disease, understanding the pathogenesis of disease, identifying molecular targets for drug treatment and conducting individualized pharmacotherapy. -
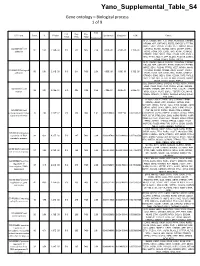
Supplemental Tables4.Pdf
Yano_Supplemental_Table_S4 Gene ontology – Biological process 1 of 9 Fold List Pop Pop GO Term Count % PValue Bonferroni Benjamini FDR Genes Total Hits Total Enrichment DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0007155~cell CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0022610~biological CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DCC, ENAH, PLXNA2, CAPZA2, ATP5B, ASTN1, PAX6, ZEB2, CDH2, CDH4, GLI3, CD24A, EPHB1, NRCAM, GO:0006928~cell CTTNBP2, EDNRB, APP, PTK2, ETV1, CLASP2, STRBP, 36 4.04 3.46E-07 510 205 7436 2.56 7.28E-04 3.64E-04 5.98E-04 motion NRG1, DCLK1, PLAT, SGPL1, TGFBR1, EVL, MYH9, YWHAE, NCKAP1, CTNNA2, SEMA6A, EPHA4, NDEL1, FYN, LRP6 PLXNA2, ADCY5, PAX6, GLI3, CTNNB1, LPHN2, EDNRB, LPHN3, APP, CSNK2A1, GPR45, NRG1, RAPGEF1, WWOX, SGPL1, TLE4, SPEN, NCAM1, DDR1, GRB10, GRM3, GNAQ, HIPK1, GNB1, HIPK2, PYGO1, GO:0007166~cell RNF138, ROR2, CNTN1, -

Serum Albumin OS=Homo Sapiens
Protein Name Cluster of Glial fibrillary acidic protein OS=Homo sapiens GN=GFAP PE=1 SV=1 (P14136) Serum albumin OS=Homo sapiens GN=ALB PE=1 SV=2 Cluster of Isoform 3 of Plectin OS=Homo sapiens GN=PLEC (Q15149-3) Cluster of Hemoglobin subunit beta OS=Homo sapiens GN=HBB PE=1 SV=2 (P68871) Vimentin OS=Homo sapiens GN=VIM PE=1 SV=4 Cluster of Tubulin beta-3 chain OS=Homo sapiens GN=TUBB3 PE=1 SV=2 (Q13509) Cluster of Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 (P60709) Cluster of Tubulin alpha-1B chain OS=Homo sapiens GN=TUBA1B PE=1 SV=1 (P68363) Cluster of Isoform 2 of Spectrin alpha chain, non-erythrocytic 1 OS=Homo sapiens GN=SPTAN1 (Q13813-2) Hemoglobin subunit alpha OS=Homo sapiens GN=HBA1 PE=1 SV=2 Cluster of Spectrin beta chain, non-erythrocytic 1 OS=Homo sapiens GN=SPTBN1 PE=1 SV=2 (Q01082) Cluster of Pyruvate kinase isozymes M1/M2 OS=Homo sapiens GN=PKM PE=1 SV=4 (P14618) Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 Clathrin heavy chain 1 OS=Homo sapiens GN=CLTC PE=1 SV=5 Filamin-A OS=Homo sapiens GN=FLNA PE=1 SV=4 Cytoplasmic dynein 1 heavy chain 1 OS=Homo sapiens GN=DYNC1H1 PE=1 SV=5 Cluster of ATPase, Na+/K+ transporting, alpha 2 (+) polypeptide OS=Homo sapiens GN=ATP1A2 PE=3 SV=1 (B1AKY9) Fibrinogen beta chain OS=Homo sapiens GN=FGB PE=1 SV=2 Fibrinogen alpha chain OS=Homo sapiens GN=FGA PE=1 SV=2 Dihydropyrimidinase-related protein 2 OS=Homo sapiens GN=DPYSL2 PE=1 SV=1 Cluster of Alpha-actinin-1 OS=Homo sapiens GN=ACTN1 PE=1 SV=2 (P12814) 60 kDa heat shock protein, mitochondrial OS=Homo -

Identification of Potential Key Genes and Pathway Linked with Sporadic Creutzfeldt-Jakob Disease Based on Integrated Bioinformatics Analyses
medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Identification of potential key genes and pathway linked with sporadic Creutzfeldt-Jakob disease based on integrated bioinformatics analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 , Iranna Kotturshetti 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. 3. Department of Ayurveda, Rajiv Gandhi Education Society`s Ayurvedic Medical College, Ron, Karnataka 562209, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India NOTE: This preprint reports new research that has not been certified by peer review and should not be used to guide clinical practice. medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Abstract Sporadic Creutzfeldt-Jakob disease (sCJD) is neurodegenerative disease also called prion disease linked with poor prognosis. The aim of the current study was to illuminate the underlying molecular mechanisms of sCJD. The mRNA microarray dataset GSE124571 was downloaded from the Gene Expression Omnibus database. Differentially expressed genes (DEGs) were screened. -

MYO5A Rabbit Pab
Leader in Biomolecular Solutions for Life Science MYO5A Rabbit pAb Catalog No.: A9830 Basic Information Background Catalog No. This gene is one of three myosin V heavy-chain genes, belonging to the myosin gene A9830 superfamily. Myosin V is a class of actin-based motor proteins involved in cytoplasmic vesicle transport and anchorage, spindle-pole alignment and mRNA translocation. The Observed MW protein encoded by this gene is abundant in melanocytes and nerve cells. Mutations in 240kDa this gene cause Griscelli syndrome type-1 (GS1), Griscelli syndrome type-3 (GS3) and neuroectodermal melanolysosomal disease, or Elejalde disease. Multiple alternatively Calculated MW spliced transcript variants encoding different isoforms have been reported, but the full- 212kDa/215kDa/218kDa length nature of some variants has not been determined. Category Primary antibody Applications WB,IHC,IF Cross-Reactivity Human, Mouse, Rat Recommended Dilutions Immunogen Information WB 1:500 - 1:2000 Gene ID Swiss Prot 4644 Q9Y4I1 IHC 1:50 - 1:100 Immunogen 1:50 - 1:100 IF Recombinant fusion protein containing a sequence corresponding to amino acids 950-1150 of human MYO5A (NP_000250.3). Synonyms MYO5A;GS1;MYH12;MYO5;MYR12;myosin VA Contact Product Information www.abclonal.com Source Isotype Purification Rabbit IgG Affinity purification Storage Store at -20℃. Avoid freeze / thaw cycles. Buffer: PBS with 0.02% sodium azide,50% glycerol,pH7.3. Validation Data Western blot analysis of extracts of various cell lines, using MYO5A antibody (A9830) at 1:1000 dilution. Secondary antibody: HRP Goat Anti-Rabbit IgG (H+L) (AS014) at 1:10000 dilution. Lysates/proteins: 25ug per lane. Blocking buffer: 3% nonfat dry milk in TBST. -

The Kinesin-2 Family Member KIF3C Regulates Microtubule Dynamics and Is Required for Axon Growth and Regeneration
The Journal of Neuroscience, July 10, 2013 • 33(28):11329–11345 • 11329 Development/Plasticity/Repair The Kinesin-2 Family Member KIF3C Regulates Microtubule Dynamics and Is Required for Axon Growth and Regeneration Laura F. Gumy,1,2 Daniel J. Chew,1 Elena Tortosa,2 Eugene A. Katrukha,2 Lukas C. Kapitein,2 Aviva M. Tolkovsky,1 Casper C. Hoogenraad,2 and James W. Fawcett1 1Centre for Brain Repair, University of Cambridge, Cambridge CB2 0PY, United Kingdom, and 2Division of Cell Biology, Faculty of Science, University of Utrecht, 3584CH Utrecht, The Netherlands Axonregenerationafterinjuryrequirestheextensivereconstruction,reorganization,andstabilizationofthemicrotubulecytoskeletonin the growth cones. Here, we identify KIF3C as a key regulator of axonal growth and regeneration by controlling microtubule dynamics and organization in the growth cone. KIF3C is developmentally regulated. Rat embryonic sensory axons and growth cones contain undetect- able levels of KIF3C protein that is locally translated immediately after injury. In adult neurons, KIF3C is axonally transported from the cell body and is enriched at the growth cone where it preferentially binds to tyrosinated microtubules. Functionally, the interaction of KIF3CwithEB3isnecessaryforitslocalizationatthemicrotubuleplus-endsinthegrowthcone.DepletionofKIF3Cinadultneuronsleads to an increase in stable, overgrown and looped microtubules because of a strong decrease in the microtubule frequency of catastrophes, suggesting that KIF3C functions as a microtubule-destabilizing factor. Adult axons lacking KIF3C, by RNA interference or KIF3C gene knock-out, display an impaired axonal outgrowth in vitro and a delayed regeneration after injury both in vitro and in vivo. Murine KIF3C knock-out embryonic axons grow normally but do not regenerate after injury because they are unable to locally translate KIF3C. These data show that KIF3C is an injury-specific kinesin that contributes to axon growth and regeneration by regulating and organizing the microtubule cytoskeleton in the growth cone. -

CYP2F1 (NM 000774) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC223097 CYP2F1 (NM_000774) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: CYP2F1 (NM_000774) Human Tagged ORF Clone Tag: Myc-DDK Symbol: CYP2F1 Synonyms: C2F1; CYP2F; CYPIIF1 Vector: pCMV6-Entry (PS100001) E. coli Selection: Kanamycin (25 ug/mL) Cell Selection: Neomycin This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 4 CYP2F1 (NM_000774) Human Tagged ORF Clone – RC223097 ORF Nucleotide >RC223097 representing NM_000774 Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGGACAGCATAAGCACAGCCATCTTACTCCTGCTCCTGGCTCTCGTCTGTCTGCTCCTGACCCTAAGCT CAAGAGATAAGGGAAAGCTGCCTCCGGGACCCAGACCCCTCTCAATCCTGGGAAACCTGCTGCTGCTTTG CTCCCAAGACATGCTGACTTCTCTCACTAAGCTGAGCAAGGAGTATGGCTCCATGTACACAGTGCACCTG GGACCCAGGCGGGTGGTGGTCCTCAGCGGGTACCAAGCTGTGAAGGAGGCCCTGGTGGACCAGGGAGAGG AGTTTAGTGGCCGCGGTGACTACCCTGCCTTTTTCAACTTTACCAAGGGCAATGGCATCGCCTTCTCCAG TGGGGATCGATGGAAGGTCCTGAGACAGTTCTCTATCCAGATTCTACGGAATTTCGGGATGGGGAAGAGA AGCATTGAGGAGCGAATCCTAGAGGAGGGCAGCTTCCTGCTGGCGGAGCTGCGGAAAACTGAAGGCGAGC CCTTTGACCCCACGTTTGTGCTGAGTCGCTCAGTGTCCAACATTATCTGTTCCGTGCTCTTCGGCAGCCG CTTCGACTATGATGATGAGCGTCTGCTCACCATTATCCGCCTTATCAATGACAACTTCCAAATCATGAGC -

Comprehensive Analysis of Expression, Prognosis and Immune Infltrates for Kinesin Superfamily Members in Human Pancreatic Adenocarcinoma
Comprehensive Analysis of Expression, Prognosis and Immune Inltrates for Kinesin Superfamily Members in Human Pancreatic Adenocarcinoma Zehao Chen ( [email protected] ) Wuhan University https://orcid.org/0000-0001-9440-0782 Jie Wang Wuhan University Jian Xu Wuhan University Wenjie Zhu Wuhan University Jianxin Jiang Wuhan University Research Keywords: Kinesin superfamily, Pancreatic Adenocarcinoma, Expression, Prognostic value, Immune Inltrates Posted Date: June 16th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-550619/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/26 Abstract Background Kinesin superfamily (KIFs) has a long-reported signicant inuence on the initiation, development, and progress of pancreatic adenocarcinoma (PAAD). However, the expression level of different KIFs in PAAD and its relationship with the prognosis and immune inltration in patients with PAAD have not been fully elucidated. Methods Comprehensive bioinformatics analyses were done using data from UCSC XENA data hubs, TCGA databaseOncomine databases, GEO datasets, GEPIA, GTEx, The human protein atlas, Kaplan Meier plotter, cBioPortal, STRING and KEGG database. Then, the relationship between KIFs expression and tumor immune inltrates was studied by using the TIMER database. Results A total of 24 differentially expressed KIFs at transcriptional levels were identied between tumor tissues and normal tissues with 1 (KIF1A) downregulated and 23 (KIF2A, KIF2C, KIF3A, KIF3B, KIF3C, KIF4A, KIF5B, KIF7, KIF9, KIF10, KIF11, KIF13A, KIF13B, KIF15, KIF16B, KIF18B, KIF20A, KIF20B, KIF21B, KIF22, KIF23, KIF26B, KIFC1) overexpressed by GEPIA. The protein expression level of KIF3A, KIF3B, KIF4A, KIF5B, KIF7, KIF9, CENPE, KIF11, KIF13A, KIF13B, KIF15, KIF16B, KIF18B, KIF20A, KIF20B, KIF21B, KIF22, KIF23, KIF26B, KIFC1 are higher in PAAD tissues than in the adjacent tissues, which is almost the same as the results of KIFs at transcriptional levels in PAAD.