Spectrin Binding Motifs Control Scribble Cortical Dynamics And
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Data
Supplemental Figure 1: In vitro viability data A C HDQ-P1 basal-like cells HDQ-P1 untreated HDQ-P1 40 0.04 uM MEKi 35 30 25 MCF7 20 15 CAL85-1 %BrdU 10 Incorporation 5 MDA-MB-231 0 MDA-MB-468 0 6E-05 3E-04 0.002 0.008 0.04 0.2 1 5 25 MEKi (ȝM) T47D luminal cells 50 T-47D untreated T-47D 40 0.2uM MEKi 30 20 PD0325901 (ȝM) %BrdU 10 Incorporation 0 0 6E-05 3E-04 0.002 0.008 0.04 0.2 1 5 25 MEKi (ȝM) B D HDQ-P1 Vehicle Treated 1PM MEKi 100 G1: 45.2% 1500 G1: 62.5% MCF7 1000 S: 4.2% S: 3.4% 75 BT474 800 G2/M: 34.9% 1000 G2/M: 27.4% MDA-MB-231 34.9 27.4 50 600 HDQ-P1 # Cells 4.17 #Cells 3.36 Cell viability (%) 400 CAL85-1 45.2 500 62.5 25 0.001 0.01 0.1 1 10 200 MEK inhibitor 2 (PM) 0 0 0 200 400 600 800 1000 0 200 400 600 800 1000 Propidium Iodide Propidium Iodide EGFR B A WEGFR Levels 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 --- MDA-MB-134VI Basal-like HDQ-P1 BT549 Basal-like Her2 DU4475 CAL51 + HCC1806 Supplemental Figure2:PTENandEGFRinBreastLines - MDA-MB-435s CAL85-1 + HCC70 CAL120 -- MX1 CAL148 CAL-120 + CAL85-1 HCC70 -- SW527 HCC1143 CAL-51 HCC1395 + MDA-MB-231 HCC1937 + HCC1143 + BT20 MDA-MB-231 + Hs578T MDA-MB-436 + MDA-MB-468 MDA-MB-468 - HCC1395 MFM223 + BT-549 - CAL-148 AU565 + HCC1937 BT474 + HCC38 EFM192A + MDA-MB-436 + HCC1954 HCC1419 + AU565 ------ HCC1569 BT474 Her2 orLuminal EFM-192A HCC1954 HCC1419 HCC2218 HCC1569 HCC202 MDA-MB-361 HCC2218 MDA-MB-453 + JIM-T --- SKBR3 KPL4 MDA-MB-361 UACC812 MDA-MB-453 UACC893 + SKBR3 ZR75-30 ---------- UACC-812 -

Defining Functional Interactions During Biogenesis of Epithelial Junctions
ARTICLE Received 11 Dec 2015 | Accepted 13 Oct 2016 | Published 6 Dec 2016 | Updated 5 Jan 2017 DOI: 10.1038/ncomms13542 OPEN Defining functional interactions during biogenesis of epithelial junctions J.C. Erasmus1,*, S. Bruche1,*,w, L. Pizarro1,2,*, N. Maimari1,3,*, T. Poggioli1,w, C. Tomlinson4,J.Lees5, I. Zalivina1,w, A. Wheeler1,w, A. Alberts6, A. Russo2 & V.M.M. Braga1 In spite of extensive recent progress, a comprehensive understanding of how actin cytoskeleton remodelling supports stable junctions remains to be established. Here we design a platform that integrates actin functions with optimized phenotypic clustering and identify new cytoskeletal proteins, their functional hierarchy and pathways that modulate E-cadherin adhesion. Depletion of EEF1A, an actin bundling protein, increases E-cadherin levels at junctions without a corresponding reinforcement of cell–cell contacts. This unexpected result reflects a more dynamic and mobile junctional actin in EEF1A-depleted cells. A partner for EEF1A in cadherin contact maintenance is the formin DIAPH2, which interacts with EEF1A. In contrast, depletion of either the endocytic regulator TRIP10 or the Rho GTPase activator VAV2 reduces E-cadherin levels at junctions. TRIP10 binds to and requires VAV2 function for its junctional localization. Overall, we present new conceptual insights on junction stabilization, which integrate known and novel pathways with impact for epithelial morphogenesis, homeostasis and diseases. 1 National Heart and Lung Institute, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. 2 Computing Department, Imperial College London, London SW7 2AZ, UK. 3 Bioengineering Department, Faculty of Engineering, Imperial College London, London SW7 2AZ, UK. 4 Department of Surgery & Cancer, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Myopathy Genes (HGNC) Neuropathy (HGNC) Neuromuscular Disease
Myopathy Genes Neuropathy Neuromuscular Disease (HGNC) (HGNC) (HGNC) ABHD5 ABCA1 ADCK3 ACTG2 ACO2 AGRN AGK AGXT ALS2 ALDOA AIFM1 ANG AMER1 ALAD AP4B1 ANO5 AMACR AP4E1 AR AP1S1 AP4M1 AUH APTX AP4S1 B4GALT1 AR AP5Z1 CACNA1S ATL3 ATM CASQ1 B4GALNT1 ATXN10 CCDC78 BAG3 ATXN7 CHCHD10 BRP44L BEAN1 CHRNA1 C12orf65 C9orf72 CHRNB1 C19orf12 CACNB4 CHRND C1NH CAPN3 CHRNE CECR1 CHAT CLPB CISD2 CHKB COL6A1 CLCF1 CHMP2B COL6A2 CLCN2 CHRNG COL6A3 CLP1 CLCN1 COLQ CMT2G COL9A3 CTNS CMT2H COQ2 DGUOK CMTDIA COQ6 DNA2 CMTX2 COQ9 DNAJB6 CMTX3 COX15 DNAJC19 COASY CPT1A DNM2 COX6A1 CYP7B1 DPM2 CPOX DAG1 DYSF CYP27A1 DDHD2 EMD CYP2U1 DOK7 EPG5 DARS2 DPAGT1 FAM111B DCAF8 DPM3 FBXL4 DDHD1 DUX4 FKBP14 DFNX5 ECEL1 FKRP DHTKD1 ERBB3 FLH1 DIAPH3 ERLIN2 FLNC DNAJB2 FA2H HNRNPA1 DNAJC3 FKTN HNRNPDL ELOVL5 FUS HNRPA2B1 ERCC8 G6PC KLHL40 FAH GFPT1 KLHL41 FAM126A GLE1 LAMA2 FBN1 GYS2 LDB3 FMR1 HSPD1 LMOD3 FXN IFRD1 MEGF10 GALC INF2 MGME1 GBE1 ISPD MTAP GJC2 ITGA7 MTMR14 GP1BA ITPR1 MYF6 HADHA KCNA1 MYH14 HADHB KCNC3 MYLK2 HFE KCNE3 NARS2 HINT1 KCNJ18 NEB HK1 KCNJ2 ORAI1 HMBS KIAA0196 PRKAG2 HSD17B4 KIF21A PTEN HSN1B L1CAM RBCK1 IARS2 LAMB2 RET IGHMBP2 LARGE RMND1 KCNJ10 MCCC2 SCN4A KIF5A MRE11A SERAC1 LRSAM1 MRPL3 SGCA LYST MTO1 SIL1 MANBA MTPAP SPEG MARS MTTP STAC3 MTATP6 MUSK STIM1 MYH14 MYBPC3 SYNE1 MYOT MYH3 SYNE2 NAMSD MYH8 TAZ NF2 NF1 TIA1 NGLY1 NIPA1 TMEM43 NMSR NOP56 TNPO3 NOTCH3 OPTN TNXB OPA1 PDSS2 TPM2 OPA3 PDYN TRPV4 OTOF PFN1 UBA1 PDK3 PHKA2 VCP PDSS1 PHKG2 XDH PEX10 PHOX2A ACADS PEX2 PIP5K1C ACADVL PMM2 PLEC ACTA1 PNPLA6 PLP1 AGL PPOX POMGNT1 AMPD1 PRICKLE1 -

Cldn19 Clic2 Clmp Cln3
NewbornDx™ Advanced Sequencing Evaluation When time to diagnosis matters, the NewbornDx™ Advanced Sequencing Evaluation from Athena Diagnostics delivers rapid, 5- to 7-day results on a targeted 1,722-genes. A2ML1 ALAD ATM CAV1 CLDN19 CTNS DOCK7 ETFB FOXC2 GLUL HOXC13 JAK3 AAAS ALAS2 ATP1A2 CBL CLIC2 CTRC DOCK8 ETFDH FOXE1 GLYCTK HOXD13 JUP AARS2 ALDH18A1 ATP1A3 CBS CLMP CTSA DOK7 ETHE1 FOXE3 GM2A HPD KANK1 AASS ALDH1A2 ATP2B3 CC2D2A CLN3 CTSD DOLK EVC FOXF1 GMPPA HPGD K ANSL1 ABAT ALDH3A2 ATP5A1 CCDC103 CLN5 CTSK DPAGT1 EVC2 FOXG1 GMPPB HPRT1 KAT6B ABCA12 ALDH4A1 ATP5E CCDC114 CLN6 CUBN DPM1 EXOC4 FOXH1 GNA11 HPSE2 KCNA2 ABCA3 ALDH5A1 ATP6AP2 CCDC151 CLN8 CUL4B DPM2 EXOSC3 FOXI1 GNAI3 HRAS KCNB1 ABCA4 ALDH7A1 ATP6V0A2 CCDC22 CLP1 CUL7 DPM3 EXPH5 FOXL2 GNAO1 HSD17B10 KCND2 ABCB11 ALDOA ATP6V1B1 CCDC39 CLPB CXCR4 DPP6 EYA1 FOXP1 GNAS HSD17B4 KCNE1 ABCB4 ALDOB ATP7A CCDC40 CLPP CYB5R3 DPYD EZH2 FOXP2 GNE HSD3B2 KCNE2 ABCB6 ALG1 ATP8A2 CCDC65 CNNM2 CYC1 DPYS F10 FOXP3 GNMT HSD3B7 KCNH2 ABCB7 ALG11 ATP8B1 CCDC78 CNTN1 CYP11B1 DRC1 F11 FOXRED1 GNPAT HSPD1 KCNH5 ABCC2 ALG12 ATPAF2 CCDC8 CNTNAP1 CYP11B2 DSC2 F13A1 FRAS1 GNPTAB HSPG2 KCNJ10 ABCC8 ALG13 ATR CCDC88C CNTNAP2 CYP17A1 DSG1 F13B FREM1 GNPTG HUWE1 KCNJ11 ABCC9 ALG14 ATRX CCND2 COA5 CYP1B1 DSP F2 FREM2 GNS HYDIN KCNJ13 ABCD3 ALG2 AUH CCNO COG1 CYP24A1 DST F5 FRMD7 GORAB HYLS1 KCNJ2 ABCD4 ALG3 B3GALNT2 CCS COG4 CYP26C1 DSTYK F7 FTCD GP1BA IBA57 KCNJ5 ABHD5 ALG6 B3GAT3 CCT5 COG5 CYP27A1 DTNA F8 FTO GP1BB ICK KCNJ8 ACAD8 ALG8 B3GLCT CD151 COG6 CYP27B1 DUOX2 F9 FUCA1 GP6 ICOS KCNK3 ACAD9 ALG9 -
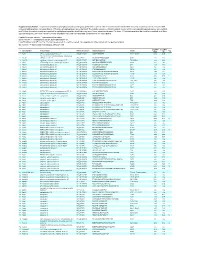
Supplementary Table 1
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding -

Biological Models of Colorectal Cancer Metastasis and Tumor Suppression
BIOLOGICAL MODELS OF COLORECTAL CANCER METASTASIS AND TUMOR SUPPRESSION PROVIDE MECHANISTIC INSIGHTS TO GUIDE PERSONALIZED CARE OF THE COLORECTAL CANCER PATIENT By Jesse Joshua Smith Dissertation Submitted to the Faculty of the Graduate School of Vanderbilt University In partial fulfillment of the requirements For the degree of DOCTOR OF PHILOSOPHY In Cell and Developmental Biology May, 2010 Nashville, Tennessee Approved: Professor R. Daniel Beauchamp Professor Robert J. Coffey Professor Mark deCaestecker Professor Ethan Lee Professor Steven K. Hanks Copyright 2010 by Jesse Joshua Smith All Rights Reserved To my grandparents, Gladys and A.L. Lyth and Juanda Ruth and J.E. Smith, fully supportive and never in doubt. To my amazing and enduring parents, Rebecca Lyth and Jesse E. Smith, Jr., always there for me. .my sure foundation. To Jeannine, Bill and Reagan for encouragement, patience, love, trust and a solid backing. To Granny George and Shawn for loving support and care. And To my beautiful wife, Kelly, My heart, soul and great love, Infinitely supportive, patient and graceful. ii ACKNOWLEDGEMENTS This work would not have been possible without the financial support of the Vanderbilt Medical Scientist Training Program through the Clinical and Translational Science Award (Clinical Investigator Track), the Society of University Surgeons-Ethicon Scholarship Fund and the Surgical Oncology T32 grant and the Vanderbilt Medical Center Section of Surgical Sciences and the Department of Surgical Oncology. I am especially indebted to Drs. R. Daniel Beauchamp, Chairman of the Section of Surgical Sciences, Dr. James R. Goldenring, Vice Chairman of Research of the Department of Surgery, Dr. Naji N. -

Identification of the Fatty Acid Synthase Interaction Network Via Itraq-Based Proteomics Indicates the Potential Molecular Mecha
Huang et al. Cancer Cell Int (2020) 20:332 https://doi.org/10.1186/s12935-020-01409-2 Cancer Cell International PRIMARY RESEARCH Open Access Identifcation of the fatty acid synthase interaction network via iTRAQ-based proteomics indicates the potential molecular mechanisms of liver cancer metastasis Juan Huang1, Yao Tang1, Xiaoqin Zou1, Yi Lu1, Sha She1, Wenyue Zhang1, Hong Ren1, Yixuan Yang1,2* and Huaidong Hu1,2* Abstract Background: Fatty acid synthase (FASN) is highly expressed in various types of cancer and has an important role in carcinogenesis and metastasis. To clarify the mechanisms of FASN in liver cancer invasion and metastasis, the FASN protein interaction network in liver cancer was identifed by targeted proteomic analysis. Methods: Wound healing and Transwell assays was performed to observe the efect of FASN during migration and invasion in liver cancer. Isobaric tags for relative and absolute quantitation (iTRAQ)-based mass spectrometry were used to identify proteins interacting with FASN in HepG2 cells. Diferential expressed proteins were validated by co-immunoprecipitation, western blot analyses and confocal microscopy. Western blot and reverse transcription- quantitative polymerase chain reaction (RT-qPCR) were performed to demonstrate the mechanism of FASN regulating metastasis. Results: FASN knockdown inhibited migration and invasion of HepG2 and SMMC7721 cells. A total of, 79 proteins interacting with FASN were identifed. Additionally, gene ontology term enrichment analysis indicated that the majority of biological regulation and cellular processes that the FASN-interacting proteins were associated with. Co- precipitation and co-localization of FASN with fascin actin-bundling protein 1 (FSCN1), signal-induced proliferation- associated 1 (SIPA1), spectrin β, non-erythrocytic 1 (SPTBN1) and CD59 were evaluated. -

Common Genetic Aberrations Associated with Metabolic Interferences in Human Type-2 Diabetes and Acute Myeloid Leukemia: a Bioinformatics Approach
International Journal of Molecular Sciences Article Common Genetic Aberrations Associated with Metabolic Interferences in Human Type-2 Diabetes and Acute Myeloid Leukemia: A Bioinformatics Approach Theodora-Christina Kyriakou 1, Panagiotis Papageorgis 1,2 and Maria-Ioanna Christodoulou 3,* 1 Tumor Microenvironment, Metastasis and Experimental Therapeutics Laboratory, Basic and Translational Cancer Research Center, Department of Life Sciences, European University Cyprus, Nicosia 2404, Cyprus; [email protected] (T.-C.K.); [email protected] (P.P.) 2 European University Cyprus Research Center, Nicosia 2404, Cyprus 3 Tumor Immunology and Biomarkers Laboratory, Basic and Translational Cancer Research Center, Department of Life Sciences, European University Cyprus, Nicosia 2404, Cyprus * Correspondence: [email protected] Abstract: Type-2 diabetes mellitus (T2D) is a chronic metabolic disorder, associated with an increased risk of developing solid tumors and hematological malignancies, including acute myeloid leukemia (AML). However, the genetic background underlying this predisposition remains elusive. We herein aimed at the exploration of the genetic variants, related transcriptomic changes and disturbances in metabolic pathways shared by T2D and AML, utilizing bioinformatics tools and repositories, as well as publicly available clinical datasets. Our approach revealed that rs11709077 and rs1801282, on PPARG, rs11108094 on USP44, rs6685701 on RPS6KA1 and rs7929543 on AC118942.1 comprise Citation: Kyriakou, T.-C.; common SNPs susceptible to the two diseases and, together with 64 other co-inherited proxy SNPs, Papageorgis, P.; Christodoulou, M.-I. may affect the expression patterns of metabolic genes, such as USP44, METAP2, PPARG, TIMP4 and Common Genetic Aberrations RPS6KA1, in adipose tissue, skeletal muscle, liver, pancreas and whole blood. -

Actinin-4 Reveals a Mechanism for Regulating Its F-Actin-Binding Affinity
Disease-associated mutant ␣-actinin-4 reveals a mechanism for regulating its F-actin-binding affinity Astrid Weins*, Johannes S. Schlondorff*, Fumihiko Nakamura†, Bradley M. Denker*, John H. Hartwig†, Thomas P. Stossel†, and Martin R. Pollak*‡ *Renal and †Translational Medicine Divisions, Department of Medicine, Brigham and Women’s Hospital and Harvard Medical School, Boston, MA 02115 Edited by Thomas D. Pollard, Yale University, New Haven, CT, and approved August 23, 2007 (received for review March 16, 2007) ␣-Actinin-4 is a widely expressed protein that employs an actin- them cell motility, endocytosis, and adhesion (7–9). Cultured binding site with two calponin homology domains to crosslink podocytes deficient in one of the nonmuscle isotypes, ␣-actinin-4, actin filaments (F-actin) in a Ca2؉-sensitive manner in vitro.An exhibit defective substrate adhesion (10), which is consistent with inherited, late-onset form of kidney failure is caused by point the well documented localization of ␣-actinin at focal adhesion sites mutations in the ␣-actinin-4 actin-binding domain. Here we show (11), and is therefore probably responsible for the impaired trans- that ␣-actinin-4/F-actin aggregates, observed in vivo in podocytes lational locomotion of the ␣-actinin-4-deficient cells (7). of humans and mice with disease, likely form as a direct result of Five distinct point mutations in the ␣-actinin-4 head domain the increased actin-binding affinity of the protein. We document causing focal segmental glomerulosclerosis in affected humans all that exposure of a buried actin-binding site 1 in mutant ␣-actinin-4 mediate increased actin binding (12, 13). However, this effect has causes an increase in its actin-binding affinity, abolishes its Ca2؉ not yet been studied in detail, and the mechanism of disease has so regulation in vitro, and diverts its normal localization from actin far remained elusive. -

Supplementary Material Contents
Supplementary Material Contents Immune modulating proteins identified from exosomal samples.....................................................................2 Figure S1: Overlap between exosomal and soluble proteomes.................................................................................... 4 Bacterial strains:..............................................................................................................................................4 Figure S2: Variability between subjects of effects of exosomes on BL21-lux growth.................................................... 5 Figure S3: Early effects of exosomes on growth of BL21 E. coli .................................................................................... 5 Figure S4: Exosomal Lysis............................................................................................................................................ 6 Figure S5: Effect of pH on exosomal action.................................................................................................................. 7 Figure S6: Effect of exosomes on growth of UPEC (pH = 6.5) suspended in exosome-depleted urine supernatant ....... 8 Effective exosomal concentration....................................................................................................................8 Figure S7: Sample constitution for luminometry experiments..................................................................................... 8 Figure S8: Determining effective concentration ......................................................................................................... -

Genomics of Inherited Bone Marrow Failure and Myelodysplasia Michael
Genomics of inherited bone marrow failure and myelodysplasia Michael Yu Zhang A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy University of Washington 2015 Reading Committee: Mary-Claire King, Chair Akiko Shimamura Marshall Horwitz Program Authorized to Offer Degree: Molecular and Cellular Biology 1 ©Copyright 2015 Michael Yu Zhang 2 University of Washington ABSTRACT Genomics of inherited bone marrow failure and myelodysplasia Michael Yu Zhang Chair of the Supervisory Committee: Professor Mary-Claire King Department of Medicine (Medical Genetics) and Genome Sciences Bone marrow failure and myelodysplastic syndromes (BMF/MDS) are disorders of impaired blood cell production with increased leukemia risk. BMF/MDS may be acquired or inherited, a distinction critical for treatment selection. Currently, diagnosis of these inherited syndromes is based on clinical history, family history, and laboratory studies, which directs the ordering of genetic tests on a gene-by-gene basis. However, despite extensive clinical workup and serial genetic testing, many cases remain unexplained. We sought to define the genetic etiology and pathophysiology of unclassified bone marrow failure and myelodysplastic syndromes. First, to determine the extent to which patients remained undiagnosed due to atypical or cryptic presentations of known inherited BMF/MDS, we developed a massively-parallel, next- generation DNA sequencing assay to simultaneously screen for mutations in 85 BMF/MDS genes. Querying 71 pediatric and adult patients with unclassified BMF/MDS using this assay revealed 8 (11%) patients with constitutional, pathogenic mutations in GATA2 , RUNX1 , DKC1 , or LIG4 . All eight patients lacked classic features or laboratory findings for their syndromes.