Type 2 Diabetes Modifies Skeletal Muscle Gene Expression Response to Gastric Bypass Surgery
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Sanjay Kumar Gupta
The human CCHC-type Zinc Finger Nucleic Acid Binding Protein (CNBP) binds to the G-rich elements in target mRNA coding sequences and promotes translation Das humane CCHC-Typ-Zinkfinger-Nukleinsäure-Binde-Protein (CNBP) bindet an G-reiche Elemente in der kodierenden Sequenz seiner Ziel-mRNAs und fördert deren Translation Doctoral thesis for a doctoral degree at the Graduate School of Life Sciences, Julius-Maximilians-Universität WürzBurg, Section: Biomedicine suBmitted By Sanjay Kumar Gupta from Varanasi, India WürzBurg, 2016 1 Submitted on: …………………………………………………………..…….. Office stamp Members of the Promotionskomitee: Chairperson: Prof. Dr. Alexander Buchberger Primary Supervisor: Dr. Stefan Juranek Supervisor (Second): Prof. Dr. Utz Fischer Supervisor (Third): Dr. Markus Landthaler Date of Public Defence: …………………………………………….………… Date of Receipt of Certificates: ………………………………………………. 2 Summary The genetic information encoded with in the genes are transcribed and translated to give rise to the functional proteins, which are building block of a cell. At first, it was thought that the regulation of gene expression particularly occurs at the level of transcription By various transcription factors. Recent discoveries have shown the vital role of gene regulation at the level of RNA also known as post-transcriptional gene regulation (PTGR). Apart from non-coding RNAs e.g. micro RNAs, various RNA Binding proteins (RBPs) play essential role in PTGR. RBPs have been implicated in different stages of mRNA life cycle ranging from splicing, processing, transport, localization and decay. In last 20 years studies have shown the presence of hundreds of RBPs across eukaryotic systems many of which are widely conserved. Given the rising numBer of RBPs and their link to human diseases it is quite evident that RBPs have major role in cellular processes and their regulation. -

PDF Output of CLIC (Clustering by Inferred Co-Expression)
PDF Output of CLIC (clustering by inferred co-expression) Dataset: Num of genes in input gene set: 7 Total number of genes: 16493 CLIC PDF output has three sections: 1) Overview of Co-Expression Modules (CEMs) Heatmap shows pairwise correlations between all genes in the input query gene set. Red lines shows the partition of input genes into CEMs, ordered by CEM strength. Each row shows one gene, and the brightness of squares indicates its correlations with other genes. Gene symbols are shown at left side and on the top of the heatmap. 2) Details of each CEM and its expansion CEM+ Top panel shows the posterior selection probability (dataset weights) for top GEO series datasets. Bottom panel shows the CEM genes (blue rows) as well as expanded CEM+ genes (green rows). Each column is one GEO series dataset, sorted by their posterior probability of being selected. The brightness of squares indicates the gene's correlations with CEM genes in the corresponding dataset. CEM+ includes genes that co-express with CEM genes in high-weight datasets, measured by LLR score. 3) Details of each GEO series dataset and its expression profile: Top panel shows the detailed information (e.g. title, summary) for the GEO series dataset. Bottom panel shows the background distribution and the expression profile for CEM genes in this dataset. Overview of Co-Expression Modules (CEMs) with Dataset Weighting Scale of average Pearson correlations Num of Genes in Query Geneset: 7. Num of CEMs: 1. 0.0 0.2 0.4 0.6 0.8 1.0 Ctr9 Rtf1 Paf1 Cdc73 Leo1 Wdr61 Pex2 Ctr9 Rtf1 -

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -

Targeting MYCN in Neuroblastoma by BET Bromodomain Inhibition
Published OnlineFirst February 21, 2013; DOI: 10.1158/2159-8290.CD-12-0418 RESEARCH ARTICLE Targeting MYCN in Neuroblastoma by BET Bromodomain Inhibition Alexandre Puissant1,3, Stacey M. Frumm1,3, Gabriela Alexe1,3,5,6, Christopher F. Bassil1,3, Jun Qi2, Yvan H. Chanthery8, Erin A. Nekritz8, Rhamy Zeid2, William Clay Gustafson8, Patricia Greninger7, Matthew J. Garnett10, Ultan McDermott10, Cyril H. Benes7, Andrew L. Kung1,3, William A. Weiss8,9, James E. Bradner2,4, and Kimberly Stegmaier1,3,6 Downloaded from cancerdiscovery.aacrjournals.org on October 2, 2021. © 2013 American Association for Cancer Research. 15-CD-12-0418_p308-323.indd 1 22/02/13 12:15 AM Published OnlineFirst February 21, 2013; DOI: 10.1158/2159-8290.CD-12-0418 A BSTRACT Bromodomain inhibition comprises a promising therapeutic strategy in cancer, particularly for hematologic malignancies. To date, however, genomic biomarkers to direct clinical translation have been lacking. We conducted a cell-based screen of genetically defined cancer cell lines using a prototypical inhibitor of BET bromodomains. Integration of genetic features with chemosensitivity data revealed a robust correlation between MYCN amplification and sensitivity to bromodomain inhibition. We characterized the mechanistic and translational significance of this finding in neuroblastoma, a childhood cancer with frequent amplification of MYCN. Genome-wide expression analysis showed downregulation of the MYCN transcriptional program accompanied by suppression of MYCN transcription. Functionally, bromodomain-mediated inhibition of MYCN impaired growth and induced apoptosis in neuroblastoma. BRD4 knockdown phenocopied these effects, establishing BET bromodomains as transcriptional regulators of MYCN. BET inhibition conferred a significant survival advantage in 3 in vivo neuroblastoma models, providing a compelling rationale for developing BET bro- modomain inhibitors in patients with neuroblastoma. -

Genome-Wide Determination of On-Target and Off-Target Characteristics for RNA-Guided DNA Methylation by Dcas9 Methyltransferases.” Gigascience 7 (3): Giy011
Genome-wide determination of on- target and off-target characteristics for RNA-guided DNA methylation by dCas9 methyltransferases The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Lin, L., Y. Liu, F. Xu, J. Huang, T. F. Daugaard, T. S. Petersen, B. Hansen, et al. 2018. “Genome-wide determination of on-target and off-target characteristics for RNA-guided DNA methylation by dCas9 methyltransferases.” GigaScience 7 (3): giy011. doi:10.1093/ gigascience/giy011. http://dx.doi.org/10.1093/gigascience/giy011. Published Version doi:10.1093/gigascience/giy011 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:37068160 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA GigaScience, 7, 2018, 1–19 doi: 10.1093/gigascience/giy011 Advance Access Publication Date: 19 February 2018 Research RESEARCH Genome-wide determination of on-target and off-target characteristics for RNA-guided DNA methylation by dCas9 methyltransferases Lin Lin1,2,†,YongLiu1,†, Fengping Xu3,4,5,†,JinrongHuang3,4,5,†, Tina Fuglsang Daugaard1, Trine Skov Petersen1, Bettina Hansen1, Lingfei Ye3,QingZhou3,4, Fang Fang3,4,LingYang3,4, Shengting Li1,3, Lasse Fløe1, Kristopher Torp Jensen1, Ellen Shrock6, Fang Chen3,4,5, Huanming Yang3,4,7,JianWang3,4,XinLiu3,4,XunXu3,4,∗, Lars Bolund1,3,4,2,8,9, Anders Lade Nielsen1 and Yonglun Luo1,3,4,2,8,9,10,∗ 1Department of Biomedicine, Aarhus University, Aarhus, Denmark, 2Danish Regenerative Engineering Alliance for Medicine, Department of Biomedicine, Aarhus University, Aarhus, Denmark, 3BGI-Shenzhen, Shenzhen 518083, China, 4China National GeneBank-Shenzhen, BGI-Research, Shenzhen 518083, China, 5Department of Biology, University of Copenhagen, Copenhagen, Denmark, 6Department of Genetics, Harvard Medical School, Boston, MA, USA, 7James D. -

Literature Mining Sustains and Enhances Knowledge Discovery from Omic Studies
LITERATURE MINING SUSTAINS AND ENHANCES KNOWLEDGE DISCOVERY FROM OMIC STUDIES by Rick Matthew Jordan B.S. Biology, University of Pittsburgh, 1996 M.S. Molecular Biology/Biotechnology, East Carolina University, 2001 M.S. Biomedical Informatics, University of Pittsburgh, 2005 Submitted to the Graduate Faculty of School of Medicine in partial fulfillment of the requirements for the degree of Doctor of Philosophy University of Pittsburgh 2016 UNIVERSITY OF PITTSBURGH SCHOOL OF MEDICINE This dissertation was presented by Rick Matthew Jordan It was defended on December 2, 2015 and approved by Shyam Visweswaran, M.D., Ph.D., Associate Professor Rebecca Jacobson, M.D., M.S., Professor Songjian Lu, Ph.D., Assistant Professor Dissertation Advisor: Vanathi Gopalakrishnan, Ph.D., Associate Professor ii Copyright © by Rick Matthew Jordan 2016 iii LITERATURE MINING SUSTAINS AND ENHANCES KNOWLEDGE DISCOVERY FROM OMIC STUDIES Rick Matthew Jordan, M.S. University of Pittsburgh, 2016 Genomic, proteomic and other experimentally generated data from studies of biological systems aiming to discover disease biomarkers are currently analyzed without sufficient supporting evidence from the literature due to complexities associated with automated processing. Extracting prior knowledge about markers associated with biological sample types and disease states from the literature is tedious, and little research has been performed to understand how to use this knowledge to inform the generation of classification models from ‘omic’ data. Using pathway analysis methods to better understand the underlying biology of complex diseases such as breast and lung cancers is state-of-the-art. However, the problem of how to combine literature- mining evidence with pathway analysis evidence is an open problem in biomedical informatics research. -

Mitochondrial Genetics
Mitochondrial genetics Patrick Francis Chinnery and Gavin Hudson* Institute of Genetic Medicine, International Centre for Life, Newcastle University, Central Parkway, Newcastle upon Tyne NE1 3BZ, UK Introduction: In the last 10 years the field of mitochondrial genetics has widened, shifting the focus from rare sporadic, metabolic disease to the effects of mitochondrial DNA (mtDNA) variation in a growing spectrum of human disease. The aim of this review is to guide the reader through some key concepts regarding mitochondria before introducing both classic and emerging mitochondrial disorders. Sources of data: In this article, a review of the current mitochondrial genetics literature was conducted using PubMed (http://www.ncbi.nlm.nih.gov/pubmed/). In addition, this review makes use of a growing number of publically available databases including MITOMAP, a human mitochondrial genome database (www.mitomap.org), the Human DNA polymerase Gamma Mutation Database (http://tools.niehs.nih.gov/polg/) and PhyloTree.org (www.phylotree.org), a repository of global mtDNA variation. Areas of agreement: The disruption in cellular energy, resulting from defects in mtDNA or defects in the nuclear-encoded genes responsible for mitochondrial maintenance, manifests in a growing number of human diseases. Areas of controversy: The exact mechanisms which govern the inheritance of mtDNA are hotly debated. Growing points: Although still in the early stages, the development of in vitro genetic manipulation could see an end to the inheritance of the most severe mtDNA disease. Keywords: mitochondria/genetics/mitochondrial DNA/mitochondrial disease/ mtDNA Accepted: April 16, 2013 Mitochondria *Correspondence address. The mitochondrion is a highly specialized organelle, present in almost all Institute of Genetic Medicine, International eukaryotic cells and principally charged with the production of cellular Centre for Life, Newcastle energy through oxidative phosphorylation (OXPHOS). -
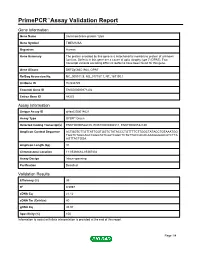
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name transmembrane protein 126A Gene Symbol TMEM126A Organism Human Gene Summary The protein encoded by this gene is a mitochondrial membrane protein of unknown function. Defects in this gene are a cause of optic atrophy type 7 (OPA7). Two transcript variants encoding different isoforms have been found for this gene. Gene Aliases DKFZp586C1924, OPA7 RefSeq Accession No. NC_000011.9, NG_017157.1, NT_167190.1 UniGene ID Hs.533725 Ensembl Gene ID ENSG00000171202 Entrez Gene ID 84233 Assay Information Unique Assay ID qHsaCID0014621 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000528105, ENST00000304511, ENST00000532180 Amplicon Context Sequence ACTGGTCTTGTTATTGGTGGTCTATACCCTGTTTTCTTGGCTATACCTGTAAATGG TGGTCTAGCAGCCAGGTATCAATCAGCTCTGTTACCACACAAAGGGAACATCTTA AGTTACTGGA Amplicon Length (bp) 91 Chromosome Location 11:85366682-85367402 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 99 R2 0.9997 cDNA Cq 21.12 cDNA Tm (Celsius) 80 gDNA Cq 35.93 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 1/4 PrimePCR™Assay Validation Report TMEM126A, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 2/4 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -

Supplementary File 1
Supplementary Materials Figure S1. RNA sequencing quality control report. The upper panel showed per base sequence quality of (a) miRNA from 786-O and ACHN, and (b) RNA from 786-O and ACHN; the lower panel showed the per sequence quality scores of (c) miRNA from 786-O and ACHN, and (d) RNA from 786-O and ACHN. The quality score from 15 to 99 bp fell into the green background, indicating good quality calls. The majority read > 35 indicated good sequencing quality. Table S1. The list of miRNA with significant changes (ACHN vs 786-O) #miRNA Fold Change #miRNA Fold Change #miRNA Fold Change hsa-let-7a-2-3p -490.00 hsa-miR-192-5p 21.37 hsa-miR-323a-3p 135.76 hsa-let-7c-3p -10.30 hsa-miR-194-3p 31.18 hsa-miR-323b-3p 53.75 hsa-miR-100-3p -26.39 hsa-miR-194-5p 21.87 hsa-miR-326 -16.46 hsa-miR-100-5p -63.93 hsa-miR-195-5p 8.31 hsa-miR-329-3p 101.81 hsa-miR-105-5p -288.00 hsa-miR-196a-3p 5.28 hsa-miR-329-5p 436 hsa-miR-1185-1-3p 32.08 hsa-miR-199b-5p -9.75 hsa-miR-335-3p 73.67 hsa-miR-1185-2-3p 37.22 hsa-miR-203a-3p 222.25 hsa-miR-335-5p 71.58 hsa-miR-1185-5p 91.13 hsa-miR-205-5p -5.12 hsa-miR-337-3p 1696 hsa-miR-1197 7.38 hsa-miR-20b-5p 5.28 hsa-miR-337-5p 12141 hsa-miR-125b-1-3p -301.58 hsa-miR-211-5p 118 hsa-miR-34b-3p 1110.62 hsa-miR-125b-2-3p 5.91 hsa-miR-215-5p 18.88 hsa-miR-34b-5p 796.38 hsa-miR-1262 16.28 hsa-miR-218-5p 14.37 hsa-miR-34c-3p 10751 hsa-miR-1266-5p 6.62 hsa-miR-224-5p 118 hsa-miR-34c-5p 836.81 hsa-miR-127-3p 1723.39 hsa-miR-2277-3p 6.65 hsa-miR-3614-5p 141 hsa-miR-127-5p 4334.00 hsa-miR-2682-3p 6.63 hsa-miR-3617-5p -140 hsa-miR-1271-5p -

Novel Gene Discovery in Primary Ciliary Dyskinesia
Novel Gene Discovery in Primary Ciliary Dyskinesia Mahmoud Raafat Fassad Genetics and Genomic Medicine Programme Great Ormond Street Institute of Child Health University College London A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy University College London 1 Declaration I, Mahmoud Raafat Fassad, confirm that the work presented in this thesis is my own. Where information has been derived from other sources, I confirm that this has been indicated in the thesis. 2 Abstract Primary Ciliary Dyskinesia (PCD) is one of the ‘ciliopathies’, genetic disorders affecting either cilia structure or function. PCD is a rare recessive disease caused by defective motile cilia. Affected individuals manifest with neonatal respiratory distress, chronic wet cough, upper respiratory tract problems, progressive lung disease resulting in bronchiectasis, laterality problems including heart defects and adult infertility. Early diagnosis and management are essential for better respiratory disease prognosis. PCD is a highly genetically heterogeneous disorder with causal mutations identified in 36 genes that account for the disease in about 70% of PCD cases, suggesting that additional genes remain to be discovered. Targeted next generation sequencing was used for genetic screening of a cohort of patients with confirmed or suggestive PCD diagnosis. The use of multi-gene panel sequencing yielded a high diagnostic output (> 70%) with mutations identified in known PCD genes. Over half of these mutations were novel alleles, expanding the mutation spectrum in PCD genes. The inclusion of patients from various ethnic backgrounds revealed a striking impact of ethnicity on the composition of disease alleles uncovering a significant genetic stratification of PCD in different populations. -

RNA Splicing in the Transition from B Cells
RNA Splicing in the Transition from B Cells to Antibody-Secreting Cells: The Influences of ELL2, Small Nuclear RNA, and Endoplasmic Reticulum Stress This information is current as of September 24, 2021. Ashley M. Nelson, Nolan T. Carew, Sage M. Smith and Christine Milcarek J Immunol 2018; 201:3073-3083; Prepublished online 8 October 2018; doi: 10.4049/jimmunol.1800557 Downloaded from http://www.jimmunol.org/content/201/10/3073 Supplementary http://www.jimmunol.org/content/suppl/2018/10/05/jimmunol.180055 http://www.jimmunol.org/ Material 7.DCSupplemental References This article cites 44 articles, 16 of which you can access for free at: http://www.jimmunol.org/content/201/10/3073.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision by guest on September 24, 2021 • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2018 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology RNA Splicing in the Transition from B Cells to Antibody-Secreting Cells: The Influences of ELL2, Small Nuclear RNA, and Endoplasmic Reticulum Stress Ashley M. -

Combinatorial Strategies Using CRISPR/Cas9 for Gene Mutagenesis in Adult Mice
Combinatorial strategies using CRISPR/Cas9 for gene mutagenesis in adult mice Avery C. Hunker A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy University of Washington 2019 Reading Committee: Larry S. Zweifel, Chair Sheri J. Mizumori G. Stanley McKnight Program Authorized to Offer Degree: Pharmacology 2 © Copyright 2019 Avery C. Hunker 3 University of Washington ABSTRACT Combinatorial strategies using CRISPR/Cas9 for gene mutagenesis in adult mice Avery C. Hunker Chair of the Supervisory Committee: Larry Zweifel Department of Pharmacology A major challenge to understanding how genes modulate complex behaviors is the inability to restrict genetic manipulations to defined cell populations or circuits. To circumvent this, we created a simple strategy for limiting gene knockout to specific cell populations using a viral-mediated, conditional CRISPR/SaCas9 system in combination with intersectional genetic strategies. A small single guide RNA (sgRNA) directs Staphylococcus aureus CRISPR-associated protein (SaCas9) to unique sites on DNA in a Cre-dependent manner resulting in double strand breaks and gene mutagenesis in vivo. To validate this technique we targeted nine different genes of diverse function in distinct cell types in mice and performed an array of analyses to confirm gene mutagenesis and subsequent protein loss, including IHC, cell-type specific DNA sequencing, electrophysiology, Western blots, and behavior. We show that these vectors are as efficient as conventional conditional gene knockout and provide a viable alternative to complex genetic crosses. This strategy provides additional benefits of 4 targeting gene mutagenesis to cell types previously difficult to isolate, and the ability to target genes in specific neural projections for gene inactivation.