Insertion Polymorphisms of Mobile Genetic Elements in Sexual and Asexual Populations of Daphnia Pulex
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
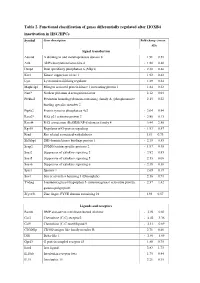
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -
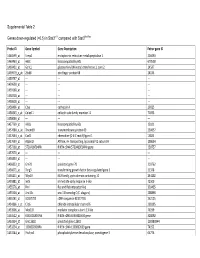
0.5) in Stat3∆/∆ Compared with Stat3flox/Flox
Supplemental Table 2 Genes down-regulated (<0.5) in Stat3∆/∆ compared with Stat3flox/flox Probe ID Gene Symbol Gene Description Entrez gene ID 1460599_at Ermp1 endoplasmic reticulum metallopeptidase 1 226090 1460463_at H60c histocompatibility 60c 670558 1460431_at Gcnt1 glucosaminyl (N-acetyl) transferase 1, core 2 14537 1459979_x_at Zfp68 zinc finger protein 68 24135 1459747_at --- --- --- 1459608_at --- --- --- 1459168_at --- --- --- 1458718_at --- --- --- 1458618_at --- --- --- 1458466_at Ctsa cathepsin A 19025 1458345_s_at Colec11 collectin sub-family member 11 71693 1458046_at --- --- --- 1457769_at H60a histocompatibility 60a 15101 1457680_a_at Tmem69 transmembrane protein 69 230657 1457644_s_at Cxcl1 chemokine (C-X-C motif) ligand 1 14825 1457639_at Atp6v1h ATPase, H+ transporting, lysosomal V1 subunit H 108664 1457260_at 5730409E04Rik RIKEN cDNA 5730409E04Rik gene 230757 1457070_at --- --- --- 1456893_at --- --- --- 1456823_at Gm70 predicted gene 70 210762 1456671_at Tbrg3 transforming growth factor beta regulated gene 3 21378 1456211_at Nlrp10 NLR family, pyrin domain containing 10 244202 1455881_at Ier5l immediate early response 5-like 72500 1455576_at Rinl Ras and Rab interactor-like 320435 1455304_at Unc13c unc-13 homolog C (C. elegans) 208898 1455241_at BC037703 cDNA sequence BC037703 242125 1454866_s_at Clic6 chloride intracellular channel 6 209195 1453906_at Med13l mediator complex subunit 13-like 76199 1453522_at 6530401N04Rik RIKEN cDNA 6530401N04 gene 328092 1453354_at Gm11602 predicted gene 11602 100380944 1453234_at -

4-6 Weeks Old Female C57BL/6 Mice Obtained from Jackson Labs Were Used for Cell Isolation
Methods Mice: 4-6 weeks old female C57BL/6 mice obtained from Jackson labs were used for cell isolation. Female Foxp3-IRES-GFP reporter mice (1), backcrossed to B6/C57 background for 10 generations, were used for the isolation of naïve CD4 and naïve CD8 cells for the RNAseq experiments. The mice were housed in pathogen-free animal facility in the La Jolla Institute for Allergy and Immunology and were used according to protocols approved by the Institutional Animal Care and use Committee. Preparation of cells: Subsets of thymocytes were isolated by cell sorting as previously described (2), after cell surface staining using CD4 (GK1.5), CD8 (53-6.7), CD3ε (145- 2C11), CD24 (M1/69) (all from Biolegend). DP cells: CD4+CD8 int/hi; CD4 SP cells: CD4CD3 hi, CD24 int/lo; CD8 SP cells: CD8 int/hi CD4 CD3 hi, CD24 int/lo (Fig S2). Peripheral subsets were isolated after pooling spleen and lymph nodes. T cells were enriched by negative isolation using Dynabeads (Dynabeads untouched mouse T cells, 11413D, Invitrogen). After surface staining for CD4 (GK1.5), CD8 (53-6.7), CD62L (MEL-14), CD25 (PC61) and CD44 (IM7), naïve CD4+CD62L hiCD25-CD44lo and naïve CD8+CD62L hiCD25-CD44lo were obtained by sorting (BD FACS Aria). Additionally, for the RNAseq experiments, CD4 and CD8 naïve cells were isolated by sorting T cells from the Foxp3- IRES-GFP mice: CD4+CD62LhiCD25–CD44lo GFP(FOXP3)– and CD8+CD62LhiCD25– CD44lo GFP(FOXP3)– (antibodies were from Biolegend). In some cases, naïve CD4 cells were cultured in vitro under Th1 or Th2 polarizing conditions (3, 4). -

Downloaded from USCS Tables (
bioRxiv preprint doi: https://doi.org/10.1101/318329; this version posted February 4, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. The selection arena in early human blastocysts resolves the pluripotent inner cell mass Manvendra Singh1, Thomas J. Widmann2, Vikas Bansal7, Jose L. Cortes2, Gerald G. Schumann3, Stephanie Wunderlich4, Ulrich Martin4, Marta Garcia-Canadas2, Jose L. Garcia- Perez2,5*, Laurence D. Hurst6*#, Zsuzsanna Izsvák1* 1 Max-Delbrück-Center for Molecular Medicine in the Helmholtz Society, Robert-Rössle- Strasse 10, 13125 Berlin, Germany. 2 GENYO. Centre for Genomics and Oncological Research: Pfizer/University of Granada/Andalusian Regional Government, PTS Granada, 18016 Granada, Spain. 3 Paul-Ehrlich-Institute, Division of Medical Biotechnology, Paul-Ehrlich-Strasse 51-59, 63225 Langen, Germany. 4Center for Regenerative Medicine Hannover Medical School (MHH) Carl-Neuberg-Str.1, Building J11, D-30625 Hannover, Germany 5Institute of Genetics and Molecular Medicine (IGMM), University of Edinburgh, Crewe Road, Edinburgh EH4 2XU, United Kingdom 6 The Milner Centre for Evolution, Department of Biology and Biochemistry, University of Bath, Bath, Somerset, UK, BA2 7AY. 7 Institute of Medical Systems Biology, Center for Molecular Neurobiology (ZMNH), University Medical Center Hamburg-Eppendorf (UKE), 20246, Hamburg, Germany. *Corresponding authors *Zsuzsanna Izsvák Max Delbrück Center for Molecular Medicine Robert Rössle Strasse 10, 13092 Berlin, Germany Telefon: +49 030-9406-3510 Fax: +49 030-9406-2547 email: [email protected] http://www.mdcberlin.de/en/research/research_teams/mobile_dna/index.html and *Laurence D. -

UNIVERSITY of CALIFORNIA RIVERSIDE Investigations Into The
UNIVERSITY OF CALIFORNIA RIVERSIDE Investigations into the Role of TAF1-mediated Phosphorylation in Gene Regulation A Dissertation submitted in partial satisfaction of the requirements for the degree of Doctor of Philosophy in Cell, Molecular and Developmental Biology by Brian James Gadd December 2012 Dissertation Committee: Dr. Xuan Liu, Chairperson Dr. Frank Sauer Dr. Frances M. Sladek Copyright by Brian James Gadd 2012 The Dissertation of Brian James Gadd is approved Committee Chairperson University of California, Riverside Acknowledgments I am thankful to Dr. Liu for her patience and support over the last eight years. I am deeply indebted to my committee members, Dr. Frank Sauer and Dr. Frances Sladek for the insightful comments on my research and this dissertation. Thanks goes out to CMDB, especially Dr. Bachant, Dr. Springer and Kathy Redd for their support. Thanks to all the members of the Liu lab both past and present. A very special thanks to the members of the Sauer lab, including Silvia, Stephane, David, Matt, Stephen, Ninuo, Toby, Josh, Alice, Alex and Flora. You have made all the years here fly by and made them so enjoyable. From the Sladek lab I want to thank Eugene, John, Linh and Karthi. Special thanks go out to all the friends I’ve made over the years here. Chris, Amber, Stephane and David, thank you so much for feeding me, encouraging me and keeping me sane. Thanks to the brothers for all your encouragement and prayers. To any I haven’t mentioned by name, I promise I haven’t forgotten all you’ve done for me during my graduate years. -

PRODUCT SPECIFICATION Prest Antigen TEX9
PrEST Antigen TEX9 Product Datasheet PrEST Antigen PRODUCT SPECIFICATION Product Name PrEST Antigen TEX9 Product Number APrEST81497 Gene Description testis expressed 9 Corresponding Anti-TEX9 (HPA039415) Antibodies Description Recombinant protein fragment of Human TEX9 Amino Acid Sequence Recombinant Protein Epitope Signature Tag (PrEST) antigen sequence: QEELDNVVCECNKKEDEIQNLKSQVKNFEEDFMRQQRTINMQQSQVEKYK TLFEEANKKYDGLQQQLSSVERELENKRRLQKQAASSQSAT Fusion Tag N-terminal His6ABP (ABP = Albumin Binding Protein derived from Streptococcal Protein G) Expression Host E. coli Purification IMAC purification Predicted MW 28 kDa including tags Usage Suitable as control in WB and preadsorption assays using indicated corresponding antibodies. Purity >80% by SDS-PAGE and Coomassie blue staining Buffer PBS and 1M Urea, pH 7.4. Unit Size 100 µl Concentration Lot dependent Storage Upon delivery store at -20°C. Avoid repeated freeze/thaw cycles. Notes Gently mix before use. Optimal concentrations and conditions for each application should be determined by the user. Product of Sweden. For research use only. Not intended for pharmaceutical development, diagnostic, therapeutic or any in vivo use. No products from Atlas Antibodies may be resold, modified for resale or used to manufacture commercial products without prior written approval from Atlas Antibodies AB. Warranty: The products supplied by Atlas Antibodies are warranted to meet stated product specifications and to conform to label descriptions when used and stored properly. Unless otherwise stated, this warranty is limited to one year from date of sales for products used, handled and stored according to Atlas Antibodies AB's instructions. Atlas Antibodies AB's sole liability is limited to replacement of the product or refund of the purchase price. All products are supplied for research use only. -

TEX9 and Eif3b Functionally Synergize to Promote the Progression Of
Xu et al. BMC Cancer (2019) 19:875 https://doi.org/10.1186/s12885-019-6071-9 RESEARCHARTICLE Open Access TEX9 and eIF3b functionally synergize to promote the progression of esophageal squamous cell carcinoma Fengkai Xu1†, Shu Zhang2†, Zhonghe Liu1†, Jie Gu1, Yin Li1, Lin Wang1, Wei Mao1, Qiaoliang Zhu1, Huankai Shou1, Di Ge1 and Chunlai Lu1* Abstract Background: Esophageal squamous cell carcinoma (ESCC) is one of the most frequent malignant digestive tumors around the world. We previously demonstrated that eIF3b could promote the progression of ESCC. The exact mechanisms underlying these effects remained unknown. Methods: Quantitative proteomics was applied to detect the potential targets of Eukaryotic translation initiation factor 3 subunit b (eIF3b). RT-qPCR and Western blot were performed to detect the expression of targeted gene and pathway related genes. RNA-immunoprecipitation was applied to verify the binding of eIF3b with targeted gene. Moreover, CCK-8 assay, colony-formation assay, transwell assay, flow cytometry for cell apoptosis and tumor xenograft assay were performed to analyze the regulation of the targeted gene on the bio-function of ESCC cells. Results: Quantitative proteomics data showed that Testis-expressed protein 9 (TEX9) expression was positively associated with eIF3b expression. RT-qPCR and Western blot results confirmed the quantitative proteomics data and demonstrated that TEX9 expression was positively correlated with TNM stage in ESCC. Furtherly, RNA- immunoprecipitation confirmed that eIF3b binding to TEX9 mRNA. The bio-function related assay demonstrated that TEX9 and eIF3b functionally synergized to promote the proliferation and migration, and inhibited the apoptosis of ESCC cells. In the analysis of mechanism, we revealed that TEX9 and eIF3b promoted the progression of ESCC through the activation of AKT signaling pathway. -

Termination of RNA Polymerase II Transcription by the 5’-3’ Exonuclease Xrn2
TERMINATION OF RNA POLYMERASE II TRANSCRIPTION BY THE 5’-3’ EXONUCLEASE XRN2 by MICHAEL ANDRES CORTAZAR OSORIO B.S., Universidad del Valle – Colombia, 2011 A thesis submitted to the Faculty of the Graduate School of the University of Colorado in partial fulfillment of the requirements for the degree of Doctor of Philosophy Molecular Biology Program 2018 This thesis for the Doctor of Philosophy degree by Michael Andrés Cortázar Osorio has been approved for the Molecular Biology Program by Mair Churchill, Chair Richard Davis Jay Hesselberth Thomas Blumenthal James Goodrich David Bentley, Advisor Date: Aug 17, 2018 ii Cortázar Osorio, Michael Andrés (Ph.D., Molecular Biology) Termination of RNA polymerase II transcription by the 5’-3’ exonuclease Xrn2 Thesis directed by Professor David L. Bentley ABSTRACT Termination of transcription occurs when RNA polymerase (pol) II dissociates from the DNA template and releases a newly-made mRNA molecule. Interestingly, an active debate fueled by conflicting reports over the last three decades is still open on which of the two main models of termination of RNA polymerase II transcription does in fact operate at 3’ ends of genes. The torpedo model indicates that the 5’-3’ exonuclease Xrn2 targets the nascent transcript for degradation after cleavage at the polyA site and chases pol II for termination. In contrast, the allosteric model asserts that transcription through the polyA signal induces a conformational change of the elongation complex and converts it into a termination-competent complex. In this thesis, I propose a unified allosteric-torpedo mechanism. Consistent with a polyA site-dependent conformational change of the elongation complex, I found that pol II transitions at the polyA site into a mode of slow transcription elongation that is accompanied by loss of Spt5 phosphorylation in the elongation complex. -

Genomics of Inherited Bone Marrow Failure and Myelodysplasia Michael
Genomics of inherited bone marrow failure and myelodysplasia Michael Yu Zhang A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy University of Washington 2015 Reading Committee: Mary-Claire King, Chair Akiko Shimamura Marshall Horwitz Program Authorized to Offer Degree: Molecular and Cellular Biology 1 ©Copyright 2015 Michael Yu Zhang 2 University of Washington ABSTRACT Genomics of inherited bone marrow failure and myelodysplasia Michael Yu Zhang Chair of the Supervisory Committee: Professor Mary-Claire King Department of Medicine (Medical Genetics) and Genome Sciences Bone marrow failure and myelodysplastic syndromes (BMF/MDS) are disorders of impaired blood cell production with increased leukemia risk. BMF/MDS may be acquired or inherited, a distinction critical for treatment selection. Currently, diagnosis of these inherited syndromes is based on clinical history, family history, and laboratory studies, which directs the ordering of genetic tests on a gene-by-gene basis. However, despite extensive clinical workup and serial genetic testing, many cases remain unexplained. We sought to define the genetic etiology and pathophysiology of unclassified bone marrow failure and myelodysplastic syndromes. First, to determine the extent to which patients remained undiagnosed due to atypical or cryptic presentations of known inherited BMF/MDS, we developed a massively-parallel, next- generation DNA sequencing assay to simultaneously screen for mutations in 85 BMF/MDS genes. Querying 71 pediatric and adult patients with unclassified BMF/MDS using this assay revealed 8 (11%) patients with constitutional, pathogenic mutations in GATA2 , RUNX1 , DKC1 , or LIG4 . All eight patients lacked classic features or laboratory findings for their syndromes. -

Molecular Signatures of Membrane Protein Complexes Underlying Muscular Dystrophy*□S
crossmark Research Author’s Choice © 2016 by The American Society for Biochemistry and Molecular Biology, Inc. This paper is available on line at http://www.mcponline.org Molecular Signatures of Membrane Protein Complexes Underlying Muscular Dystrophy*□S Rolf Turk‡§¶ʈ**, Jordy J. Hsiao¶, Melinda M. Smits¶, Brandon H. Ng¶, Tyler C. Pospisil‡§¶ʈ**, Kayla S. Jones‡§¶ʈ**, Kevin P. Campbell‡§¶ʈ**, and Michael E. Wright¶‡‡ Mutations in genes encoding components of the sar- The muscular dystrophies are hereditary diseases charac- colemmal dystrophin-glycoprotein complex (DGC) are re- terized primarily by the progressive degeneration and weak- sponsible for a large number of muscular dystrophies. As ness of skeletal muscle. Most are caused by deficiencies in such, molecular dissection of the DGC is expected to both proteins associated with the cell membrane (i.e. the sarco- reveal pathological mechanisms, and provides a biologi- lemma in skeletal muscle), and typical features include insta- cal framework for validating new DGC components. Es- bility of the sarcolemma and consequent death of the myofi- tablishment of the molecular composition of plasma- ber (1). membrane protein complexes has been hampered by a One class of muscular dystrophies is caused by mutations lack of suitable biochemical approaches. Here we present in genes that encode components of the sarcolemmal dys- an analytical workflow based upon the principles of pro- tein correlation profiling that has enabled us to model the trophin-glycoprotein complex (DGC). In differentiated skeletal molecular composition of the DGC in mouse skeletal mus- muscle, this structure links the extracellular matrix to the cle. We also report our analysis of protein complexes in intracellular cytoskeleton. -

TEX9 (D-2): Sc-398987
SANTA CRUZ BIOTECHNOLOGY, INC. TEX9 (D-2): sc-398987 BACKGROUND APPLICATIONS TEX9 (testis-expressed sequence 9 protein) is a 391 amino acid protein. The TEX9 (D-2) is recommended for detection of TEX9 of mouse, rat and human gene that encodes TEX9 contains roughly 80,448 bases and maps to human origin by Western Blotting (starting dilution 1:100, dilution range 1:100- chromosome 15q21.3. Housing approximately 106 million base pairs and 1:1000), immunoprecipitation [1-2 µg per 100-500 µg of total protein (1 ml encoding more than 700 genes, chromosome 15 makes up about 3% of the of cell lysate)], immunofluorescence (starting dilution 1:50, dilution range human genome. Angelman and Prader-Willi syndromes are associated with 1:50-1:500) and solid phase ELISA (starting dilution 1:30, dilution range loss of function or deletion of genes in the 15q11-q13 region. In the case of 1:30-1:3000). Angelman syndrome, this loss is due to inactivity of the maternal 15q11-q13 Suitable for use as control antibody for TEX9 siRNA (h): sc-90152, TEX9 encoded UBE3A gene in the brain by either chromosomal deletion or mutation. siRNA (m): sc-154227, TEX9 shRNA Plasmid (h): sc-90152-SH, TEX9 shRNA In cases of Prader-Willi syndrome, there is a partial or complete deletion of Plasmid (m): sc-154227-SH, TEX9 shRNA (h) Lentiviral Particles: sc-90152-V this region from the paternal copy of chromosome 15. Tay-Sachs disease is a and TEX9 shRNA (m) Lentiviral Particles: sc-154227-V. lethal disorder associated with mutations of the HEXA gene, which is encoded by chromosome 15. -

Full-Text.Pdf
Systematic Evaluation of Genes and Genetic Variants Associated with Type 1 Diabetes Susceptibility This information is current as Ramesh Ram, Munish Mehta, Quang T. Nguyen, Irma of September 23, 2021. Larma, Bernhard O. Boehm, Flemming Pociot, Patrick Concannon and Grant Morahan J Immunol 2016; 196:3043-3053; Prepublished online 24 February 2016; doi: 10.4049/jimmunol.1502056 Downloaded from http://www.jimmunol.org/content/196/7/3043 Supplementary http://www.jimmunol.org/content/suppl/2016/02/19/jimmunol.150205 Material 6.DCSupplemental http://www.jimmunol.org/ References This article cites 44 articles, 5 of which you can access for free at: http://www.jimmunol.org/content/196/7/3043.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision by guest on September 23, 2021 • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Systematic Evaluation of Genes and Genetic Variants Associated with Type 1 Diabetes Susceptibility Ramesh Ram,*,† Munish Mehta,*,† Quang T.