Final Report Umpqua Chub Population Estimate USFWS Grant # E-2-54
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
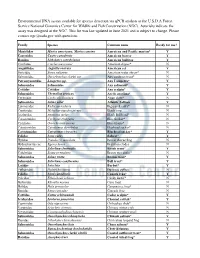
Edna Assay Development
Environmental DNA assays available for species detection via qPCR analysis at the U.S.D.A Forest Service National Genomics Center for Wildlife and Fish Conservation (NGC). Asterisks indicate the assay was designed at the NGC. This list was last updated in June 2021 and is subject to change. Please contact [email protected] with questions. Family Species Common name Ready for use? Mustelidae Martes americana, Martes caurina American and Pacific marten* Y Castoridae Castor canadensis American beaver Y Ranidae Lithobates catesbeianus American bullfrog Y Cinclidae Cinclus mexicanus American dipper* N Anguillidae Anguilla rostrata American eel Y Soricidae Sorex palustris American water shrew* N Salmonidae Oncorhynchus clarkii ssp Any cutthroat trout* N Petromyzontidae Lampetra spp. Any Lampetra* Y Salmonidae Salmonidae Any salmonid* Y Cottidae Cottidae Any sculpin* Y Salmonidae Thymallus arcticus Arctic grayling* Y Cyrenidae Corbicula fluminea Asian clam* N Salmonidae Salmo salar Atlantic Salmon Y Lymnaeidae Radix auricularia Big-eared radix* N Cyprinidae Mylopharyngodon piceus Black carp N Ictaluridae Ameiurus melas Black Bullhead* N Catostomidae Cycleptus elongatus Blue Sucker* N Cichlidae Oreochromis aureus Blue tilapia* N Catostomidae Catostomus discobolus Bluehead sucker* N Catostomidae Catostomus virescens Bluehead sucker* Y Felidae Lynx rufus Bobcat* Y Hylidae Pseudocris maculata Boreal chorus frog N Hydrocharitaceae Egeria densa Brazilian elodea N Salmonidae Salvelinus fontinalis Brook trout* Y Colubridae Boiga irregularis Brown tree snake* -

Umpqua Bibliography by Groups I. Possible
Umpqua Bibliography by Groups Introduction This bibliography is about the Umpqua River Basin, particularly its hydrology and its fisheries. Some films, photographs and archival materials are included along with books, articles, maps and web pages. The bibliography was built using local, regional and national library catalogs, major online databases, theses and dissertations, watershed assessments and Federal endangered species reports. Its historical coverage begins with Joel Palmer’s 1847 Journal of Travels over the Rocky Mountains and ends in early 2008. I have attempted to cover major physical features of the Umpqua Basin, as well as important aquatic species and critical issues facing the area. Trying to build a comprehensive collection of references on any subject in the age of information is, to use an image Joel Palmer would understand, like trying to catch a greased pig. It is ever squirming out of your grasp. This collection of references is necessarily incomplete, but I’d be grateful to learn of any major omissions. --Susan Gilmont Updated September, 2008 I. Possible Candidates for Digitization: Grey Literature, Important Or Uncataloged Reports on the Umpqua River (Sorted by Date) Roth, A. R. A survey of the waters of the South Umpqua Ranger District : Umpqua National Forest . Portland, Or.: USDA Forest Service, Pacific Northwest Region; 1937. Call Number: OSU Libraries: Valley SH35.O7 U65 Notes: Baseline temperature data for the Umpqua Oregon State Game Commission. Lower Umpqua River Study : Summary : Annual Report. 1949. Call Number: Available from NMFS Environmental and Technical Svcs. Division, 525 NE Oregon St. Portland, OR 97232 (? dated reference). State Library or StreamNet should have this. -

Oregonichthys Kalawatseti) in the North Umpqua After Almost a Century
REOCCURRENCE OF THE UMPQUA CHUB (OREGONICHTHYS KALAWATSETI) IN THE NORTH UMPQUA AFTER ALMOST A CENTURY Authors: Penaluna, Brooke E, and Ellenburg, Loretta Source: Northwestern Naturalist, 100(3) : 208-210 Published By: Society for Northwestern Vertebrate Biology URL: https://doi.org/10.1898/1051-1733-100.3.208 BioOne Complete (complete.BioOne.org) is a full-text database of 200 subscribed and open-access titles in the biological, ecological, and environmental sciences published by nonprofit societies, associations, museums, institutions, and presses. Your use of this PDF, the BioOne Complete website, and all posted and associated content indicates your acceptance of BioOne’s Terms of Use, available at www.bioone.org/terms-of-use. Usage of BioOne Complete content is strictly limited to personal, educational, and non - commercial use. Commercial inquiries or rights and permissions requests should be directed to the individual publisher as copyright holder. BioOne sees sustainable scholarly publishing as an inherently collaborative enterprise connecting authors, nonprofit publishers, academic institutions, research libraries, and research funders in the common goal of maximizing access to critical research. Downloaded From: https://bioone.org/journals/Northwestern-Naturalist on 27 May 2020 Terms of Use: https://bioone.org/terms-of-use Access provided by United States Department of Agriculture National Agricultural Library (NAL) GENERAL NOTES NORTHWESTERN NATURALIST 100:208–210 WINTER 2019 REOCCURRENCE OF THE UMPQUA CHUB (OREGONICHTHYS KALAWATSETI) IN THE NORTH UMPQUA AFTER ALMOST A CENTURY BROOKE EPENALUNA AND LORETTA ELLENBURG ABSTRACT—We document the 1st record in 93 y of Subsequent surveys to the same sites throughout Umpqua Chub (Oregonichthys kalawatseti) in the North the basin in 1998 (Simon and Markle 1999), and Umpqua River. -

Molecular Systematics of Western North American Cyprinids (Cypriniformes: Cyprinidae)
Zootaxa 3586: 281–303 (2012) ISSN 1175-5326 (print edition) www.mapress.com/zootaxa/ ZOOTAXA Copyright © 2012 · Magnolia Press Article ISSN 1175-5334 (online edition) urn:lsid:zoobank.org:pub:0EFA9728-D4BB-467E-A0E0-0DA89E7E30AD Molecular systematics of western North American cyprinids (Cypriniformes: Cyprinidae) SUSANA SCHÖNHUTH 1, DENNIS K. SHIOZAWA 2, THOMAS E. DOWLING 3 & RICHARD L. MAYDEN 1 1 Department of Biology, Saint Louis University, 3507 Laclede Avenue, St. Louis, MO 63103, USA. E-mail S.S: [email protected] ; E-mail RLM: [email protected] 2 Department of Biology and Curator of Fishes, Monte L. Bean Life Science Museum, Brigham Young University, Provo, UT 84602, USA. E-mail: [email protected] 3 School of Life Sciences, Arizona State University, Tempe, AZ 85287-4501, USA. E-mail: [email protected] Abstract The phylogenetic or evolutionary relationships of species of Cypriniformes, as well as their classification, is in a era of flux. For the first time ever, the Order, and constituent Families are being examined for relationships within a phylogenetic context. Relevant findings as to sister-group relationships are largely being inferred from analyses of both mitochondrial and nuclear DNA sequences. Like the vast majority of Cypriniformes, due to an overall lack of any phylogenetic investigation of these fishes since Hennig’s transformation of the discipline, changes in hypotheses of relationships and a natural classification of the species should not be of surprise to anyone. Basically, for most taxa no properly supported phylogenetic hypothesis has ever been done; and this includes relationships with reasonable taxon and character sampling of even families and subfamilies. -

Fishtraits: a Database on Ecological and Life-History Traits of Freshwater
FishTraits database Traits References Allen, D. M., W. S. Johnson, and V. Ogburn-Matthews. 1995. Trophic relationships and seasonal utilization of saltmarsh creeks by zooplanktivorous fishes. Environmental Biology of Fishes 42(1)37-50. [multiple species] Anderson, K. A., P. M. Rosenblum, and B. G. Whiteside. 1998. Controlled spawning of Longnose darters. The Progressive Fish-Culturist 60:137-145. [678] Barber, W. E., D. C. Williams, and W. L. Minckley. 1970. Biology of the Gila Spikedace, Meda fulgida, in Arizona. Copeia 1970(1):9-18. [485] Becker, G. C. 1983. Fishes of Wisconsin. University of Wisconsin Press, Madison, WI. Belk, M. C., J. B. Johnson, K. W. Wilson, M. E. Smith, and D. D. Houston. 2005. Variation in intrinsic individual growth rate among populations of leatherside chub (Snyderichthys copei Jordan & Gilbert): adaptation to temperature or length of growing season? Ecology of Freshwater Fish 14:177-184. [349] Bonner, T. H., J. M. Watson, and C. S. Williams. 2006. Threatened fishes of the world: Cyprinella proserpina Girard, 1857 (Cyprinidae). Environmental Biology of Fishes. In Press. [133] Bonnevier, K., K. Lindstrom, and C. St. Mary. 2003. Parental care and mate attraction in the Florida flagfish, Jordanella floridae. Behavorial Ecology and Sociobiology 53:358-363. [410] Bortone, S. A. 1989. Notropis melanostomus, a new speices of Cyprinid fish from the Blackwater-Yellow River drainage of northwest Florida. Copeia 1989(3):737-741. [575] Boschung, H.T., and R. L. Mayden. 2004. Fishes of Alabama. Smithsonian Books, Washington. [multiple species] 1 FishTraits database Breder, C. M., and D. E. Rosen. 1966. Modes of reproduction in fishes. -

Thesis-1967D-R418d.Pdf (6.356Mb)
DESCRIPTIONS AND ANAllSES OF 'l'HE CEPHAUC LATERAL-LINE SYSTEMS OF 'l'BE CYPRINID GDmS HYBOPSIS ·~ , HARLEY f!AYNE ~O Bachelor of Science Oklah~ Stat• Univer~it7 Stillli!B.ter, Oklahoma 1961 !'aster or Science Oklahoma State University _Stillwater, Oklahoma l.963 Submitted to the faculty of the GradWtte College of th~ Oklahoma State tfniver~ity in partial tul:f'illmm.t of th~ requir~nts for ths degrc,e of DOCTOR OF PHILOSOPHY Juq; 1967 DESCRIPTIONS AND ANALYSES OF THE CEPHALIC LATERAL-LINE SYSTEMS OF THE CYPRJNID GENUS HYBOPSIS Thesis Approved: ~.o.m~ n nDean iJ&Ve,of the Graduate --- College 11 OKLAHOMA STATE UNIVERSITY LIBRARY ·1 . ~ ·1 JAN 16 191W } PRE.FACE Use of the cephalic lateral-line system as an ind.ex to habitats knowno and ecological requirements of fishes ha.111. long been. This lateral-line study o:t.Bzbopsis was aonduate~ to; (1) correlate lateral- .. line structure with species ecologies; (2) predict probable ecologies of little-known species 0£ Bybopsis;, (3) investigate the neuromast- bone :relationships in Hzbopsis; and (4) erect tentative phylogenies for some subgenera of ljtybopsiso I am indebted ta Dr. Rudolph J. Mill.er, 'filY' major advisor, tor his asaietanoe and encouragement during the courae or this •tu:iy and his critical examination of this manuscript. Dre. George Ao Moli>re, L. Herbert Bruneau, Br.van Po Gla11, am Roy w. Jones aerved on ?113' advisory committee and also evaluated the manuscript. Sincere thanks i1 pereon aJ4r extemad to Dr. George A. Moore 11 because it waa his initial in fluence years ago which aroused~ interests in fish morphology . -

Taxonomic Analyses of the Oregon Chub, Oregonichthys Crameri
AN ABSTRACT OF THE THESIS OF Debra T. Bills for the degree of Master of Science in Fisheries and Wildlife presented on June 1.1989. Title: Taxonomic Analyses of the Oregon Chub ( Oregonichthys crameri) Redacted for Privacy Abstract approved: tkWuglas F. Markle The Oregon chub. Oregonichthys crameri. one of Oregon's endemic freshwater fishes.is restricted to the Willamette and Umpqua rivers and their tributaries. Specimens from the two rivers have traditionally been considered the same species. The goal of this study was to re-examine that assumption and determine if the Willamette Oregon chub and the Umpqua Oregon chub represent different taxa. The Oregon chub were examined through three sets of data: meristic. morphometric. and electrophoretic. Univariate (General Linear Model) and multivariate (Principal Components Analysis and Discriminant Function Analysis) analyses were performed. Univariate analyses resulted in significant differences between the Willamette and the Umpqua in the meristic characters: precaudal vertebrae. caudal vertebrae. the difference between precaudal and caudal vertebrae. pelvic fin rays, rudimentary dorsal rays, total dorsal rays, upper procurrent caudal rays, total caudal rays, maxillary barbels, and cephalic lateral line counts; in the morphometric characters: body width, head depth, eye length, and anal base length; and in the electrophoretic character, lactate dehydrogenase in muscle tissue. Discriminant function analysis based on meristic characters correctly classifies 90% of the Willamette and 65% of the Umpqua specimens. Discriminant function analysis based on morphometric characters correctly classifies over 89% of the Willamette and over 84% of the Umpqua specimens. Descriptions of the Willamette and Umpqua Oregon chub subspecies are given. Taxonomic Analyses of the Oregon Chub (Oregonichthys crameri) by Debra T. -

United States Department of the Interior
United States Department of the Interior FISH AND WILDLIFE SERVICE Oregon Fish and Wildlife Office 2600 SE 98th Avenue, Suite 100 Portland, Oregon 97266 Phone: (503) 231-6179 FAX: (503) 231-6195 Reply To: 8330.F0047(09) File Name: CREP BO 2009_final.doc TS Number: 09-314 TAILS: 13420-2009-F-0047 Doc Type: Final Don Howard, Acting State Executive Director U.S. Department of Agriculture Farm Service Agency, Oregon State Office 7620 SW Mohawk St. Tualatin, OR 97062-8121 Dear Mr. Howard, This letter transmits the U.S. Fish and Wildlife Service’s (Service) Biological and Conference Opinion (BO) and includes our written concurrence based on our review of the proposed Oregon Conservation Reserve Enhancement Program (CREP) to be administered by the Farm Service Agency (FSA) throughout the State of Oregon, and its effects on Federally-listed species in accordance with section 7 of the Endangered Species Act (Act) of 1973, as amended (16 U.S.C. 1531 et seq.). Your November 24, 2008 request for informal and formal consultation with the Service, and associated Program Biological Assessment for the Oregon Conservation Reserve Enhancement Program (BA), were received on November 24, 2008. We received your letter providing a 90-day extension on March 26, 2009 based on the scope and complexity of the program and the related species that are covered, which we appreciated. This Concurrence and BO covers a period of approximately 10 years, from the date of issuance through December 31, 2019. The BA also includes species that fall within the jurisdiction of the National Oceanic and Atmospheric Administration’s Fisheries Service (NOAA Fisheries Service). -

Phylogeography of Rhinichthys Cataractae (Teleostei
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Texas A&M University PHYLOGEOGRAPHY OF RHINICHTHYS CATARACTAE (TELEOSTEI: CYPRINIDAE): PRE-GLACIAL COLONIZATION ACROSS THE CONTINENTAL DIVIDE AND PLEISTOCENE DIVERSIFICATION WITHIN THE RIO GRANDE DRAINAGE A Thesis by DAE-MIN KIM Submitted to the Office of Graduate and Professional Studies of Texas A&M University in partial fulfillment of the requirements for the degree of MASTER OF SCIENCE Chair of Committee, Kevin W. Conway Committee Members, Gary P. Garrett John R. Gold Gary Voelker Head of Department, Michael Masser December 2013 Major Subject: Wildlife and Fisheries Sciences Copyright 2013 Dae-Min Kim ABSTRACT The longnose dace, Rhinichthys cataractae, is a primary freshwater fish inhibiting riffle habitats in small headwater rivers and streams across the North American continent, including drainages east and west of the Continental Divide. Phylogenetic analyses of 1140 base pairs (bp) of the mitochondrially encoded cytochrome b gene and 2298-2346 bp of the nuclear-encoded genes S7 and RAG1 were obtained from 87 individuals of R. cataractae (collected throughout its range) and from several close relatives recovered a monophyletic R. cataractae species-group that contained R. evermanni, R. sp ―Millicoma dace‖ and R. cataractae; a monophyletic R. cataractae was not recovered. Within the R. cataractae species-group, two well-supported clades were identified, including a western clade (containing R. evermanni, R. sp. ―Millicoma dace‖ and individuals of R. cataractae from Pacific slope drainages) and an eastern clade (containing individuals of R. cataractae from Arctic, Atlantic, and Gulf slope drainages). Within the eastern clade of R. -

Conservation Status of Imperiled North American Freshwater And
FEATURE: ENDANGERED SPECIES Conservation Status of Imperiled North American Freshwater and Diadromous Fishes ABSTRACT: This is the third compilation of imperiled (i.e., endangered, threatened, vulnerable) plus extinct freshwater and diadromous fishes of North America prepared by the American Fisheries Society’s Endangered Species Committee. Since the last revision in 1989, imperilment of inland fishes has increased substantially. This list includes 700 extant taxa representing 133 genera and 36 families, a 92% increase over the 364 listed in 1989. The increase reflects the addition of distinct populations, previously non-imperiled fishes, and recently described or discovered taxa. Approximately 39% of described fish species of the continent are imperiled. There are 230 vulnerable, 190 threatened, and 280 endangered extant taxa, and 61 taxa presumed extinct or extirpated from nature. Of those that were imperiled in 1989, most (89%) are the same or worse in conservation status; only 6% have improved in status, and 5% were delisted for various reasons. Habitat degradation and nonindigenous species are the main threats to at-risk fishes, many of which are restricted to small ranges. Documenting the diversity and status of rare fishes is a critical step in identifying and implementing appropriate actions necessary for their protection and management. Howard L. Jelks, Frank McCormick, Stephen J. Walsh, Joseph S. Nelson, Noel M. Burkhead, Steven P. Platania, Salvador Contreras-Balderas, Brady A. Porter, Edmundo Díaz-Pardo, Claude B. Renaud, Dean A. Hendrickson, Juan Jacobo Schmitter-Soto, John Lyons, Eric B. Taylor, and Nicholas E. Mandrak, Melvin L. Warren, Jr. Jelks, Walsh, and Burkhead are research McCormick is a biologist with the biologists with the U.S. -

Smallmouth Bass (Micropterus Dolomieu) ERSS
Smallmouth Bass (Micropterus dolomieu) Ecological Risk Screening Summary U.S. Fish & Wildlife Service, February 2019 Web Version, 8/26/2019 Photo: Marcus Rosten/USFWS. Public domain – Government work. 1 Native Range and Status in the United States Native Range From Froese and Pauly (2019a): “North America: St. Lawrence-Great Lakes system, Hudson Bay and Mississippi River basins from southern Quebec in Canada to North Dakota and south to northern Alabama and eastern Oklahoma in the USA.” “Native species ranging from Manitoba to Nova Scotia [Canada]; introduced into other provinces [Coker et al. 2001].” Status in the United States From Froese and Pauly (2019a): “Widely transplanted throughout USA except in Atlantic and Gulf Slope drainages, where rare south of Virginia and east of central Texas [Page and Burr 1991].” “Imported into Hawaii in 1953 for sportfishing; became established in Nu'uanu Reservoir No.3 in 1956; presently well established in the Ho'omaluhia Reservoir, Kaukonahua Stream, Nu'uanu Stream, and Maunawili Stream on O'ahu and in various streams on the island of Kaua'I; law 1 prohibits release of this species in areas where they had not previously occurred [Yamamoto and Tagawa 2000].” According to Fuller et al. (2019), nonindigenous occurrences of Micropterus dolomieu have been reported in the following States, with range of years and hydrologic units in parentheses: Alabama (1971-2008; Black Warrior-Tombigbee, Cahaba, Coosa-Tallapoosa, Middle Coosa, Middle Tombigbee-Chickasaw, Mobile-Tombigbee, Sipsey Fork, Upper Tallapoosa) -

California's Freshwater Biodiversity
CALIFORNIA’S FRESHWATER BIODIVERSITY IN A CONTINENTAL CONTEXT Science for Conservation Technical Brief November 2009 Department of Conservation Science The Nature Conservancy, California Freshwater Biodiversity in California 2 California’s Freshwater Ecoregions in a Continental Context A Science for Conservation Technical Brief The Nature Conservancy, California The Freshwater Conservation Challenge Worldwide, freshwater species and habitats are, on average, more imperiled than their terrestrial or marine counterparts. In continental North America alone, 40% of freshwater fish are at risk of extinction or already extinct (Jelks et al. 2008). Despite concerns over the health of the world’s freshwater species and systems (Millennium Ecosystem Assessment 2005), there have been few attempts to systematically describe patterns of freshwater biodiversity on Earth. This is due in part to the lack of comprehensive, synthesized data on the distributions of freshwater species (Abell 2008). Without a robust biodiversity foundation, conservationists face challenges in setting freshwater protection priorities and agendas at the global, continental and regional scales. Freshwater Ecoregions of the World Project To fill this void, in 2008, World Wildlife Fund-US, The Nature Conservancy, and more than 130 scientists participated in the Freshwater Ecoregions of the World (FEOW) project. FEOW identified 426 freshwater ecoregions and provided information on freshwater biogeography and biodiversity; similar analyses exist for the terrestrial and marine realms. Until this effort, global biodiversity classification and planning efforts had been characterized using land-based parameters. FEOW is the first attempt to describe the world from a freshwater perspective. With this information, scientists and conservationists can more clearly compare freshwater biota and their conservation needs across large geographies.