Comparing Three Ecosystem Models of the Tasman and Golden Bays, New Zealand
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
World Bank Document
GLOBAL ENVIRONMENT 33977 FACILITY Public Disclosure Authorized Public Disclosure Authorized Quarterly Operational Report April 1995 Public Disclosure Authorized GEF Public Disclosure Authorized development,agencies, national institutions, (GEF) is a financial tions, bilateral T mechanismhe Global Environment that provides Facility grant and concessional funds non-governmental organizations (NGOs), private sector to developing countries for projects and activities that aim entities, and academic institutions. The GEF also comprises to protect the global environment. GEF resources are avail- a Small Grants Programme available for projects in the able for projects and other activities that address climate four focal areas that are put forward by grassroots groups change, loss of biological diversity, pollution of international and NGOs in developing countries. waters, and depletion of the ozone layer. Countries can The Quarterly Operational Report is designed to pro- obtain GEF funds if they are eligible to borrow from the vide a comprehensive review of, and a status report on, the World Bank (IBRD and/or IDA) or receive technical assis- GEE work program. A brief description of each of the GEE's tance grants from UNDP through a country program. projects organized alphabetically by region can be Responsibility for implementing GEF activities is found on pages 8-J8. Each description lists the name of the shared by the United Nations Development Programme UNDP, UNEP or World Bank Task Manager responsible for (UNDP), the United Nations Environment Programme the project. Inquiries about specific projects should be (UNEP) and the World Bank. UNDP is responsible for referred to the responsible Task Manager. Their telephone technical assistance activities, capacity building, and the and fax numbers can be found on pages 63 and 64. -

New Zealand Fishes a Field Guide to Common Species Caught by Bottom, Midwater, and Surface Fishing Cover Photos: Top – Kingfish (Seriola Lalandi), Malcolm Francis
New Zealand fishes A field guide to common species caught by bottom, midwater, and surface fishing Cover photos: Top – Kingfish (Seriola lalandi), Malcolm Francis. Top left – Snapper (Chrysophrys auratus), Malcolm Francis. Centre – Catch of hoki (Macruronus novaezelandiae), Neil Bagley (NIWA). Bottom left – Jack mackerel (Trachurus sp.), Malcolm Francis. Bottom – Orange roughy (Hoplostethus atlanticus), NIWA. New Zealand fishes A field guide to common species caught by bottom, midwater, and surface fishing New Zealand Aquatic Environment and Biodiversity Report No: 208 Prepared for Fisheries New Zealand by P. J. McMillan M. P. Francis G. D. James L. J. Paul P. Marriott E. J. Mackay B. A. Wood D. W. Stevens L. H. Griggs S. J. Baird C. D. Roberts‡ A. L. Stewart‡ C. D. Struthers‡ J. E. Robbins NIWA, Private Bag 14901, Wellington 6241 ‡ Museum of New Zealand Te Papa Tongarewa, PO Box 467, Wellington, 6011Wellington ISSN 1176-9440 (print) ISSN 1179-6480 (online) ISBN 978-1-98-859425-5 (print) ISBN 978-1-98-859426-2 (online) 2019 Disclaimer While every effort was made to ensure the information in this publication is accurate, Fisheries New Zealand does not accept any responsibility or liability for error of fact, omission, interpretation or opinion that may be present, nor for the consequences of any decisions based on this information. Requests for further copies should be directed to: Publications Logistics Officer Ministry for Primary Industries PO Box 2526 WELLINGTON 6140 Email: [email protected] Telephone: 0800 00 83 33 Facsimile: 04-894 0300 This publication is also available on the Ministry for Primary Industries website at http://www.mpi.govt.nz/news-and-resources/publications/ A higher resolution (larger) PDF of this guide is also available by application to: [email protected] Citation: McMillan, P.J.; Francis, M.P.; James, G.D.; Paul, L.J.; Marriott, P.; Mackay, E.; Wood, B.A.; Stevens, D.W.; Griggs, L.H.; Baird, S.J.; Roberts, C.D.; Stewart, A.L.; Struthers, C.D.; Robbins, J.E. -
Dictionary.Pdf
THE SEAFARER’S WORD A Maritime Dictionary A B C D E F G H I J K L M N O P Q R S T U V W X Y Z Ranger Hope © 2007- All rights reserved A ● ▬ A: Code flag; Diver below, keep well clear at slow speed. Aa.: Always afloat. Aaaa.: Always accessible - always afloat. A flag + three Code flags; Azimuth or bearing. numerals: Aback: When a wind hits the front of the sails forcing the vessel astern. Abaft: Toward the stern. Abaft of the beam: Bearings over the beam to the stern, the ships after sections. Abandon: To jettison cargo. Abandon ship: To forsake a vessel in favour of the life rafts, life boats. Abate: Diminish, stop. Able bodied seaman: Certificated and experienced seaman, called an AB. Abeam: On the side of the vessel, amidships or at right angles. Aboard: Within or on the vessel. About, go: To manoeuvre to the opposite sailing tack. Above board: Genuine. Able bodied seaman: Advanced deckhand ranked above ordinary seaman. Abreast: Alongside. Side by side Abrid: A plate reinforcing the top of a drilled hole that accepts a pintle. Abrolhos: A violent wind blowing off the South East Brazilian coast between May and August. A.B.S.: American Bureau of Shipping classification society. Able bodied seaman Absorption: The dissipation of energy in the medium through which the energy passes, which is one cause of radio wave attenuation. Abt.: About Abyss: A deep chasm. Abyssal, abysmal: The greatest depth of the ocean Abyssal gap: A narrow break in a sea floor rise or between two abyssal plains. -

British Family Names
cs 25o/ £22, Cornrll IBniwwitg |fta*g BOUGHT WITH THE INCOME FROM THE SAGE ENDOWMENT FUND THE GIFT OF Hcnrti W~ Sage 1891 A.+.xas.Q7- B^llll^_ DATE DUE ,•-? AUG 1 5 1944 !Hak 1 3 1^46 Dec? '47T Jan 5' 48 ft e Univeral, CS2501 .B23 " v Llb«"y Brit mii!Sm?nS,£& ori8'" and m 3 1924 olin 029 805 771 The original of this book is in the Cornell University Library. There are no known copyright restrictions in the United States on the use of the text. http://www.archive.org/details/cu31924029805771 BRITISH FAMILY NAMES. : BRITISH FAMILY NAMES ftbetr ©riain ano fIDeaning, Lists of Scandinavian, Frisian, Anglo-Saxon, and Norman Names. HENRY BARBER, M.D. (Clerk), "*• AUTHOR OF : ' FURNESS AND CARTMEL NOTES,' THE CISTERCIAN ABBEY OF MAULBRONN,' ( SOME QUEER NAMES,' ' THE SHRINE OF ST. BONIFACE AT FULDA,' 'POPULAR AMUSEMENTS IN GERMANY,' ETC. ' "What's in a name ? —Romeo and yuliet. ' I believe now, there is some secret power and virtue in a name.' Burton's Anatomy ofMelancholy. LONDON ELLIOT STOCK, 62, PATERNOSTER ROW, E.C. 1894. 4136 CONTENTS. Preface - vii Books Consulted - ix Introduction i British Surnames - 3 nicknames 7 clan or tribal names 8 place-names - ii official names 12 trade names 12 christian names 1 foreign names 1 foundling names 1 Lists of Ancient Patronymics : old norse personal names 1 frisian personal and family names 3 names of persons entered in domesday book as HOLDING LANDS temp. KING ED. CONFR. 37 names of tenants in chief in domesday book 5 names of under-tenants of lands at the time of the domesday survey 56 Norman Names 66 Alphabetical List of British Surnames 78 Appendix 233 PREFACE. -
2018 Final LOFF W/ Ref and Detailed Info
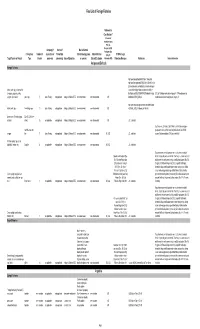
Final List of Foreign Fisheries Rationale for Classification ** (Presence of mortality or injury (P/A), Co- Occurrence (C/O), Company (if Source of Marine Mammal Analogous Gear Fishery/Gear Number of aquaculture or Product (for Interactions (by group Marine Mammal (A/G), No RFMO or Legal Target Species or Product Type Vessels processor) processing) Area of Operation or species) Bycatch Estimates Information (N/I)) Protection Measures References Detailed Information Antigua and Barbuda Exempt Fisheries http://www.fao.org/fi/oldsite/FCP/en/ATG/body.htm http://www.fao.org/docrep/006/y5402e/y5402e06.htm,ht tp://www.tradeboss.com/default.cgi/action/viewcompan lobster, rock, spiny, demersal fish ies/searchterm/spiny+lobster/searchtermcondition/1/ , (snappers, groupers, grunts, ftp://ftp.fao.org/fi/DOCUMENT/IPOAS/national/Antigua U.S. LoF Caribbean spiny lobster trap/ pot >197 None documented, surgeonfish), flounder pots, traps 74 Lewis Fishing not applicable Antigua & Barbuda EEZ none documented none documented A/G AndBarbuda/NPOA_IUU.pdf Caribbean mixed species trap/pot are category III http://www.nmfs.noaa.gov/pr/interactions/fisheries/tabl lobster, rock, spiny free diving, loops 19 Lewis Fishing not applicable Antigua & Barbuda EEZ none documented none documented A/G e2/Atlantic_GOM_Caribbean_shellfish.html Queen conch (Strombus gigas), Dive (SCUBA & free molluscs diving) 25 not applicable not applicable Antigua & Barbuda EEZ none documented none documented A/G U.S. trade data Southeastern U.S. Atlantic, Gulf of Mexico, and Caribbean snapper- handline, hook and grouper and other reef fish bottom longline/hook-and-line/ >5,000 snapper line 71 Lewis Fishing not applicable Antigua & Barbuda EEZ none documented none documented N/I, A/G U.S. -

Benthic Habitat Classes and Trawl Fishing Disturbance in New Zealand Waters Shallower Than 250 M
Benthic habitat classes and trawl fishing disturbance in New Zealand waters shallower than 250 m New Zealand Aquatic Environment and Biodiversity Report No.144 S.J. Baird, J. Hewitt, B.A. Wood ISSN 1179-6480 (online) ISBN 978-0-477-10532-3 (online) January 2015 Requests for further copies should be directed to: Publications Logistics Officer Ministry for Primary Industries PO Box 2526 WELLINGTON 6140 Email: [email protected] Telephone: 0800 00 83 33 Facsimile: 04-894 0300 This publication is also available on the Ministry for Primary Industries websites at: http://www.mpi.govt.nz/news-resources/publications.aspx http://fs.fish.govt.nz go to Document library/Research reports © Crown Copyright - Ministry for Primary Industries Contents EXECUTIVE SUMMARY 1 1. INTRODUCTION 3 The study area 3 2. COASTAL BENTHIC HABITAT CLASSES 4 2.1 Introduction 4 2.2 Habitat class definitions 6 2.3 Sensitivity of the habitat to fishing disturbance 10 3. SPATIAL PATTERN OF BOTTOM-CONTACTING TRAWL FISHING ACTIVITY 11 3.1 Bottom-contact trawl data 12 3.2 Spatial distribution of trawl data 21 3.3 Trawl footprint within the study area 26 3.4 Overlap of five-year trawl footprint on habitats within 250 m 32 3.5 GIS output from the overlay of the trawl footprint and habitat classes 37 4. SUMMARY OF NON-TRAWL BOTTOM-CONTACT FISHING METHODS IN THE STUDY AREA 38 5. DISCUSSION 39 6. ACKNOWLEDGMENTS 41 7. REFERENCES 42 APPENDIX 1: AREAS CLOSED TO FISHING WITHIN THE STUDY AREA 46 APPENDIX 2: MAPS SHOWING THE DISTRIBUTION OF THE DATA INPUTS FOR THE BENTHIC HABITAT DESCRIPTORS 49 APPENDIX 3: SENSITIVITY TO FISHING DISTURBANCE 53 APPENDIX 4: TRAWL FISHING DATA 102 APPENDIX 5: CELL-BASED TRAWL SUMMARIES 129 APPENDIX 6: TRAWL FOOTPRINT SUMMARY 151 APPENDIX 7: TRAWL FOOTPRINT – HABITAT OVERLAY 162 APPENDIX 8: SUMMARY OF DREDGE OYSTER AND SCALLOP EFFORT DATA WITHIN 250 M, 1 OCTOBER 2007–30 SEPTEMBER 2012 165 APPENDIX 9: SUMMARY OF DANISH SEINE EFFORT 181 EXECUTIVE SUMMARY Baird, S.J.; Hewitt, J.E.; Wood, B.A. -

Intrinsic Vulnerability in the Global Fish Catch
The following appendix accompanies the article Intrinsic vulnerability in the global fish catch William W. L. Cheung1,*, Reg Watson1, Telmo Morato1,2, Tony J. Pitcher1, Daniel Pauly1 1Fisheries Centre, The University of British Columbia, Aquatic Ecosystems Research Laboratory (AERL), 2202 Main Mall, Vancouver, British Columbia V6T 1Z4, Canada 2Departamento de Oceanografia e Pescas, Universidade dos Açores, 9901-862 Horta, Portugal *Email: [email protected] Marine Ecology Progress Series 333:1–12 (2007) Appendix 1. Intrinsic vulnerability index of fish taxa represented in the global catch, based on the Sea Around Us database (www.seaaroundus.org) Taxonomic Intrinsic level Taxon Common name vulnerability Family Pristidae Sawfishes 88 Squatinidae Angel sharks 80 Anarhichadidae Wolffishes 78 Carcharhinidae Requiem sharks 77 Sphyrnidae Hammerhead, bonnethead, scoophead shark 77 Macrouridae Grenadiers or rattails 75 Rajidae Skates 72 Alepocephalidae Slickheads 71 Lophiidae Goosefishes 70 Torpedinidae Electric rays 68 Belonidae Needlefishes 67 Emmelichthyidae Rovers 66 Nototheniidae Cod icefishes 65 Ophidiidae Cusk-eels 65 Trachichthyidae Slimeheads 64 Channichthyidae Crocodile icefishes 63 Myliobatidae Eagle and manta rays 63 Squalidae Dogfish sharks 62 Congridae Conger and garden eels 60 Serranidae Sea basses: groupers and fairy basslets 60 Exocoetidae Flyingfishes 59 Malacanthidae Tilefishes 58 Scorpaenidae Scorpionfishes or rockfishes 58 Polynemidae Threadfins 56 Triakidae Houndsharks 56 Istiophoridae Billfishes 55 Petromyzontidae -

Determining the Diet of New Zealand King Shag Using DNA Metabarcoding
Determining the diet of New Zealand king shag using DNA metabarcoding New Zealand King Shag, (Leucocarbo carunculatus) on Blumine Island, Marlborough Sounds, New Zealand in 2016 (Wikipedia commons). Aimee van der Reis & Andrew Jeffs Report Prepared For: Department of Conservation, Conservation Services Programme, Project BCBC2019-05. DOC MarineDRAFT Science Advisors Graeme Taylor and Dr Karen Middlemiss. November 2020 Reports from Auckland UniServices Limited should only be used for the purposes for which they were commissioned. If it is proposed to use a report prepared by Auckland UniServices Limited for a different purpose or in a different context from that intended at the time of commissioning the work, then UniServices should be consulted to verify whether the report is being correctly interpreted. In particular it is requested that, where quoted, conclusions given in Auckland UniServices reports should be stated in full. INTRODUCTION The New Zealand king shag (Leucocarbo carunculatus) is an endemic seabird that is classed as nationally endangered (Miskelly et al., 2008). The population is confined to a small number of colonies located around the coastal margins of the outer Marlborough Sounds (South Island, New Zealand); with surveys suggesting the population is currently stable (~800 individuals surveyed in 2020; Aquaculture New Zealand, 2020; Schuckard et al., 2015). Monitoring the colonies has become a priority and research is being conducted to better understand their population dynamics and basic ecology to improve the management of the population, particularly in relation to human activities such as fishing, aquaculture and land use (Fisher & Boren, 2012). The diet of the New Zealand king shag is strongly linked to the waters surrounding their colonies and it has been suggested that anthropogenic activities, such as marine farm structures, may displace foraging habitat that could affect the population of New Zealand king shag (Fisher & Boren, 2012). -

Worse Things Happen at Sea: the Welfare of Wild-Caught Fish
[ “One of the sayings of the Holy Prophet Muhammad(s) tells us: ‘If you must kill, kill without torture’” (Animals in Islam, 2010) Worse things happen at sea: the welfare of wild-caught fish Alison Mood fishcount.org.uk 2010 Acknowledgments Many thanks to Phil Brooke and Heather Pickett for reviewing this document. Phil also helped to devise the strategy presented in this report and wrote the final chapter. Cover photo credit: OAR/National Undersea Research Program (NURP). National Oceanic and Atmospheric Administration/Dept of Commerce. 1 Contents Executive summary 4 Section 1: Introduction to fish welfare in commercial fishing 10 10 1 Introduction 2 Scope of this report 12 3 Fish are sentient beings 14 4 Summary of key welfare issues in commercial fishing 24 Section 2: Major fishing methods and their impact on animal welfare 25 25 5 Introduction to animal welfare aspects of fish capture 6 Trawling 26 7 Purse seining 32 8 Gill nets, tangle nets and trammel nets 40 9 Rod & line and hand line fishing 44 10 Trolling 47 11 Pole & line fishing 49 12 Long line fishing 52 13 Trapping 55 14 Harpooning 57 15 Use of live bait fish in fish capture 58 16 Summary of improving welfare during capture & landing 60 Section 3: Welfare of fish after capture 66 66 17 Processing of fish alive on landing 18 Introducing humane slaughter for wild-catch fish 68 Section 4: Reducing welfare impact by reducing numbers 70 70 19 How many fish are caught each year? 20 Reducing suffering by reducing numbers caught 73 Section 5: Towards more humane fishing 81 81 21 Better welfare improves fish quality 22 Key roles for improving welfare of wild-caught fish 84 23 Strategies for improving welfare of wild-caught fish 105 Glossary 108 Worse things happen at sea: the welfare of wild-caught fish 2 References 114 Appendix A 125 fishcount.org.uk 3 Executive summary Executive Summary 1 Introduction Perhaps the most inhumane practice of all is the use of small bait fish that are impaled alive on There is increasing scientific acceptance that fish hooks, as bait for fish such as tuna. -

Marine Ecology Progress Series 530:223
The following supplement accompanies the article Economic incentives and overfishing: a bioeconomic vulnerability index William W. L. Cheung*, U. Rashid Sumaila *Corresponding author: [email protected] Marine Ecology Progress Series 530: 223–232 (2015) Supplement Table S1. Country level discount rate used in the analysis Country/Territory Discount rate (%) Albania 13.4 Algeria 8.0 Amer Samoa 11.9 Andaman Is 10.0 Angola 35.0 Anguilla 10.0 Antigua Barb 10.9 Argentina 8.6 Aruba 11.3 Ascension Is 10.0 Australia 6.5 Azores Is 7.0 Bahamas 5.3 Bahrain 8.1 Baker Howland Is 7.0 Bangladesh 15.1 Barbados 9.7 Belgium 3.8 Belize 14.3 Benin 10.0 Bermuda 7.0 Bosnia Herzg 10.0 Bouvet Is 7.0 Br Ind Oc Tr 7.0 Br Virgin Is 10.0 Brazil 50.0 Brunei Darsm 10.0 Country/Territory Discount rate (%) Bulgaria 9.2 Cambodia 16.9 Cameroon 16.0 Canada 8.0 Canary Is 7.0 Cape Verde 12.3 Cayman Is 7.0 Channel Is 7.0 Chile 7.8 China Main 5.9 Christmas I. 10.0 Clipperton Is 7.0 Cocos Is 10.0 Colombia 14.3 Comoros 10.8 Congo Dem Rep 16.0 Congo Rep 16.0 Cook Is. 10.0 Costa Rica 19.9 Cote d'Ivoire 10.0 Croatia 10.0 Crozet Is 7.0 Cuba 10.0 Cyprus 6.8 Denmark 7.0 Desventuradas Is 10.0 Djibouti 11.2 Dominica 9.5 Dominican Rp 19.8 East Timor 10.0 Easter Is 10.0 Ecuador 9.4 Egypt 12.8 El Salvador 10.0 Eq Guinea 16.0 Eritrea 10.0 Estonia 10.0 Faeroe Is 7.0 Falkland Is 7.0 Fiji 6.2 Finland 7.0 Fr Guiana 10.0 Fr Moz Ch Is 10.0 Country/Territory Discount rate (%) Fr Polynesia 10.0 France 4.0 Gabon 16.0 Galapagos Is 10.0 Gambia 30.9 Gaza Strip 10.0 Georgia 20.3 Germany (Baltic) 7.0 Germany (North Sea) 7.0 Ghana 10.0 Gibraltar 7.0 Greece 7.0 Greenland 7.0 Grenada 9.9 Guadeloupe 10.0 Guam 7.0 Guatemala 12.9 Guinea 10.0 GuineaBissau 10.0 Guyana 14.6 Haiti 43.8 Heard Is 7.0 Honduras 17.6 Hong Kong 7.4 Iceland 17.3 India 11.7 Indonesia 16.0 Iran 15.0 Iraq 14.1 Ireland 2.7 Isle of Man 7.0 Israel 6.9 Italy 5.8 Jamaica 17.5 Jan Mayen 7.0 Japan (Pacific Coast) 10.0 Japan (Sea of Japan) 10.0 Jarvis Is 10.0 Johnston I. -

From Cray Rings to Closure : Aspects of the Tasmanian Fishing Industry To
/ FROM CRAY RINGS TO CLOSURE ASPECTS OF THE TASMANIAN FISHING INDUSTRY TO CIRCA 1970 BY PETER L. STOREY B.A. ,, •' ..'-· ' Submitted as part of the fulfilment of the requirement for the degree of Master of Humanities, University of Tasmania, December 1993. •"'' \ This thesis contains no material which has been accepted for the award of any other higher degree or graduate diploma in any university and, to the best of my .knowledge and belief, the thesis contains no material previously published or written by another person except where due reference is made in text of the thesis. tTK ABSTRACT The history of the Tasmanian fishing industry is traced in general terms from settlement to 1925, and in greater detail to circa 1970. The development of the industry is reviewed emphasising changes in structure, the roles of management boards and the effects of government policy. A number of public enquiries and ·their recommendations are analysed to gain an insight into the industry at various times. The roles of regional centres and their increasing participation are investigated as are the development of the west coast fisheries. Events such as the great depression, the second world war, and the emergence and decline of export markets are examined in the light of their effects on the industry in general, and fishermen in particular. The organisation of fishermen into professional associations and commercial co-operatives, their expansion, and in some cases demise, is also explored. The attitudes of fishermen to resource management, conservation, and the effects of pollution are reviewed, as are their responses to fisheries and marine regulations Particular attention is paid to the concept of fishing effort and this is examined in the light of changing fishing technology, increasing capitalisation and the availability and dependence on.-bank finance. -

61661147.Pdf
Resource Inventory of Marine and Estuarine Fishes of the West Coast and Alaska: A Checklist of North Pacific and Arctic Ocean Species from Baja California to the Alaska–Yukon Border OCS Study MMS 2005-030 and USGS/NBII 2005-001 Project Cooperation This research addressed an information need identified Milton S. Love by the USGS Western Fisheries Research Center and the Marine Science Institute University of California, Santa Barbara to the Department University of California of the Interior’s Minerals Management Service, Pacific Santa Barbara, CA 93106 OCS Region, Camarillo, California. The resource inventory [email protected] information was further supported by the USGS’s National www.id.ucsb.edu/lovelab Biological Information Infrastructure as part of its ongoing aquatic GAP project in Puget Sound, Washington. Catherine W. Mecklenburg T. Anthony Mecklenburg Report Availability Pt. Stephens Research Available for viewing and in PDF at: P. O. Box 210307 http://wfrc.usgs.gov Auke Bay, AK 99821 http://far.nbii.gov [email protected] http://www.id.ucsb.edu/lovelab Lyman K. Thorsteinson Printed copies available from: Western Fisheries Research Center Milton Love U. S. Geological Survey Marine Science Institute 6505 NE 65th St. University of California, Santa Barbara Seattle, WA 98115 Santa Barbara, CA 93106 [email protected] (805) 893-2935 June 2005 Lyman Thorsteinson Western Fisheries Research Center Much of the research was performed under a coopera- U. S. Geological Survey tive agreement between the USGS’s Western Fisheries