Downloaded from the PFAM Database50
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

AGC Kinases in Mtor Signaling, in Mike Hall and Fuyuhiko Tamanoi: the Enzymes, Vol
Provided for non-commercial research and educational use only. Not for reproduction, distribution or commercial use. This chapter was originally published in the book, The Enzymes, Vol .27, published by Elsevier, and the attached copy is provided by Elsevier for the author's benefit and for the benefit of the author's institution, for non-commercial research and educational use including without limitation use in instruction at your institution, sending it to specific colleagues who know you, and providing a copy to your institution’s administrator. All other uses, reproduction and distribution, including without limitation commercial reprints, selling or licensing copies or access, or posting on open internet sites, your personal or institution’s website or repository, are prohibited. For exceptions, permission may be sought for such use through Elsevier's permissions site at: http://www.elsevier.com/locate/permissionusematerial From: ESTELA JACINTO, AGC Kinases in mTOR Signaling, In Mike Hall and Fuyuhiko Tamanoi: The Enzymes, Vol. 27, Burlington: Academic Press, 2010, pp.101-128. ISBN: 978-0-12-381539-2, © Copyright 2010 Elsevier Inc, Academic Press. Author's personal copy 7 AGC Kinases in mTOR Signaling ESTELA JACINTO Department of Physiology and Biophysics UMDNJ-Robert Wood Johnson Medical School, Piscataway New Jersey, USA I. Abstract The mammalian target of rapamycin (mTOR), a protein kinase with homology to lipid kinases, orchestrates cellular responses to growth and stress signals. Various extracellular and intracellular inputs to mTOR are known. mTOR processes these inputs as part of two mTOR protein com- plexes, mTORC1 or mTORC2. Surprisingly, despite the many cellular functions that are linked to mTOR, there are very few direct mTOR substrates identified to date. -

A Calcium- and Calmodulin-Dependent Kinase I␣/ Microtubule Affinity Regulating Kinase 2 Signaling Cascade Mediates Calcium-Dependent Neurite Outgrowth
The Journal of Neuroscience, April 18, 2007 • 27(16):4413–4423 • 4413 Cellular/Molecular A Calcium- and Calmodulin-Dependent Kinase I␣/ Microtubule Affinity Regulating Kinase 2 Signaling Cascade Mediates Calcium-Dependent Neurite Outgrowth Nataliya V. Uboha,1 Marc Flajolet,2 Angus C. Nairn,1,2 and Marina R. Picciotto1 1Department of Psychiatry, Yale University School of Medicine, New Haven, Connecticut 06508, and 2Laboratory of Molecular and Cellular Neuroscience, The Rockefeller University, New York, New York 10021 Calcium is a critical regulator of neuronal differentiation and neurite outgrowth during development, as well as synaptic plasticity in adulthood. Calcium- and calmodulin-dependent kinase I (CaMKI) can regulate neurite outgrowth; however, the signal transduction cascades that lead to its physiological effects have not yet been elucidated. CaMKI␣ was therefore used as bait in a yeast two-hybrid assay and microtubule affinity regulating kinase 2 (MARK2)/Par-1b was identified as an interacting partner of CaMKI in three independent screens. The interaction between CaMKI and MARK2 was confirmed in vitro and in vivo by coimmunoprecipitation. CaMKI binds MARK2 within its kinase domain, but only if it is activated by calcium and calmodulin. Expression of CaMKI and MARK2 in Neuro-2A (N2a) cells and in primary hippocampal neurons promotes neurite outgrowth, an effect dependent on the catalytic activities of these enzymes. In addition, decreasing MARK2 activity blocks the ability of the calcium ionophore ionomycin to promote neurite outgrowth. Finally, CaMKI phosphorylates MARK2 on novel sites within its kinase domain. Mutation of these phosphorylation sites decreases both MARK2 kinase activity and its ability to promote neurite outgrowth. -
Protein Kinases Phosphorylation/Dephosphorylation Protein Phosphorylation Is One of the Most Important Mechanisms of Cellular Re
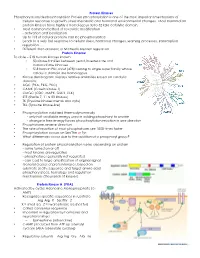
Protein Kinases Phosphorylation/dephosphorylation Protein phosphorylation is one of the most important mechanisms of cellular responses to growth, stress metabolic and hormonal environmental changes. Most mammalian protein kinases have highly a homologous 30 to 32 kDa catalytic domain. • Most common method of reversible modification - activation and localization • Up to 1/3 of cellular proteins can be phosphorylated • Leads to a very fast response to cellular stress, hormonal changes, learning processes, transcription regulation .... • Different than allosteric or Michealis Menten regulation Protein Kinome To date – 518 human kinases known • 50 kinase families between yeast, invertebrate and mammaliane kinomes • 518 human PKs, most (478) belong to single super family whose catalytic domain are homologous. • Kinase dendrogram displays relative similarities based on catalytic domains. • AGC (PKA, PKG, PKC) • CAMK (Casein kinase 1) • CMGC (CDC, MAPK, GSK3, CLK) • STE (Sterile 7, 11 & 20 kinases) • TK (Tryosine kinases memb and cyto) • TKL (Tyrosine kinase-like) • Phosphorylation stabilized thermodynamically - only half available energy used in adding phosphoryl to protein - change in free energy forces phosphorylation reaction in one direction • Phosphatases reverse direction • The rate of reaction of most phosphatases are 1000 times faster • Phosphorylation occurs on Ser/The or Tyr • What differences occur due to the addition of a phosphoryl group? • Regulation of protein phosphorylation varies depending on protein - some turned on or off -

Camk Iialpha) (C6974
Anti-CaM Kinase IIa (CaMK IIa) produced in rabbit, IgG fraction of antiserum Catalog Number C6974 Product Description residue in the autoinhibitory domain 7 (Thr286 in Anti-CaM Kinase IIa (CaMK IIa) is produced in rabbit CaMKIIa and Thr287 in CaMKIIb). Autophosphorylation using as immunogen a synthetic peptide of CaMKIIa at Thr286 has been shown to be required for (KWQIVHFHRSGAPSVLPH) corresponding to the LTP and learning.8 CaMKII activation results in C-terminal region of rat CaM Kinase IIa (amino acids switching of the kinase to a Ca2+/CaM- independent 461-478), conjugated to KLH. This sequence is state and its translocation to the PSD.9,10 identical in human, mouse and chicken CaM Kinase IIa PSD-associated CaMKII in turn phosphorylates and has limited homology (50-60%) with CaM Kinase ionotropic glutamate receptors (e.g. NMDAR, AMPA-R), IIb, g and d subunits. Whole antiserum is purified to thus providing a mechanism for increased synaptic 9-12 provide an IgG fraction of antiserum. signaling during LTP. Anti-CaM Kinase IIa recognizes rat CaM Kinase IIa Reagent (50 kDa). Applications include the detection and Supplied as a solution in 0.01 M phosphate buffered localization of CaM Kinase IIa (50 kDa) by saline, pH 7.4, containing 15 mM sodium azide. immunoblotting. Staining of CaM Kinase IIa in immunoblotting is specifically inhibited with the CaM Precautions and Disclaimer Kinase IIa immunizing peptide. This product is for R&D use only, not for drug, household, or other uses. Please consult the Material Ca2+/Calmodulin dependent protein kinase II (CaMKII) Safety Data Sheet for information regarding hazards belongs to the family of Ser/Thr protein kinases and safe handling practices. -

Intracellular Calcium Regulates Amp-Activated Protein Kinase Activity in an Oscillation-Dependent Manner
INTRACELLULAR CALCIUM REGULATES AMP-ACTIVATED PROTEIN KINASE ACTIVITY IN AN OSCILLATION-DEPENDENT MANNER Sungkwon Park, Eric M. England, Haibo Zhu, Jason M. Scheffler, Steve C. Kasten, Tracy L. Scheffler, and * David E. Gerrard Department of Animal and Poultry Sciences, Virginia Tech, Blacksburg, VA, 24061, USA *Corresponding author (phone: +1-540-231-9157; fax: +1-540-231-3010; e-mail: [email protected]) Abstract—Skeletal muscle calcium signaling is important for muscle contraction, as well as regulates many cellular processes. Calcium-regulated calmodulin dependent kinase kinase (CaMKK) has recently been identified as upstream regulator of AMP-activated protein kinase (AMPK), which is energy regulator in skeletal muscle. Although there is evidence that cytosolic calcium regulates AMPK through a series of pathways, the molecular mechanisms by which calcium regulates AMPK are poorly understood. The objective of this study is to understand the function of calcium oscillations on AMPK activity and define the specific calcium-regulated signaling molecules in this pathway. AMPK activity was increased by 2 folds in muscles from mice treated with AICAR (known AMPK activator). Administration of caffeine (calcium releasing agent) for 10 d decreased AICAR-induced AMPK activity to control level. This repressed AMPK activity was blocked by dantrolene, a ryanodine receptor stabilizer. Different calcium frequencies were simulated in C2C12 myotubes by alternating media containing caffeine and dantrolene. Changes in intracellular calcium levels were confirmed by fluorescent calcium indicator, Fura2. To define the function of calcium signaling, CaMKK or CaMK was knocked down. Low frequency calcium stimulations had a positive effect on AICAR-induced AMPK activity, whereas continuous high calcium level decreases AMPK activity suggesting a biphasic control of AMPK activity by calcium. -

A-Camkii Controls the Growth of Human Osteosarcoma by Regulating Cell Cycle Progression Kaiyu Yuan1, Leland WK Chung2, Gene P Siegal3 and Majd Zayzafoon1
Laboratory Investigation (2007) 87, 938–950 & 2007 USCAP, Inc All rights reserved 0023-6837/07 $30.00 a-CaMKII controls the growth of human osteosarcoma by regulating cell cycle progression Kaiyu Yuan1, Leland WK Chung2, Gene P Siegal3 and Majd Zayzafoon1 Osteosarcoma is the most frequent type of primary bone cancer in children and adolescents. These malignant osteoid forming tumors are characterized by their uncontrolled hyperproliferation. Here, we investigate the role of Ca2 þ / calmodulin-dependent protein kinase II (CaMKII) in the growth of human osteosarcoma. We show that a-CaMKII is expressed in human osteosarcoma cell lines and in primary osteosarcoma tissue derived from patients. The pharmaco- logic inhibition of CaMKII in MG-63 and 143B human osteosarcoma cells by KN-93 resulted in an 80 and 70% decrease in proliferation, respectively, and induced cell cycle arrest in the G0/G1 phase. The in vivo administration of KN-93 to mice xenografted with human osteosarcoma cells significantly decreased intratibial and subcutaneous tumor growth. Mechanistically, KN-93 and a-CaMKII siRNA increased p21(CIP/KIP) gene expression, protein levels, and decreased the phosphorylation of retinoblastoma protein and E2F transactivation. Furthermore, the inhibition of CaMKII decreased membrane-bound Tiam1 and GTP-bound Rac1, which are known to be involved in p21 expression and tumor growth in a variety of solid malignant neoplasms. Our results suggest that CaMKII plays a critical role in the growth of osteosarcoma, and its inhibition could be an attractive therapeutic target to combat conventional high-grade osteosarcoma in children. Laboratory Investigation (2007) 87, 938–950; doi:10.1038/labinvest.3700658; published online 16 July 2007 KEYWORDS: osteosarcoma; CaMKII; cell cycle; osteoblasts; p21; Rac1 Osteosarcomas are among the most frequent primary bone cycle progression. -

Novel Regulation of Mtor Complex 1 Signaling by Site-Specific Mtor Phosphorylation
Novel Regulation of mTOR Complex 1 Signaling by Site-Specific mTOR Phosphorylation by Bilgen Ekim Üstünel A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Cell and Developmental Biology) in The University of Michigan 2012 Doctoral Committee: Assistant Professor Diane C. Fingar, Chair Associate Professor Billy Tsai Associate Professor Anne B. Vojtek Assistant Professor Patrick J. Hu Assistant Professor Ken Inoki “Our true mentor in life is science.” (“Hayatta en hakiki mürşit ilimdir.”) Mustafa Kemal Atatürk, the founder of Turkish Republic © Bilgen Ekim Üstünel 2012 Acknowledgements This thesis would not have been possible without the enormous support and encouragement of my Ph.D. advisor Diane C. Fingar. I am sincerely thankful for her research insight and guidance during my Ph.D. training. I would like to express my great appreciation to Billy Tsai, Anne B. Vojtek, Ken Inoki, and Patrick J. Hu for serving on my thesis committee, whose advice and help have been valuable. I would like to thank all members of the Fingar, Tsai, and Verhey labs for the discussion in our group meetings. I also would like to thank the CDB administrative staff, especillay Kristen Hug, for their help. I thank Ed Feener for performing the liquid chromatography tandem mass spectrometry analysis to identify novel phosphorylation sites on mTOR and Steve Riddle for performing the in vitro kinome screen to identify candidate kinases for mTOR S2159 phosphorylation site. I thank Brian Magnuson, Hugo A. Acosta-Jaquez, and Jennifer A. Keller for contributing to my first-author paper published in Molecular and Cellular Biology Journal in 2011. -

Regulation of Ca2+/Calmodulin-Dependent Protein Kinase Kinase Β by Camp Signaling
Regulatory phosphorylation of CaMKKβ by PKA Regulation of Ca2+/calmodulin-dependent protein kinase kinase β by cAMP signaling Shota Takabatake,1,* Satomi Ohtsuka,1,* Takeyuki Sugawara, 2 Naoya Hatano,1 Naoki Kanayama,1 Masaki Magari,1 Hiroyuki Sakagami,2 and Hiroshi Tokumitsu1,** 1Applied Cell Biology, Graduate School of Interdisciplinary Science and Engineering in Health Systems, Okayama University, Okayama 700-8530 Japan, 2Department of Anatomy, Kitasato University School of Medicine, Sagamihara, Kanagawa, 252-0374 Japan **To whom correspondence should be addressed: Hiroshi Tokumitsu, Ph.D. Applied Cell Biology, Graduate School of Interdisciplinary Science and Engineering in Health Systems, Okayama University, 3-1-1 Tsushima-naka, Kita-ku, Okayama 700-8530, Japan. Tel/FAX: +81-86-251-8197; E-mail: [email protected] Notes: *S. T. and S. O. contributed equally to this work. Running title: Regulatory phosphorylation of CaMKKβ by PKA The abbreviations used are: CaMKKβ, Ca2+/CaM-dependent protein kinase kinase β; AID, autoinhibitory domain; CaM, calmodulin; CaMK, Ca2+/CaM-dependent protein kinase; AMPK, 5’AMP-activated protein kinase; PKA, cAMP-dependent protein kinase; CDK5, cyclin-dependent kinase 5; GSK3, glycogen synthase kinase 3; DAPK, death-associated kinase ConfliCt of interest: The authors declare that they have no conflict of interest with the contents of this article. 1 Regulatory phosphorylation of CaMKKβ by PKA ABSTRACT BACKGROUND: Ca2+/calmodulin-dependent protein kinase kinase (CaMKK) is a pivotal activator of CaMKI, CaMKIV and 5’-AMP-activated protein kinase (AMPK), controlling Ca2+-dependent intracellular signaling including various neuronal, metabolic and pathophysiological responses. Recently, we demonstrated that CaMKKβ is feedback phosphorylated at Thr144 by the downstream AMPK, resulting in the conversion of CaMKKβ into Ca2+/CaM-dependent enzyme. -

MTOR Signaling and Metabolism in Early T Cell Development
G C A T T A C G G C A T genes Review MTOR Signaling and Metabolism in Early T Cell Development Guy Werlen, Ritika Jain and Estela Jacinto * Department of Biochemistry and Molecular Biology, Robert Wood Johnson Medical School, Rutgers University, Piscataway, NJ 08854, USA; [email protected] (G.W.); [email protected] (R.J.) * Correspondence: [email protected]; Tel.: +1-732-235-4476 Abstract: The mechanistic target of rapamycin (mTOR) controls cell fate and responses via its func- tions in regulating metabolism. Its role in controlling immunity was unraveled by early studies on the immunosuppressive properties of rapamycin. Recent studies have provided insights on how metabolic reprogramming and mTOR signaling impact peripheral T cell activation and fate. The contribution of mTOR and metabolism during early T-cell development in the thymus is also emerging and is the subject of this review. Two major T lineages with distinct immune functions and peripheral homing organs diverge during early thymic development; the αβ- and γδ-T cells, which are defined by their respective TCR subunits. Thymic T-regulatory cells, which have immunosup- pressive functions, also develop in the thymus from positively selected αβ-T cells. Here, we review recent findings on how the two mTOR protein complexes, mTORC1 and mTORC2, and the signaling molecules involved in the mTOR pathway are involved in thymocyte differentiation. We discuss emerging views on how metabolic remodeling impacts early T cell development and how this can be mediated via mTOR signaling. Keywords: mTOR; mTORC1; mTORC2; thymocytes; T lymphocytes; early T cell development; T-cell metabolism Citation: Werlen, G.; Jain, R.; Jacinto, E. -

Dual-Specificity, Tyrosine Phosphorylation-Regulated Kinases
International Journal of Molecular Sciences Review Dual-Specificity, Tyrosine Phosphorylation-Regulated Kinases (DYRKs) and cdc2-Like Kinases (CLKs) in Human Disease, an Overview Mattias F. Lindberg and Laurent Meijer * Perha Pharmaceuticals, Perharidy Peninsula, 29680 Roscoff, France; [email protected] * Correspondence: [email protected] Abstract: Dual-specificity tyrosine phosphorylation-regulated kinases (DYRK1A, 1B, 2-4) and cdc2- like kinases (CLK1-4) belong to the CMGC group of serine/threonine kinases. These protein ki- nases are involved in multiple cellular functions, including intracellular signaling, mRNA splicing, chromatin transcription, DNA damage repair, cell survival, cell cycle control, differentiation, ho- mocysteine/methionine/folate regulation, body temperature regulation, endocytosis, neuronal development, synaptic plasticity, etc. Abnormal expression and/or activity of some of these kinases, DYRK1A in particular, is seen in many human nervous system diseases, such as cognitive deficits associated with Down syndrome, Alzheimer’s disease and related diseases, tauopathies, demen- tia, Pick’s disease, Parkinson’s disease and other neurodegenerative diseases, Phelan-McDermid syndrome, autism, and CDKL5 deficiency disorder. DYRKs and CLKs are also involved in dia- betes, abnormal folate/methionine metabolism, osteoarthritis, several solid cancers (glioblastoma, breast, and pancreatic cancers) and leukemias (acute lymphoblastic leukemia, acute megakaryoblas- Citation: Lindberg, M.F.; Meijer, L. tic leukemia), viral infections (influenza, HIV-1, HCMV, HCV, CMV, HPV), as well as infections Dual-Specificity, Tyrosine caused by unicellular parasites (Leishmania, Trypanosoma, Plasmodium). This variety of pathological Phosphorylation-Regulated Kinases implications calls for (1) a better understanding of the regulations and substrates of DYRKs and (DYRKs) and cdc2-Like Kinases CLKs and (2) the development of potent and selective inhibitors of these kinases and their evaluation (CLKs) in Human Disease, an as therapeutic drugs. -

Phosphorylation-Dependent Sub-Functionalization of the Calcium-Dependent Protein Kinase CPK28
bioRxiv preprint doi: https://doi.org/10.1101/2020.10.16.338442; this version posted October 17, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Phosphorylation-dependent sub-functionalization of 2 the calcium-dependent protein kinase CPK28 3 4 5 Melissa Bredow1,#, Kyle W. Bender2,#,a, Alexandra Johnson Dingee1,b, Danalyn R. 6 Holmes1,c, Alysha Thomson1,d, Danielle Ciren1,e, Cailun A. S. Tanney1,f, Katherine E. 7 Dunning1,3, Marco Trujillo3, Steven C. Huber2, and Jacqueline Monaghan1,* 8 9 10 1 Department of Biology, Queen’s University, Kingston, Canada 11 2 Department of Plant Biology, School of Integrative Biology, University of Illinois- 12 Urbana-Champaign, Urbana-Champaign, United States of America 13 3 Department of Cell Biology, University of Freiburg, Freiburg, Germany 14 15 # These authors contributed equally to this work 16 * Corresponding author: [email protected] 17 18 19 a Current address: University of Zurich, Department of Plant and Microbial Biology, 20 Zurich, Switzerland 21 b Current address: Faculty of Law, Western University, London, Canada 22 c Current address: Center for Plant Molecular Biology, University of Tuebingen, 23 Tuebingen, Germany 24 d Current address: Faculty of Medicine, McMaster University, Hamilton, Canada 25 e Current address: Cold Spring Harbor Laboratory, Cold Spring Harbor, United States of 26 America 27 f Current address: Department of Plant Science, McGill University, Montreal, Canada 28 29 30 Keywords: calcium-dependent protein kinases, calcium-dependent protein kinase 28, 31 botrytis-induced kinase 1, phosphorylation, immune signaling, stem elongation, 32 biochemical mechanism 33 34 35 36 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.10.16.338442; this version posted October 17, 2020. -

Structural Basis and Prediction of Substrate Specificity in Protein Serine͞threonine Kinases
Structural basis and prediction of substrate specificity in protein serine͞threonine kinases Ross I. Brinkworth, Robert A. Breinl, and Bostjan Kobe* Department of Biochemistry and Molecular Biology and Institute for Molecular Bioscience, University of Queensland, Brisbane, Queensland 4072, Australia Edited by Susan S. Taylor, University of California at San Diego, La Jolla, CA, and approved November 18, 2002 (received for review July 16, 2002) The large number of protein kinases makes it impractical to an automated prediction of optimal substrate peptides by using determine their specificities and substrates experimentally. Using only the amino acid sequence of the protein kinase as input. To the available crystal structures, molecular modeling, and sequence explore the utility of the method, we used PREDIKIN to analyze analyses of kinases and substrates, we developed a set of rules the signaling pathways in two cellular processes in yeast, cell governing the binding of a heptapeptide substrate motif (sur- cycle control and DNA damage checkpoints, and predicted new rounding the phosphorylation site) to the kinase and implemented connections in these pathways. Our method should be generally these rules in a web-interfaced program for automated prediction applicable to identifying possible substrate proteins for protein of optimal substrate peptides, taking only the amino acid sequence serine͞threonine kinases and should aid in unraveling signaling of a protein kinase as input. We show the utility of the method by pathways in which these proteins may be involved. analyzing yeast cell cycle control and DNA damage checkpoint pathways. Our method is the only available predictive method Materials and Methods generally applicable for identifying possible substrate proteins for Identification of Binding Sites and Sequence Motifs.