Predicting Spatial Patterns of Plant Biodiversity: from Species to Communities
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Phyteuma Vagneri A. Kern. (Campanulaceae) 238-241 ©Naturhistorisches Museum Wien, Download Unter
ZOBODAT - www.zobodat.at Zoologisch-Botanische Datenbank/Zoological-Botanical Database Digitale Literatur/Digital Literature Zeitschrift/Journal: Annalen des Naturhistorischen Museums in Wien Jahr/Year: 2013 Band/Volume: 115B Autor(en)/Author(s): Pachschwöll Clemens Artikel/Article: Short Communication: Typification of Kerner names 8: Phyteuma vagneri A. Kern. (Campanulaceae) 238-241 ©Naturhistorisches Museum Wien, download unter www.biologiezentrum.at Ann. Naturhist. Mus. Wien, B 115 238-241 Wien, März 2013 Short Communication Typification of Kerner names 8: Phyteuma vagneri A. K e r n . (Campanulaceae) C. Pachschwöll* Key Words: Phyteuma vagneri, Campanulaceae, Carpathians, Romania, Ukraine. In the 1870s, Ludwig Vägner [Lajos Wagner] (1815-1888) distributed herbarium spec imens of Phyteuma (A n o n y m u s 1877, C o n e r t 1999). Of these, one accession bears the label “ Phyteuma orbiculare L. / Auf Alpentriften und Gebirgswiesen zu Körösmezö / in der Marmaross. / Juni - Juli, [sine anno], L. Vägner” another one “Flora der Marmaros / Phyteuma / Auf der Rahoer Alpe Terentin / Juni 1873, Vägner” (dozens of vouch- ers in WU-Kemer!). Vägners specimens from the historical region Märamaros, nowa- days northem Romania and southwestem Ukraine, were recognized by Kerner as a new species. V ä g n e r mentioned “Phyteuma Vagneri A . K e r n e r . A körösmezei Bliznicza, a rahöi Terentin havasokon. Junius-juliusban.” in his list of plants from Märamaros county with two localities “Blisnicza”[Bjih3hhu;h] and “Terentin” [TepeHTHH], but without a descrip tion (W a g n e r 1876, se e also B e c k 1883: 182). In 1883 K e r n e r formally described this Carpathian endemic when he distributed this plant in his “Flora exsiccata Austro-Hungarica” and dedicated it to Vägner (K e r n e r 1883, K e r n e r 1884: 107-108). -

Relationships Among Levels of Biodiversity and the Relevance of Intraspecific Diversity in Conservation – a Project Synopsis F
ARTICLE IN PRESS Perspectives in Plant Ecology, Evolution and Systematics Perspectives in Plant Ecology, Evolution and Systematics 10 (2008) 259–281 www.elsevier.de/ppees Relationships among levels of biodiversity and the relevance of intraspecific diversity in conservation – a project synopsis F. Gugerlia,Ã, T. Englischb, H. Niklfeldb, A. Tribschc,1, Z. Mirekd, M. Ronikierd, N.E. Zimmermanna, R. Holdereggera, P. Taberlete, IntraBioDiv Consortium2,3 aWSL Swiss Federal Research Institute, Zu¨rcherstrasse 111, 8903 Birmensdorf, Switzerland bDepartment of Biogeography, University of Vienna, Rennweg 14, 1030 Wien, Austria cDepartment of Systematic and Evolutionary Botany, Rennweg 14, 1030 Wien, Austria dDepartment of Vascular Plant Systematics, Institute of Botany, Polish Academy of Science, Krako´w, Lubicz 46, 31-512 Krako´w, Poland eLaboratoire d’Ecologie Alpine (LECA), CNRS UMR 5553, University Joseph Fourier, BP 53, 2233 Rue de la Piscine, 38041 Grenoble Cedex 9, France Received 11 June 2007; received in revised form 4 June 2008; accepted 9 July 2008 Abstract The importance of the conservation of all three fundamental levels of biodiversity (ecosystems, species and genes) has been widely acknowledged, but only in recent years it has become technically feasible to consider intraspecific diversity, i.e. the genetic component to biodiversity. In order to facilitate the assessment of biodiversity, considerable efforts have been made towards identifying surrogates because the efficient evaluation of regional biodiversity would help in designating important areas for nature conservation at larger spatial scales. However, we know little about the fundamental relationships among the three levels of biodiversity, which impedes the formulation of a general, widely applicable concept of biodiversity conservation through surrogates. -
(A) Journals with the Largest Number of Papers Reporting Estimates Of
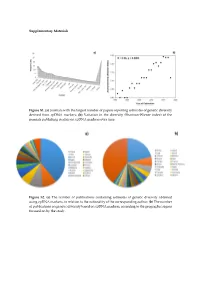
Supplementary Materials Figure S1. (a) Journals with the largest number of papers reporting estimates of genetic diversity derived from cpDNA markers; (b) Variation in the diversity (Shannon-Wiener index) of the journals publishing studies on cpDNA markers over time. Figure S2. (a) The number of publications containing estimates of genetic diversity obtained using cpDNA markers, in relation to the nationality of the corresponding author; (b) The number of publications on genetic diversity based on cpDNA markers, according to the geographic region focused on by the study. Figure S3. Classification of the angiosperm species investigated in the papers that analyzed genetic diversity using cpDNA markers: (a) Life mode; (b) Habitat specialization; (c) Geographic distribution; (d) Reproductive cycle; (e) Type of flower, and (f) Type of pollinator. Table S1. Plant species identified in the publications containing estimates of genetic diversity obtained from the use of cpDNA sequences as molecular markers. Group Family Species Algae Gigartinaceae Mazzaella laminarioides Angiospermae Typhaceae Typha laxmannii Angiospermae Typhaceae Typha orientalis Angiospermae Typhaceae Typha angustifolia Angiospermae Typhaceae Typha latifolia Angiospermae Araliaceae Eleutherococcus sessiliflowerus Angiospermae Polygonaceae Atraphaxis bracteata Angiospermae Plumbaginaceae Armeria pungens Angiospermae Aristolochiaceae Aristolochia kaempferi Angiospermae Polygonaceae Atraphaxis compacta Angiospermae Apocynaceae Lagochilus macrodontus Angiospermae Polygonaceae Atraphaxis -

Piano Di Gestione Del Sic/Zps It3310001 “Dolomiti Friulane”
Piano di Gestione del SIC/ZPS IT 3310001 “Dolomiti Friulane” – ALLEGATO 2 PIANO DI GESTIONE DEL SIC/ZPS IT3310001 “DOLOMITI FRIULANE” ALLEGATO 2 ELENCO DELLE SPECIE FLORISTICHE E SCHEDE DESCRITTIVE DELLE SPECIE DI IMPORTANZA COMUNITARIA Agosto 2012 Responsabile del Piano : Ing. Alessandro Bardi Temi Srl Piano di Gestione del SIC/ZPS IT 3310001 “Dolomiti Friulane” – ALLEGATO 2 Classe Sottoclasse Ordine Famiglia Specie 1 Lycopsida Lycopodiatae Lycopodiales Lycopodiaceae Huperzia selago (L.)Schrank & Mart. subsp. selago 2 Lycopsida Lycopodiatae Lycopodiales Lycopodiaceae Diphasium complanatum (L.) Holub subsp. complanatum 3 Lycopsida Lycopodiatae Lycopodiales Lycopodiaceae Lycopodium annotinum L. 4 Lycopsida Lycopodiatae Lycopodiales Lycopodiaceae Lycopodium clavatum L. subsp. clavatum 5 Equisetopsida Equisetatae Equisetales Equisetaceae Equisetum arvense L. 6 Equisetopsida Equisetatae Equisetales Equisetaceae Equisetum hyemale L. 7 Equisetopsida Equisetatae Equisetales Equisetaceae Equisetum palustre L. 8 Equisetopsida Equisetatae Equisetales Equisetaceae Equisetum ramosissimum Desf. 9 Equisetopsida Equisetatae Equisetales Equisetaceae Equisetum telmateia Ehrh. 10 Equisetopsida Equisetatae Equisetales Equisetaceae Equisetum variegatum Schleich. ex Weber & Mohr 11 Polypodiopsida Polypodiidae Polypodiales Adiantaceae Adiantum capillus-veneris L. 12 Polypodiopsida Polypodiidae Polypodiales Hypolepidaceae Pteridium aquilinum (L.)Kuhn subsp. aquilinum 13 Polypodiopsida Polypodiidae Polypodiales Cryptogrammaceae Phegopteris connectilis (Michx.)Watt -

151-170 Peruzzi
INFORMATORE BOTANICO ITALIANO, 42 (1) 151-170, 2010 151 Checklist dei generi e delle famiglie della flora vascolare italiana L. PERUZZI ABSTRACT - Checklist of genera and families of Italian vascular flora -A checklist of the genera and families of vascular plants occurring in Italy is presented. The families were grouped according to the main six taxonomic groups (e.g. sub- classes: Lycopodiidae, Ophioglossidae, Equisetidae, Polypodiidae, Pinidae, Magnoliidae) and put in systematic order accord- ing to the most recent criteria (e.g. APG system etc.). The genera within each family are arranged in alphabetic order and delimited through recent literature survey. 55 orders (51 native and 4 exotic) and 173 families were recorded (158 + 24), for a total of 1297 genera (1060 + 237). Families with the highest number of genera were Poaceae (126 + 23) and Asteraceae (125 + 23), followed by Apiaceae (85 + 5), Brassicaceae (64 + 5), Fabaceae (45 + 19) etc. A single genus resulted narrow endemic of Italy (Sicily): the monotypic Petagnaea (Apiaceae). Three more genera are instead endemic to Sardinia and Corse: Morisia (Brassicaceae), Castroviejoa and Nananthea (Asteraceae). Key words: classification, families, flora, genera, Italy Ricevuto il 16 Novembre 2009 Accettato il 5 Marzo 2010 INTRODUZIONE Successivamente alla “Flora d’Italia” di PIGNATTI Flora Europaea (BAGELLA, URBANI, 2006; (1982), i progressi negli studi biosistematici e filoge- BOCCHIERI, IIRITI, 2006; LASTRUCCI, RAFFAELLI, netici hanno prodotto una enorme mole di cono- 2006; ROMANO et al., 2006; MAIORCA et al., 2007; scenze circa le piante vascolari. Nella recente CORAZZI, 2008) a tentantivi di aggiormanento Checklist della flora vascolare italiana e sua integra- “misti”, basati sul totale o parziale accoglimento di zione (CONTI et al., 2005, 2007a) non vengono prese classificazioni successive, spesso non ulteriormente in considerazione le famiglie di appartenenza delle specificate (es. -

A Review of European Progress Towards the Global Strategy for Plant Conservation 2011-2020
A review of European progress towards the Global Strategy for Plant Conservation 2011-2020 1 A review of European progress towards the Global Strategy for Plant Conservation 2011-2020 The geographical area of ‘Europe’ includes the forty seven countries of the Council of Europe and Belarus: Albania, Andorra, Armenia, Austria, Azerbaijan, Belarus, Belgium, Bosnia-Herzegovina, Bulgaria, Croatia, Cyprus, Czech Republic, Denmark, Estonia, Finland, France, Georgia, Germany, Greece, Hungary, Iceland, Ireland, Italy, Latvia, Liechtenstein, Lithuania, Luxembourg, Malta, Republic of Moldova, Monaco, Montenegro, Netherlands, North Macedonia, Norway, Poland, Portugal, Romania, Russian Federation, San Marino, Serbia, Slovakia, Slovenia, Spain, Sweden, Switzerland, Turkey, Ukraine, United Kingdom. Front Cover Image: Species rich meadow with Papaver paucifoliatum, Armenia, Anna Asatryan. Disclaimer: The designations employed and the presentation of material in this publication do not imply the expression of any opinion whatsoever on the part of the copyright holders concerning the legal status of any country, territory, city or area or of its authorities, or concerning the delimitation of its frontiers or boundaries. The mentioning of specific companies or products does not imply that they are endorsed or recommended by PLANTA EUROPA or Plantlife International or preferred to others that are not mentioned – they are simply included as examples. All reasonable precautions have been taken by PLANTA EUROPA and Plantlife International to verify the information contained in this publication. However, the published material is being distributed without warranty of any kind, either expressed or implied. The responsibility for the interpretation and use of the material lies with the reader. In no event shall PLANTA EUROPA, Plantlife International or the authors be liable for any consequences whatsoever arising from its use. -

Wildlife Travel Burren 2018
The Burren 2018 species list and trip report, 7th-12th June 2018 WILDLIFE TRAVEL The Burren 2018 s 1 The Burren 2018 species list and trip report, 7th-12th June 2018 Day 1: 7th June: Arrive in Lisdoonvarna; supper at Rathbaun Hotel Arriving by a variety of routes and means, we all gathered at Caherleigh House by 6pm, sustained by a round of fresh tea, coffee and delightful home-made scones from our ever-helpful host, Dermot. After introductions and some background to the geology and floral elements in the Burren from Brian (stressing the Mediterranean component of the flora after a day’s Mediterranean heat and sun), we made our way to the Rathbaun, for some substantial and tasty local food and our first taste of Irish music from the three young ladies of Ceolan, and their energetic four-hour performance (not sure any of us had the stamina to stay to the end). Day 2: 8th June: Poulsallach At 9am we were collected by Tony, our driver from Glynn’s Coaches for the week, and following a half-hour drive we arrived at a coastal stretch of species-rich limestone pavement which represented the perfect introduction to the Burren’s flora: a stunningly beautiful mix of coastal, Mediterranean, Atlantic and Arctic-Alpine species gathered together uniquely in a natural rock garden. First impressions were of patchy grassland, sparkling with heath spotted- orchids Dactylorhiza maculata ericetorum and drifts of the ubiquitous and glowing-purple bloody crane’s-bill Geranium sanguineum, between bare rock. A closer look revealed a diverse and colourful tapestry of dozens of flowers - the yellows of goldenrod Solidago virgaurea, kidney-vetch Anthyllis vulneraria, and bird’s-foot trefoil Lotus corniculatus (and its attendant common blue butterflies Polyommatus Icarus), pink splashes of wild thyme Thymus polytrichus and the hairy local subspecies of lousewort Pedicularis sylvatica ssp. -

Review of Coverage of the National Vegetation Classification
JNCC Report No. 302 Review of coverage of the National Vegetation Classification JS Rodwell, JC Dring, ABG Averis, MCF Proctor, AJC Malloch, JHJ Schaminée, & TCD Dargie July 2000 This report should be cited as: Rodwell, JS, Dring, JC, Averis, ABG, Proctor, MCF, Malloch, AJC, Schaminée, JNJ, & Dargie TCD, 2000 Review of coverage of the National Vegetation Classification JNCC Report, No. 302 © JNCC, Peterborough 2000 For further information please contact: Habitats Advice Joint Nature Conservation Committee Monkstone House, City Road, Peterborough PE1 1JY UK ISSN 0963-8091 1 2 Contents Preface .............................................................................................................................................................. 4 Acknowledgements .......................................................................................................................................... 4 1 Introduction.............................................................................................................................................. 5 1.1 Coverage of the original NVC project......................................................................................................... 5 1.2 Generation of NVC-related data by the community of users ...................................................................... 5 2 Methodology............................................................................................................................................. 7 2.1 Reviewing the wider European scene......................................................................................................... -

Chromosomenzahlen Von Hieracium (Compositae, Cichorieae) – Teil 5
Berichte der Bayerischen Botanischen Gesellschaft 80: 141-160, 2010 141 Chromosomenzahlen von Hieracium (Compositae, Cichorieae) – Teil 5 FRANZ SCHUHWERK Zusamenfassung: Erstmals festgestellte Chromosomenzahlen (die Unterart steht in Klammern, wenn die Ploidiestufe auch für die Sammelart neu ist): Hieracium atratum ssp. atratum und ssp. zin- kenense: 2n = 36, H. benzianum: 2n = 36, H. dentatum ssp. basifoliatum: 2n = 27 und 36, H. den- tatum (ssp. dentatum, ssp. expallens und ssp. prionodes): jeweils 2n = 36, H. dentatum ssp. oblongifolium: 2n = 27, H. dollineri (ssp. dollineri): 2n = 18, H. glabratum (ssp. glabratum): 2n = 27 und 36, H. glabratum ssp. trichoneurum: 2n = 36, H. jurassicum ssp. cichoriaceum: 2n = 27, H. leucophaeum: 2n = 27, H. pallescens ssp. muroriforme: 2n = 36, H. pilosum ssp. villosiceps: 2n = 27, H. porrectum: 2n = 45, H. pospichalii (ssp. pospichalii): 2n = 27, H. scorzonerifolium (ssp. fle- xile): 2n = 36, H. scorzonerifolium ssp. pantotrichum und ssp. triglaviense: jeweils 2n = 27, H. subspeciosum (ssp. subspeciosum): 2n = 27 und („ssp. lantschfeldense“): 2n = 36, H. umbrosum (ssp. oleicolor): 2n = 27, H. valdepilosum ssp. oligophyllum: 2n = 27, H. valdepilosum ssp. ra- phiolepium: 2n = 36, H. valdepilosum (ssp. subsinuatum): 2n = 18, H. villosum ssp. glaucifrons: 2n = 27. Erstmals aus den Alpen insgesamt wird ein pentaploides, erstmals aus den Nordalpen ein di- ploides Eu-Hieracium nachgewiesen. Zur Unterscheidung von H. subglaberrimum von ähnlichen Arten wird eine Tabelle vorgelegt. Die ostalpischen Sippen von H. chondrillifolium werden als getrennt zu behandelndes H. subspe- ciosum vorgeschlagen (eine Umkombination: H. subspeciosum ssp. jabornegii). Für Hieracium scorzonerifolium ssp. pantotrichum wird erstmals die Verbreitung in Bayern dargestellt. Summary: For the first time determined chromosome numbers: Hieracium atratum ssp. -

Circumscribing Genera in the European Orchid Flora: a Subjective
Ber. Arbeitskrs. Heim. Orchid. Beiheft 8; 2012: 94 - 126 Circumscribing genera in the European orchid lora: a subjective critique of recent contributions Richard M. BATEMAN Keywords: Anacamptis, Androrchis, classiication, evolutionary tree, genus circumscription, monophyly, orchid, Orchidinae, Orchis, phylogeny, taxonomy. Zusammenfassung/Summary: BATEMAN , R. M. (2012): Circumscribing genera in the European orchid lora: a subjective critique of recent contributions. – Ber. Arbeitskrs. Heim. Orch. Beiheft 8; 2012: 94 - 126. Die Abgrenzung von Gattungen oder anderen höheren Taxa erfolgt nach modernen Ansätzen weitestgehend auf der Rekonstruktion der Stammesgeschichte (Stamm- baum-Theorie), mit Hilfe von großen Daten-Matrizen. Wenngleich aufgrund des Fortschritts in der DNS-Sequenzierungstechnik immer mehr Merkmale in der DNS identiiziert werden, ist es mindestens genauso wichtig, die Anzahl der analysierten Planzen zu erhöhen, um genaue Zuordnungen zu erschließen. Die größere Vielfalt mathematischer Methoden zur Erstellung von Stammbäumen führt nicht gleichzeitig zu verbesserten Methoden zur Beurteilung der Stabilität der Zweige innerhalb der Stammbäume. Ein weiterer kontraproduktiver Trend ist die wachsende Tendenz, diverse Datengruppen mit einzelnen Matrizen zu verquicken, die besser einzeln analysiert würden, um festzustellen, ob sie ähnliche Schlussfolgerungen bezüglich der Verwandtschaftsverhältnisse liefern. Ein Stammbaum zur Abgrenzung höherer Taxa muss nicht so robust sein, wie ein Stammbaum, aus dem man Details des Evo- lutionsmusters -

Impacts Des Dépôts Atmosphériques Azotés Sur La Biodiversité Et Le Fonctionnement Des Pelouses Subalpines Pyrénéennes
Impacts des d´ep^otsatmosph´eriquesazot´essur la biodiversit´eet le fonctionnement des pelouses subalpines pyr´en´eennes Marion Boutin To cite this version: Marion Boutin. Impacts des d´ep^otsatmosph´eriques azot´essur la biodiversit´eet le fonction- nement des pelouses subalpines pyr´en´eennes.Sciences de la Terre. Universit´ePaul Sabatier - Toulouse III, 2015. Fran¸cais. <NNT : 2015TOU30185>. <tel-01357681> HAL Id: tel-01357681 https://tel.archives-ouvertes.fr/tel-01357681 Submitted on 30 Aug 2016 HAL is a multi-disciplinary open access L'archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destin´eeau d´ep^otet `ala diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publi´esou non, lished or not. The documents may come from ´emanant des ´etablissements d'enseignement et de teaching and research institutions in France or recherche fran¸caisou ´etrangers,des laboratoires abroad, or from public or private research centers. publics ou priv´es. Avant-propos Cette thèse a été financée pour une durée de trois ans par une bourse du Ministère de l’Enseignement Supérieur et de la Recherche (contrat doctoral avec charge d’enseignement) puis pour une durée de neuf mois par le Labex TULIP, la région Midi- Pyrénées et l’ADEME (Agence de l’Environnement et de la Maitrise de l’Energie). Ces recherches ont été réalisées dans le cadre du projet ANEMONE cofinancé par la Communauté de Travail des Pyrénées (régions Midi-Pyrénées N°11051284, Aquitaine N°1262C0013 et Huesca N°CTPP13/11), l’ADEME (N°1262C0013) et l’OHM (Observatoire Homme-Milieu) du Haut Vicdessos. -

Phylogeny and Biogeography of Isophyllous Species of Campanula (Campanulaceae) in the Mediterranean Area
Systematic Botany (2006), 31(4): pp. 862–880 # Copyright 2006 by the American Society of Plant Taxonomists Phylogeny and Biogeography of Isophyllous Species of Campanula (Campanulaceae) in the Mediterranean Area JEONG-MI PARK,1 SANJA KOVACˇ IC´ ,2 ZLATKO LIBER,2 WILLIAM M. M. EDDIE,3 and GERALD M. SCHNEEWEISS4,5 1Department of Systematic and Evolutionary Botany, Institute of Botany, University of Vienna, Rennweg 14, A-1030 Vienna, Austria; 2Botanical Department and Botanical Garden of the Faculty of Science, University of Zagreb, HR-10000 Zagreb, Croatia; 3Office of Lifelong Learning, University of Edinburgh, 11 Buccleuch Place, Edinburgh, EH8 9LW, Scotland, U.K.; 4Department of Biogeography and Botanical Garden, Institute of Botany, University of Vienna, Rennweg 14, A-1030 Vienna, Austria 5Author for correspondence ([email protected]) Communicating Editor: Thomas G. Lammers ABSTRACT. Sequence data from the nuclear internal transcribed spacer (ITS) were used to infer phylogenetic relationships within a morphologically, karyologically, and geographically well-defined group of species of Campanula (Campanulaceae), the Isophylla group. Although belonging to the same clade within the highly paraphyletic Campanula, the Rapunculus clade, members of the Isophylla group do not form a monophyletic group but fall into three separate clades: (i) C. elatines and C. elatinoides in the Alps; (ii) C. fragilis s.l. and C. isophylla with an amphi-Tyrrhenian distribution; and (iii) the garganica clade with an amphi-Adriatic distribution, comprised of C. fenestrellata s.l., C. garganica s.l., C. portenschlagiana, C. poscharskyana, and C. reatina. Taxa currently classified as subspecies of C. garganica (garganica, cephallenica, acarnanica) and C. fenestrellata subsp.