Issn No: 2326-0793
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

RT² Profiler PCR Array (384-Well Format) Human Apoptosis 384HT
RT² Profiler PCR Array (384-Well Format) Human Apoptosis 384HT Cat. no. 330231 PAHS-3012ZE For pathway expression analysis Format For use with the following real-time cyclers RT² Profiler PCR Array, Applied Biosystems® models 7900HT (384-well block), Format E ViiA™ 7 (384-well block); Bio-Rad CFX384™ RT² Profiler PCR Array, Roche® LightCycler® 480 (384-well block) Format G Description The Human Apoptosis RT² Profiler PCR Array profiles the expression of 370 key genes involved in apoptosis, or programmed cell death. The array includes the TNF ligands and their receptors; members of the bcl-2 family, BIR (baculoviral IAP repeat) domain proteins, CARD domain (caspase recruitment domain) proteins, death domain proteins, TRAF (TNF receptor-associated factor) domain proteins and caspases. These 370 genes are also grouped by their functional contribution to apoptosis, either anti-apoptosis or induction of apoptosis. Using real-time PCR, you can easily and reliably analyze expression of a focused panel of genes related to apoptosis with this array. For further details, consult the RT² Profiler PCR Array Handbook. Sample & Assay Technologies Shipping and storage RT² Profiler PCR Arrays in formats E and G are shipped at ambient temperature, on dry ice, or blue ice packs depending on destination and accompanying products. For long term storage, keep plates at –20°C. Note: Ensure that you have the correct RT² Profiler PCR Array format for your real-time cycler (see table above). Note: Open the package and store the products appropriately immediately -
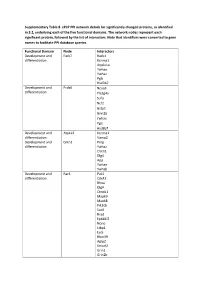
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4 -

Characterization of Genes in the CFTR-Mediated Innate Immune Response
The University of Maine DigitalCommons@UMaine Honors College 5-2012 Characterization of Genes in the CFTR-Mediated Innate Immune Response Eric Peterman Follow this and additional works at: https://digitalcommons.library.umaine.edu/honors Part of the Cell and Developmental Biology Commons, and the Molecular Biology Commons Recommended Citation Peterman, Eric, "Characterization of Genes in the CFTR-Mediated Innate Immune Response" (2012). Honors College. 71. https://digitalcommons.library.umaine.edu/honors/71 This Honors Thesis is brought to you for free and open access by DigitalCommons@UMaine. It has been accepted for inclusion in Honors College by an authorized administrator of DigitalCommons@UMaine. For more information, please contact [email protected]. CHARACTERIZATION OF GENES IN THE CFTR-MEDIATED INNATE IMMUNE RESPONSE by Eric Peterman A Thesis Submitted in Partial Fulfillment of the Requirements for a Degree with Honors (Biochemistry, Molecular and Cellular Biology) The Honors College University of Maine May 2012 Advisory Committee: Carol Kim, Professor, Molecular & Biomedical Sciences Robert Gundersen, Associate Professor, Molecular & Biomedical Sciences John Singer, Professor, Molecular & Biomedical Sciences Julie Gosse, Assistant Professor, Molecular & Biomedical Sciences Keith Hutchison, Professor, Molecular & Biomedical Sciences Mark Haggerty, Lecturer, Rezendes Preceptor for Civic Engagement Abstract: Recently, the Kim Lab has shown that the cystic fibrosis transmembrane conductance regulator (cftr) gene is responsible for mediating resistance to Pseudomonas aeruginosa in a zebrafish infection model. Using the Gene Expression Omnibus, an NCBI functional genomics data repository, it was determined that Smad3, a transcription factor in the TGF-β signaling pathway, is upregulated in the presence of P. aeruginosa. It was found that in our zebrafish model, the Smad3 paralogs Smad3a and Smad3b are upregulated following microinjection of a cftr antisense morpholino oligomer. -

Evolution of Gremlin 2 in Cetartiodactyl Mammals: Gene Loss Coincides with Lack of Upper Jaw Incisors in Ruminants
Evolution of gremlin 2 in cetartiodactyl mammals: gene loss coincides with lack of upper jaw incisors in ruminants Juan C. Opazo1, Kattina Zavala1, Paola Krall2 and Rodrigo A. Arias3 1 Instituto de Ciencias Ambientales y Evolutivas, Universidad Austral de Chile, Valdivia, Chile 2 Unidad de Nefrología, Universidad Austral de Chile, Valdivia, Chile 3 Instituto de Producción Animal, Universidad Austral de Chile, Valdivia, Chile ABSTRACT Understanding the processes that give rise to genomic variability in extant species is an active area of research within evolutionary biology. With the availability of whole genome sequences, it is possible to quantify different forms of variability such as variation in gene copy number, which has been described as an important source of genetic variability and in consequence of phenotypic variability. Most of the research on this topic has been focused on understanding the biological significance of gene duplication, and less attention has been given to the evolutionary role of gene loss. Gremlin 2 is a member of the DAN gene family and plays a significant role in tooth development by blocking the ligand-signaling pathway of BMP2 and BMP4. The goal of this study was to investigate the evolutionary history of gremlin 2 in cetartiodactyl mammals, a group that possesses highly divergent teeth morphology. Results from our analyses indicate that gremlin 2 has experienced a mixture of gene loss, gene duplication, and rate acceleration. Although the last common ancestor of cetartiodactyls possessed a single gene copy, pigs and camels are the only cetartiodactyl groups that have retained gremlin 2. According to the phyletic distribution of this gene and synteny analyses, we propose that gremlin 2 was lost in the common ancestor of ruminants and cetaceans between 56.3 and 63.5 million years ago as a product of a chromosomal rearrangement. -

14-3-3Ζ, a Novel Androgen-Responsive Gene, Is Upregulated in Prostate Cancer And
Author Manuscript Published OnlineFirst on August 17, 2012; DOI: 10.1158/1078-0432.CCR-12-0281 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. 14-3-3ζ, a novel androgen-responsive gene, is upregulated in prostate cancer and promotes prostate cancer cell proliferation and survival Taro Murata1,2 Ϯ, Ken-ichi Takayama1,3,4 Ϯ, Tomohiko Urano1,3,4, Tetsuya Fujimura2, Daisaku Ashikari1,5, Daisuke Obinata1,5, Kuniko Horie-Inoue4, Satoru Takahashi5, Yasuyoshi Ouchi3, Yukio Homma2, Satoshi Inoue1,3,4 Ϯ: These authors contributed equally to this work. 1 Department of Anti-Aging Medicine, Graduate School of Medicine, The University of Tokyo, Tokyo, Japan, 2 Department of Urology, Graduate School of Medicine, The University of Tokyo, Tokyo, Japan, 3 Department of Geriatric Medicine, Graduate School of Medicine, The University of Tokyo, Tokyo, Japan, 4 Division of Gene Regulation and Signal Transduction, Research Center for Genomic Medicine, Saitama Medical University, Saitama, Japan, 5 Department of Urology, School of Medicine, Nihon University, Tokyo, Japan Running Title: Role of 14-3-3ζ in prostate cancer Key Words: Androgen; prostate cancer; 14-3-3 protein; androgen receptor Corresponding Author: Satoshi Inoue, Department of Anti-Aging Medicine, Graduate School of Medicine, University of Tokyo, Bunkyo-ku, Tokyo 113-8655, Japan. Phone: 81-3-5800-8834; Fax: 81-3-5800-9126; E-mail: [email protected]. 1 Downloaded from clincancerres.aacrjournals.org on September 28, 2021. © 2012 American Association for Cancer Research. Author Manuscript Published OnlineFirst on August 17, 2012; DOI: 10.1158/1078-0432.CCR-12-0281 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. -

The Utility of Resolving Asthma Molecular Signatures Using Tissue-Specific Transcriptome Data 11 12 13 Debajyoti Ghosh1, Lili Ding2, Jonathan A
G3: Genes|Genomes|Genetics Early Online, published on September 8, 2020 as doi:10.1534/g3.120.401718 1 2 3 4 5 6 7 8 9 10 The utility of resolving asthma molecular signatures using tissue-specific transcriptome data 11 12 13 Debajyoti Ghosh1, Lili Ding2, Jonathan A. Bernstein1, and Tesfaye B. Mersha3* 14 15 1Immunology and Allergy, Department of Internal Medicine, University of Cincinnati, Cincinnati, OH, 16 United States of America 17 2Division of Biostatistics and Epidemiology, Cincinnati Children’s Hospital Medical Center, Department 18 of Pediatrics, University of Cincinnati, Cincinnati, OH, United States of America 19 3Division of Asthma Research, Cincinnati Children’s Hospital Medical Center, Department of Pediatrics, 20 University of Cincinnati, Cincinnati, OH, United States of America 21 22 23 24 25 Corresponding author: 26 27 Tesfaye B. Mersha, Ph.D. 28 Associate Professor 29 Cincinnati Children's Hospital Medical Center 30 Department of Pediatrics 31 University of Cincinnati 32 3333 Burnet Avenue 33 MLC 7037 34 Cincinnati, OH 45229-3026 35 Phone: (513) 803-2766 36 Fax: (513) 636-1657 37 Email: [email protected] 38 39 40 41 1 © The Author(s) 2020. Published by the Genetics Society of America. 42 Graphical Abstract 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 2 61 ABSTRACT 62 An integrative analysis focused on multi-tissue transcriptomics has not been done for asthma. Tissue- 63 specific DEGs remain undetected in many multi-tissue analyses, which influences identification of 64 disease-relevant pathways and potential drug candidates. -

1 Novel Expression Signatures Identified by Transcriptional Analysis
ARD Online First, published on October 7, 2009 as 10.1136/ard.2009.108043 Ann Rheum Dis: first published as 10.1136/ard.2009.108043 on 7 October 2009. Downloaded from Novel expression signatures identified by transcriptional analysis of separated leukocyte subsets in SLE and vasculitis 1Paul A Lyons, 1Eoin F McKinney, 1Tim F Rayner, 1Alexander Hatton, 1Hayley B Woffendin, 1Maria Koukoulaki, 2Thomas C Freeman, 1David RW Jayne, 1Afzal N Chaudhry, and 1Kenneth GC Smith. 1Cambridge Institute for Medical Research and Department of Medicine, Addenbrooke’s Hospital, Hills Road, Cambridge, CB2 0XY, UK 2Roslin Institute, University of Edinburgh, Roslin, Midlothian, EH25 9PS, UK Correspondence should be addressed to Dr Paul Lyons or Prof Kenneth Smith, Department of Medicine, Cambridge Institute for Medical Research, Addenbrooke’s Hospital, Hills Road, Cambridge, CB2 0XY, UK. Telephone: +44 1223 762642, Fax: +44 1223 762640, E-mail: [email protected] or [email protected] Key words: Gene expression, autoimmune disease, SLE, vasculitis Word count: 2,906 The Corresponding Author has the right to grant on behalf of all authors and does grant on behalf of all authors, an exclusive licence (or non-exclusive for government employees) on a worldwide basis to the BMJ Publishing Group Ltd and its Licensees to permit this article (if accepted) to be published in Annals of the Rheumatic Diseases and any other BMJPGL products to exploit all subsidiary rights, as set out in their licence (http://ard.bmj.com/ifora/licence.pdf). http://ard.bmj.com/ on September 29, 2021 by guest. Protected copyright. 1 Copyright Article author (or their employer) 2009. -

Construction and Analysis of Protein-Protein Interaction Network of Non-Alcoholic Fatty Liver Disease
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.01.406215; this version posted December 9, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Construction and Analysis of Protein-Protein Interaction Network of Non-Alcoholic Fatty Liver Disease Athina I. Amanatidou and George V. Dedoussis Department of Nutrition and Dietetics, School of Health Science and Education, Harokopio University, El. Venizelou 70, 17671, Athens, Greece Correspondence to: G. V. Dedoussis, A. I. Amanatidou, Department of Nutrition and Dietetics, School of Health Science and Education, Harokopio University, El. Venizelou 70, 17671, Athens, Greece E-mail addresses: [email protected] (G.V. Dedoussis), [email protected] (A. I. Amanatidou) Telephone: +302109549179 (G.V. Dedoussis), +306949293472 (A. I. Amanatidou) 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.12.01.406215; this version posted December 9, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Non-alcoholic fatty liver disease (NAFLD) is a disease with multidimensional complexities. Many attempts have been made over the years to treat this disease but its incidence is rising. For this reason, the need to identify and study new candidate proteins that may be associated with NAFLD is of utmost importance. Systems-based approaches such as the analysis of protein-protein interaction (PPI) network could lead to the discovery of new proteins associated with a disease that can then be translated into clinical practice. -

Overexpression of YWHAZ Relates to Tumor Cell Proliferation and Malignant Outcome of Gastric Carcinoma
FULL PAPER British Journal of Cancer (2013) 108, 1324–1331 | doi: 10.1038/bjc.2013.65 Keywords: YWHAZ (14-3-3z); gastric carcinoma; malignant outcome; prognostic factor Overexpression of YWHAZ relates to tumor cell proliferation and malignant outcome of gastric carcinoma Y Nishimura1,3, S Komatsu1,3, D Ichikawa1, H Nagata1, S Hirajima1, H Takeshita1, T Kawaguchi1, T Arita1, H Konishi1, K Kashimoto1, A Shiozaki1, H Fujiwara1, K Okamoto1, H Tsuda2 and E Otsuji1 1Division of Digestive Surgery, Department of Surgery, Kyoto Prefectural University of Medicine, 465 Kajii-cho, Kawaramachihirokoji, Kamigyo-ku, Kyoto 602-8566, Japan and 2Department of Pathology, National Cancer Center Hospital, Tokyo, Japan Background: Several studies have demonstrated that YWHAZ (14-3-3z), included in the 14-3-3 family of proteins, has been implicated in the initiation and progression of cancers. We tested whether YWHAZ acted as a cancer-promoting gene through its activation/overexpression in gastric cancer (GC). Methods: We analysed 7 GC cell lines and 141 primary tumours, which were curatively resected in our hospital between 2001 and 2003. Results: Overexpression of the YWHAZ protein was frequently detected in GC cell lines (six out of seven lines, 85.7%) and primary tumour samples of GC (72 out of 141 cases, 51%), and significantly correlated with larger tumour size, venous and lymphatic invasion, deeper tumour depth, and higher pathological stage and recurrence rate. Patients with YWHAZ-overexpressing tumours had worse overall survival rates than those with non-expressing tumours in both intensity and proportion expression-dependent manner. YWHAZ positivity was independently associated with a worse outcome in multivariate analysis (P ¼ 0.0491, hazard ratio 2.3 (1.003–5.304)). -

Signal-Oriented Pathway Analyses Reveal a Signaling Complex As A
Published OnlineFirst October 10, 2016; DOI: 10.1158/0008-5472.CAN-16-1740 Cancer Integrated Systems and Technologies Research Signal-Oriented Pathway Analyses Reveal a Signaling Complex as a Synthetic Lethal Target for p53 Mutations Songjian Lu1,2, Chunhui Cai1,2, Gonghong Yan3,4,5, Zhuan Zhou3,6, Yong Wan3,6, Vicky Chen1,2, Lujia Chen1,2, Gregory F. Cooper1,2, Lina M. Obeid7, Yusuf A. Hannun7, Adrian V. Lee2,3,4,5, and Xinghua Lu1,2 Abstract Defining processes that are synthetic lethal with p53 muta- methods-identified pathways were perturbed by SGA. In par- tions in cancer cells may reveal possible therapeutic strategies. ticular, our analyses revealed that SGA affecting TP53, PTK2, In this study, we report the development of a signal-oriented YWHAZ,andMED1 perturbed a set of signals that promote cell computational framework for cancer pathway discovery in this proliferation, anchor-free colony formation, and epithelial– context. We applied our bipartite graph–based functional mod- mesenchymal transition (EMT). These proteins formed a sig- ule discovery algorithm to identify transcriptomic modules naling complex that mediates these oncogenic processes in a abnormally expressed in multiple tumors, such that the genes coordinated fashion. Disruption of this signaling complex by in a module were likely regulated by a common, perturbed knocking down PTK2, YWHAZ, or MED1 attenuated and signal. For each transcriptomic module, we applied our weight- reversed oncogenic phenotypes caused by mutant p53 in a ed k-path merge algorithm to search for a set of somatic genome synthetic lethal manner. This signal-oriented framework for alterations (SGA) that likely perturbed the signal, that is, the searching pathways and therapeutic targets is applicable to all candidate members of the pathway that regulate the transcrip- cancer types, thus potentially impacting precision medicine in tomic module. -

MLL-AF4 Binds Directly to a BCL-2 Specific Enhancer
Experimental Hematology 2016;-:-–- MLL-AF4 binds directly to a BCL-2 specific enhancer and modulates H3K27 acetylation Laura Godfreya, Jon Kerrya, Ross Thornea, Emmanouela Repapib, James O.J. Daviesa, Marta Tapiaa, Erica Ballabioa, Jim R. Hughesa, Huimin Gengc, Marina Konoplevad, and Thomas A. Milnea aWeatherall Institute of Molecular Medicine, MRC Molecular Haematology Unit, University of Oxford, Headington, Oxford, UK; bWeatherall Institute of Molecular Medicine, Computational Biology Research Group, University of Oxford, Headington, Oxford, UK; cDepartment of Laboratory Medicine, University of California San Francisco, San Francisco, CA; dDepartment of Leukemia, University of Texas MD Anderson Cancer Center, Houston, TX (Received 27 July 2016; revised 21 October 2016; accepted 2 November 2016) Survival rates for children and adults carrying mutations in the Mixed Lineage Leukemia (MLL) gene continue to have a very poor prognosis. The most common MLL mutation in acute lympho- blastic leukemia is the t(4;11)(q21;q23) chromosome translocation that fuses MLL in-frame with the AF4 gene producing MLL-AF4 and AF4-MLL fusion proteins. Previously, we found that MLL-AF4 binds to the BCL-2 gene and directly activates it through DOT1L recruitment and increased H3K79me2/3 levels. In the study described here, we performed a detailed analysis of MLL-AF4 regulation of the entire BCL-2 family. By measuring nascent RNA production in MLL-AF4 knockdowns, we found that of all the BCL-2 family genes, MLL-AF4 directly con- trols the active transcription of both BCL-2 and MCL-1 and also represses BIM via binding of the polycomb group repressor 1 (PRC1) complex component CBX8. -

Proteomic Analysis of Necroptotic Extracellular Vesicles
bioRxiv preprint doi: https://doi.org/10.1101/2020.04.11.037192; this version posted April 12, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Proteomic analysis of necroptotic extracellular vesicles Inbar Shlomovitz1, Gali Yanovich-Arad2, Ziv Erlich1, Liat Edry-Botzer1, Sefi Zargarian1, Hadar Cohen1, Yifat Ofir-Birin3, Neta Regev-Rudzki3 and Motti Gerlic1 1. Department of Clinical Microbiology and Immunology, Sackler School of Medicine, Tel Aviv University, Tel Aviv, Israel 2. Department of Human Molecular Genetics and Biochemistry, Sackler School of Medicine, Tel Aviv University, Tel Aviv, Israel 3. Department of Biomolecular Sciences, Weizmann Institute of Science, Rehovot, Israel Corresponding author Motti Gerlic, Department of Clinical Microbiology and Immunology, Sackler School of Medicine, Tel Aviv University, Tel Aviv 69978, Israel Tel: +972-36409069, E-mail: [email protected] Abstract Necroptosis is a regulated and inflammatory form of cell death. We, and others, have previously reported that necroptotic cells release extracellular vesicles (EVs). We have found that necroptotic EVs are loaded with proteins, including the phosphorylated form of the key necroptosis-executing factor, mixed lineage kinase domain-like kinase (MLKL). However, neither the exact protein composition, nor the impact, of necroptotic EVs have been delineated. To characterize their content, EVs from necroptotic and untreated U937 cells were isolated and analyzed by mass spectrometry-based proteomics. A total of 3337 proteins were identified, sharing a high degree of similarity with exosome proteome databases, and clearly distinguishing necroptotic and control EVs.