The Utility of Resolving Asthma Molecular Signatures Using Tissue-Specific Transcriptome Data 11 12 13 Debajyoti Ghosh1, Lili Ding2, Jonathan A
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

TLR3-Dependent Activation of TLR2 Endogenous Ligands Via the Myd88 Signaling Pathway Augments the Innate Immune Response
cells Article TLR3-Dependent Activation of TLR2 Endogenous Ligands via the MyD88 Signaling Pathway Augments the Innate Immune Response 1 2, 1 3 Hellen S. Teixeira , Jiawei Zhao y, Ethan Kazmierski , Denis F. Kinane and Manjunatha R. Benakanakere 2,* 1 Department of Orthodontics, School of Dental Medicine, University of Pennsylvania, Philadelphia, PA 19004, USA; [email protected] (H.S.T.); [email protected] (E.K.) 2 Department of Periodontics, School of Dental Medicine, University of Pennsylvania, Philadelphia, PA 19004, USA; [email protected] 3 Periodontology Department, Bern Dental School, University of Bern, 3012 Bern, Switzerland; [email protected] * Correspondence: [email protected] Present address: Department of Pathology, Wayne State University School of Medicine, y 541 East Canfield Ave., Scott Hall 9215, Detroit, MI 48201, USA. Received: 30 June 2020; Accepted: 12 August 2020; Published: 17 August 2020 Abstract: The role of the adaptor molecule MyD88 is thought to be independent of Toll-like receptor 3 (TLR3) signaling. In this report, we demonstrate a previously unknown role of MyD88 in TLR3 signaling in inducing endogenous ligands of TLR2 to elicit innate immune responses. Of the various TLR ligands examined, the TLR3-specific ligand polyinosinic:polycytidylic acid (poly I:C), significantly induced TNF production and the upregulation of other TLR transcripts, in particular, TLR2. Accordingly, TLR3 stimulation also led to a significant upregulation of endogenous TLR2 ligands mainly, HMGB1 and Hsp60. By contrast, the silencing of TLR3 significantly downregulated MyD88 and TLR2 gene expression and pro-inflammatory IL1β, TNF, and IL8 secretion. The silencing of MyD88 similarly led to the downregulation of TLR2, IL1β, TNF and IL8, thus suggesting MyD88 / to somehow act downstream of TLR3. -

RT² Profiler PCR Array (384-Well Format) Human Apoptosis 384HT
RT² Profiler PCR Array (384-Well Format) Human Apoptosis 384HT Cat. no. 330231 PAHS-3012ZE For pathway expression analysis Format For use with the following real-time cyclers RT² Profiler PCR Array, Applied Biosystems® models 7900HT (384-well block), Format E ViiA™ 7 (384-well block); Bio-Rad CFX384™ RT² Profiler PCR Array, Roche® LightCycler® 480 (384-well block) Format G Description The Human Apoptosis RT² Profiler PCR Array profiles the expression of 370 key genes involved in apoptosis, or programmed cell death. The array includes the TNF ligands and their receptors; members of the bcl-2 family, BIR (baculoviral IAP repeat) domain proteins, CARD domain (caspase recruitment domain) proteins, death domain proteins, TRAF (TNF receptor-associated factor) domain proteins and caspases. These 370 genes are also grouped by their functional contribution to apoptosis, either anti-apoptosis or induction of apoptosis. Using real-time PCR, you can easily and reliably analyze expression of a focused panel of genes related to apoptosis with this array. For further details, consult the RT² Profiler PCR Array Handbook. Sample & Assay Technologies Shipping and storage RT² Profiler PCR Arrays in formats E and G are shipped at ambient temperature, on dry ice, or blue ice packs depending on destination and accompanying products. For long term storage, keep plates at –20°C. Note: Ensure that you have the correct RT² Profiler PCR Array format for your real-time cycler (see table above). Note: Open the package and store the products appropriately immediately -

MYD88 L265P Is a Marker Highly Characteristic Of, but Not Restricted To, Waldenstro¨M’S Macroglobulinemia
Leukemia (2013) 27, 1722–1728 & 2013 Macmillan Publishers Limited All rights reserved 0887-6924/13 www.nature.com/leu ORIGINAL ARTICLE MYD88 L265P is a marker highly characteristic of, but not restricted to, Waldenstro¨m’s macroglobulinemia C Jime´ nez1, E Sebastia´n1, MC Chillo´n1,2, P Giraldo3, J Mariano Herna´ndez4, F Escalante5, TJ Gonza´lez-Lo´ pez6, C Aguilera7, AG de Coca8, I Murillo3, M Alcoceba1, A Balanzategui1, ME Sarasquete1,2, R Corral1, LA Marı´n1, B Paiva1,2, EM Ocio1,2, NC Gutie´ rrez1,2, M Gonza´lez1,2, JF San Miguel1,2 and R Garcı´a-Sanz1,2 We evaluated the MYD88 L265P mutation in Waldenstro¨m’s macroglobulinemia (WM) and B-cell lymphoproliferative disorders by specific polymerase chain reaction (PCR) (sensitivity B10 À 3). No mutation was seen in normal donors, while it was present in 101/117 (86%) WM patients, 27/31 (87%) IgM monoclonal gammapathies of uncertain significance (MGUS), 3/14 (21%) splenic marginal zone lymphomas and 9/48 (19%) non-germinal center (GC) diffuse large B-cell lymphomas (DLBCLs). The mutation was absent in all 28 GC-DLBCLs, 13 DLBCLs not subclassified, 35 hairy cell leukemias, 39 chronic lymphocyticleukemias(16withM-component), 25 IgA or IgG-MGUS, 24 multiple myeloma (3 with an IgM isotype), 6 amyloidosis, 9 lymphoplasmacytic lymphomas and 1 IgM-related neuropathy. Among WM and IgM- MGUS, MYD88 L265P mutation was associated with some differences in clinical and biological characteristics, although usually minor; wild-type MYD88 cases had smaller M-component (1.77 vs 2.72 g/dl, P ¼ 0.022), more lymphocytosis (24 vs 5%, P ¼ 0.006), higher lactate dehydrogenase level (371 vs 265 UI/L, P ¼ 0.002), atypical immunophenotype (CD23 À CD27 þþFMC7 þþ), less Immunoglobulin Heavy Chain Variable gene (IGHV) somatic hypermutation (57 vs 97%, P ¼ 0.012) and less IGHV3–23 gene selection (9 vs 27%, P ¼ 0.014). -

MYD88 and Beyond: Novel Opportunities for Diagnosis, Prognosis and Treatment in Waldenstro¨M’S Macroglobulinemia
Leukemia (2014) 28, 1799–1803 & 2014 Macmillan Publishers Limited All rights reserved 0887-6924/14 www.nature.com/leu CONCISE REVIEW MYD88 and beyond: novel opportunities for diagnosis, prognosis and treatment in Waldenstro¨m’s Macroglobulinemia O Landgren and N Tageja Waldenstro¨m’s Macroglobulinemia (WM) is a rare disease of the elderly with a median age of 63–68 years at diagnosis. Despite recent progress in biological insights and therapeutics, WM remains clinically challenging to diagnose and is difficult to manage with significant morbidity and lack of established curative therapies. Recently, the use of whole-genome sequencing has helped to identify a highly recurrent somatic mutation, myeloid differentiation factor 88 [MYD88] L265P in WM. This has fueled major interest in the field and as newer evidence accumulates, it is clear that that discovery of MYD88 L265P mutation may represent an important breakthrough in understanding the pathogenesis of WM and lymphoproliferative disorders. Recent scientific work in this field has also guided the identification of new targets such as CXCR4 and PI3K-delta that may have major implications in the future treatment of WM. This review discusses the role of MYD88 L265P mutations as well as targets beyond MYD88 in the setting of pathogenesis and development of future rational therapeutic trials focusing on patients diagnosed with WM. Leukemia (2014) 28, 1799–1803; doi:10.1038/leu.2014.88 INTRODUCTION transduces signals to the NF-kB transcription factors in response to Waldenstro¨m’s Macroglobulinemia (WM) is a rare hematological IL-1R1 signaling. MYD88 has a modular structure with a Toll/IL-1R malignancy with a reported age-adjusted incidence rate of 3.4 per (TIR) domain at its COOH terminus and a death domain at its 12 11 million among men and 1.7 per million among women in the NH2 terminus. -

And Sepsis-Induced Lung Inflammation and Mediates Myd88
The Journal of Immunology Caveolin-1 Tyr14 Phosphorylation Induces Interaction with TLR4 in Endothelial Cells and Mediates MyD88-Dependent Signaling and Sepsis-Induced Lung Inflammation Hao Jiao,*,†,1 Yang Zhang,*,†,1 Zhibo Yan,* Zhen-Guo Wang,* Gongjian Liu,† Richard D. Minshall,*,‡ Asrar B. Malik,‡ and Guochang Hu*,‡ Activation of TLR4 by the endotoxin LPS is a critical event in the pathogenesis of Gram-negative sepsis. Caveolin-1, the signaling protein associated with caveolae, is implicated in regulating the lung inflammatory response to LPS; however, the mechanism is not understood. In this study, we investigated the role of caveolin-1 in regulating TLR4 signaling in endothelial cells. We observed that LPS interaction with CD14 in endothelial cells induced Src-dependent caveolin-1 phosphorylation at Tyr14. Using a TLR4-MD2- CD14–transfected HEK-293 cell line and caveolin-1–deficient (cav-12/2) mouse lung microvascular endothelial cells, we demon- strated that caveolin-1 phosphorylation at Tyr14 following LPS exposure induced caveolin-1 and TLR4 interaction and, thereby, TLR4 activation of MyD88, leading to NF-kB activation and generation of proinflammatory cytokines. Exogenous expression of phosphorylation-deficient Y14F caveolin-1 mutant in cav-12/2 mouse pulmonary vasculature rendered the mice resistant to LPS compared with reintroduction of wild-type caveolin-1. Thus, caveolin-1 Y14 phosphorylation was required for the interaction with TLR4 and activation of TLR4-MyD88 signaling and sepsis-induced lung inflammation. Inhibiting caveolin-1 Tyr14 phosphoryla- tion and resultant inactivation of TLR4 signaling in pulmonary vascular endothelial cells represent a novel strategy for preventing sepsis-induced lung inflammation and injury. -

TLR Signaling Pathways
Seminars in Immunology 16 (2004) 3–9 TLR signaling pathways Kiyoshi Takeda, Shizuo Akira∗ Department of Host Defense, Research Institute for Microbial Diseases, Osaka University, and ERATO, Japan Science and Technology Corporation, 3-1 Yamada-oka, Suita, Osaka 565-0871, Japan Abstract Toll-like receptors (TLRs) have been established to play an essential role in the activation of innate immunity by recognizing spe- cific patterns of microbial components. TLR signaling pathways arise from intracytoplasmic TIR domains, which are conserved among all TLRs. Recent accumulating evidence has demonstrated that TIR domain-containing adaptors, such as MyD88, TIRAP, and TRIF, modulate TLR signaling pathways. MyD88 is essential for the induction of inflammatory cytokines triggered by all TLRs. TIRAP is specifically involved in the MyD88-dependent pathway via TLR2 and TLR4, whereas TRIF is implicated in the TLR3- and TLR4-mediated MyD88-independent pathway. Thus, TIR domain-containing adaptors provide specificity of TLR signaling. © 2003 Elsevier Ltd. All rights reserved. Keywords: TLR; Innate immunity; Signal transduction; TIR domain 1. Introduction 2. Toll-like receptors Toll receptor was originally identified in Drosophila as an A mammalian homologue of Drosophila Toll receptor essential receptor for the establishment of the dorso-ventral (now termed TLR4) was shown to induce the expression pattern in developing embryos [1]. In 1996, Hoffmann and of genes involved in inflammatory responses [3]. In addi- colleagues demonstrated that Toll-mutant flies were highly tion, a mutation in the Tlr4 gene was identified in mouse susceptible to fungal infection [2]. This study made us strains that were hyporesponsive to lipopolysaccharide [4]. aware that the immune system, particularly the innate im- Since then, Toll receptors in mammals have been a major mune system, has a skilful means of detecting invasion by focus in the immunology field. -

A Synthetic Cd8a:Myd88 Coreceptor Enhances
Published OnlineFirst October 20, 2017; DOI: 10.1158/0008-5472.CAN-17-0653 Cancer Microenvironment and Immunology Research A Synthetic CD8a:MyD88 Coreceptor Enhances CD8þ T-cell Responses to Weakly Immunogenic and Lowly Expressed Tumor Antigens Sabina Kaczanowska1, Ann Mary Joseph1, Jitao Guo1, Alexander K Tsai1, Jackline Joy Lasola1, Kenisha Younger1, Yuji Zhang1,2, Cruz Velasco Gonzales3, and Eduardo Davila1,4 Abstract Tcell–based immunotherapies are a promising approach for and Toll-like receptor signaling-related proteins. CD8a: patients with advanced cancers. However, various obstacles MyD88–expressing T cells improved antitumor responses in limit T-cell efficacy, including suboptimal T-cell receptor mice. Enhanced antitumor activity was associated with a (TCR) activation and an immunosuppressive tumor environ- unique tumor cytokine/chemokine signature, improved T-cell ment. Here, we developed a fusion protein by linking CD8a infiltration, reduced markers of T-cell exhaustion, elevated þ and MyD88 (CD8a:MyD88) to enhance CD8 T-cell levels of proteins associated with antigen presentation, and responses to weakly immunogenic and poorly expressed fewer macrophages with an immunosuppressive phenotype in tumor antigens. CD8a:MyD88–engineered T cells exhibited tumors. Given these observations, CD8a:MyD88 represents a increased proliferation and expression of effector and costi- unique and versatile approach to help overcome immunosup- mulatory molecules in a tumor antigen–dependent manner. pression and enhance T-cell responses to tumor antigens. These effects were accompanied by elevated activation of TCR Cancer Res; 77(24); 7049–58. Ó2017 AACR. Introduction ciated macrophage (6) and Th2 cytokine accumulation hamper antitumor T-cell responses. Moreover, chronic exposure to factors T cell–based immunotherapies are one of the most promising in the TME induces the expression of receptors that foster T-cell treatments for patients with advanced cancers, including melano- exhaustion, such as Tim-3, Lag-3, and PD-1 (7). -

Induced IFN Regulatory Factor 1 Transcription Factor by Myd88 in Toll-Like Receptor-Dependent Gene Induction Program
Evidence for licensing of IFN-␥-induced IFN regulatory factor 1 transcription factor by MyD88 in Toll-like receptor-dependent gene induction program Hideo Negishi*, Yasuyuki Fujita*, Hideyuki Yanai*, Shinya Sakaguchi*, Xinshou Ouyang*, Masahiro Shinohara†, Hiroshi Takayanagi†, Yusuke Ohba*, Tadatsugu Taniguchi*‡, and Kenya Honda* *Department of Immunology, Graduate School of Medicine and Faculty of Medicine, University of Tokyo, Hongo 7-3-1, Bunkyo-ku, Tokyo 113-0033, Japan; and †Department of Cell Signaling, Graduate School, Tokyo Medical and Dental University, Yushima 1-5-45, Bunkyo-ku, Tokyo 113-8549, Japan Contributed by Tadatsugu Taniguchi, August 18, 2006 The recognition of microbial components by Toll-like receptors (TLRs) In the present study we investigated how IFN-␥-induced IRF1 initiates signal transduction pathways, which trigger the expression contributes to TLR-mediated signaling. We demonstrate that of a series of target genes. It has been reported that TLR signaling is IRF1 forms a complex with MyD88, similar to the case of IRF4, enhanced by cytokines such as IFN-␥, but the mechanisms underlying IRF5, and IRF7. We also provide evidence that IRF1 induced this enhancement remain unclear. The MyD88 adaptor, which is by IFN-␥ is activated by MyD88, which we refer to as ‘‘licensing,’’ essential for signaling by many TLRs, recruits members of the IFN and migrates rapidly into the nucleus to mediate an efficient regulatory factor (IRF) family of transcription factors, such as IRF5 and induction of IFN-, iNOS, and IL-12p35. Our study therefore IRF7, to evoke the activation of TLR target genes. In this study we revealed that IRF1 is a previously unidentified member of the demonstrate that IRF1, which is induced by IFN-␥, also interacts with multimolecular complex organized via MyD88 and that the IRF1 and is activated by MyD88 upon TLR activation. -
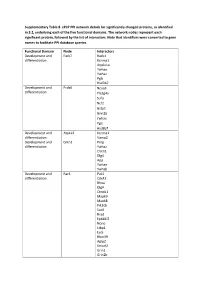
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4 -

Identification of Transcriptional Mechanisms Downstream of Nf1 Gene Defeciency in Malignant Peripheral Nerve Sheath Tumors Daochun Sun Wayne State University
Wayne State University DigitalCommons@WayneState Wayne State University Dissertations 1-1-2012 Identification of transcriptional mechanisms downstream of nf1 gene defeciency in malignant peripheral nerve sheath tumors Daochun Sun Wayne State University, Follow this and additional works at: http://digitalcommons.wayne.edu/oa_dissertations Recommended Citation Sun, Daochun, "Identification of transcriptional mechanisms downstream of nf1 gene defeciency in malignant peripheral nerve sheath tumors" (2012). Wayne State University Dissertations. Paper 558. This Open Access Dissertation is brought to you for free and open access by DigitalCommons@WayneState. It has been accepted for inclusion in Wayne State University Dissertations by an authorized administrator of DigitalCommons@WayneState. IDENTIFICATION OF TRANSCRIPTIONAL MECHANISMS DOWNSTREAM OF NF1 GENE DEFECIENCY IN MALIGNANT PERIPHERAL NERVE SHEATH TUMORS by DAOCHUN SUN DISSERTATION Submitted to the Graduate School of Wayne State University, Detroit, Michigan in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY 2012 MAJOR: MOLECULAR BIOLOGY AND GENETICS Approved by: _______________________________________ Advisor Date _______________________________________ _______________________________________ _______________________________________ © COPYRIGHT BY DAOCHUN SUN 2012 All Rights Reserved DEDICATION This work is dedicated to my parents and my wife Ze Zheng for their continuous support and understanding during the years of my education. I could not achieve my goal without them. ii ACKNOWLEDGMENTS I would like to express tremendous appreciation to my mentor, Dr. Michael Tainsky. His guidance and encouragement throughout this project made this dissertation come true. I would also like to thank my committee members, Dr. Raymond Mattingly and Dr. John Reiners Jr. for their sustained attention to this project during the monthly NF1 group meetings and committee meetings, Dr. -

Characterization of Genes in the CFTR-Mediated Innate Immune Response
The University of Maine DigitalCommons@UMaine Honors College 5-2012 Characterization of Genes in the CFTR-Mediated Innate Immune Response Eric Peterman Follow this and additional works at: https://digitalcommons.library.umaine.edu/honors Part of the Cell and Developmental Biology Commons, and the Molecular Biology Commons Recommended Citation Peterman, Eric, "Characterization of Genes in the CFTR-Mediated Innate Immune Response" (2012). Honors College. 71. https://digitalcommons.library.umaine.edu/honors/71 This Honors Thesis is brought to you for free and open access by DigitalCommons@UMaine. It has been accepted for inclusion in Honors College by an authorized administrator of DigitalCommons@UMaine. For more information, please contact [email protected]. CHARACTERIZATION OF GENES IN THE CFTR-MEDIATED INNATE IMMUNE RESPONSE by Eric Peterman A Thesis Submitted in Partial Fulfillment of the Requirements for a Degree with Honors (Biochemistry, Molecular and Cellular Biology) The Honors College University of Maine May 2012 Advisory Committee: Carol Kim, Professor, Molecular & Biomedical Sciences Robert Gundersen, Associate Professor, Molecular & Biomedical Sciences John Singer, Professor, Molecular & Biomedical Sciences Julie Gosse, Assistant Professor, Molecular & Biomedical Sciences Keith Hutchison, Professor, Molecular & Biomedical Sciences Mark Haggerty, Lecturer, Rezendes Preceptor for Civic Engagement Abstract: Recently, the Kim Lab has shown that the cystic fibrosis transmembrane conductance regulator (cftr) gene is responsible for mediating resistance to Pseudomonas aeruginosa in a zebrafish infection model. Using the Gene Expression Omnibus, an NCBI functional genomics data repository, it was determined that Smad3, a transcription factor in the TGF-β signaling pathway, is upregulated in the presence of P. aeruginosa. It was found that in our zebrafish model, the Smad3 paralogs Smad3a and Smad3b are upregulated following microinjection of a cftr antisense morpholino oligomer. -

Evolution of Gremlin 2 in Cetartiodactyl Mammals: Gene Loss Coincides with Lack of Upper Jaw Incisors in Ruminants
Evolution of gremlin 2 in cetartiodactyl mammals: gene loss coincides with lack of upper jaw incisors in ruminants Juan C. Opazo1, Kattina Zavala1, Paola Krall2 and Rodrigo A. Arias3 1 Instituto de Ciencias Ambientales y Evolutivas, Universidad Austral de Chile, Valdivia, Chile 2 Unidad de Nefrología, Universidad Austral de Chile, Valdivia, Chile 3 Instituto de Producción Animal, Universidad Austral de Chile, Valdivia, Chile ABSTRACT Understanding the processes that give rise to genomic variability in extant species is an active area of research within evolutionary biology. With the availability of whole genome sequences, it is possible to quantify different forms of variability such as variation in gene copy number, which has been described as an important source of genetic variability and in consequence of phenotypic variability. Most of the research on this topic has been focused on understanding the biological significance of gene duplication, and less attention has been given to the evolutionary role of gene loss. Gremlin 2 is a member of the DAN gene family and plays a significant role in tooth development by blocking the ligand-signaling pathway of BMP2 and BMP4. The goal of this study was to investigate the evolutionary history of gremlin 2 in cetartiodactyl mammals, a group that possesses highly divergent teeth morphology. Results from our analyses indicate that gremlin 2 has experienced a mixture of gene loss, gene duplication, and rate acceleration. Although the last common ancestor of cetartiodactyls possessed a single gene copy, pigs and camels are the only cetartiodactyl groups that have retained gremlin 2. According to the phyletic distribution of this gene and synteny analyses, we propose that gremlin 2 was lost in the common ancestor of ruminants and cetaceans between 56.3 and 63.5 million years ago as a product of a chromosomal rearrangement.