A Novel Function of YWHAZ/B-Catenin Axis in Promoting Epithelial–Mesenchymal Transition and Lung Cancer Metastasis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

RT² Profiler PCR Array (384-Well Format) Human Apoptosis 384HT
RT² Profiler PCR Array (384-Well Format) Human Apoptosis 384HT Cat. no. 330231 PAHS-3012ZE For pathway expression analysis Format For use with the following real-time cyclers RT² Profiler PCR Array, Applied Biosystems® models 7900HT (384-well block), Format E ViiA™ 7 (384-well block); Bio-Rad CFX384™ RT² Profiler PCR Array, Roche® LightCycler® 480 (384-well block) Format G Description The Human Apoptosis RT² Profiler PCR Array profiles the expression of 370 key genes involved in apoptosis, or programmed cell death. The array includes the TNF ligands and their receptors; members of the bcl-2 family, BIR (baculoviral IAP repeat) domain proteins, CARD domain (caspase recruitment domain) proteins, death domain proteins, TRAF (TNF receptor-associated factor) domain proteins and caspases. These 370 genes are also grouped by their functional contribution to apoptosis, either anti-apoptosis or induction of apoptosis. Using real-time PCR, you can easily and reliably analyze expression of a focused panel of genes related to apoptosis with this array. For further details, consult the RT² Profiler PCR Array Handbook. Sample & Assay Technologies Shipping and storage RT² Profiler PCR Arrays in formats E and G are shipped at ambient temperature, on dry ice, or blue ice packs depending on destination and accompanying products. For long term storage, keep plates at –20°C. Note: Ensure that you have the correct RT² Profiler PCR Array format for your real-time cycler (see table above). Note: Open the package and store the products appropriately immediately -

Seq2pathway Vignette
seq2pathway Vignette Bin Wang, Xinan Holly Yang, Arjun Kinstlick May 19, 2021 Contents 1 Abstract 1 2 Package Installation 2 3 runseq2pathway 2 4 Two main functions 3 4.1 seq2gene . .3 4.1.1 seq2gene flowchart . .3 4.1.2 runseq2gene inputs/parameters . .5 4.1.3 runseq2gene outputs . .8 4.2 gene2pathway . 10 4.2.1 gene2pathway flowchart . 11 4.2.2 gene2pathway test inputs/parameters . 11 4.2.3 gene2pathway test outputs . 12 5 Examples 13 5.1 ChIP-seq data analysis . 13 5.1.1 Map ChIP-seq enriched peaks to genes using runseq2gene .................... 13 5.1.2 Discover enriched GO terms using gene2pathway_test with gene scores . 15 5.1.3 Discover enriched GO terms using Fisher's Exact test without gene scores . 17 5.1.4 Add description for genes . 20 5.2 RNA-seq data analysis . 20 6 R environment session 23 1 Abstract Seq2pathway is a novel computational tool to analyze functional gene-sets (including signaling pathways) using variable next-generation sequencing data[1]. Integral to this tool are the \seq2gene" and \gene2pathway" components in series that infer a quantitative pathway-level profile for each sample. The seq2gene function assigns phenotype-associated significance of genomic regions to gene-level scores, where the significance could be p-values of SNPs or point mutations, protein-binding affinity, or transcriptional expression level. The seq2gene function has the feasibility to assign non-exon regions to a range of neighboring genes besides the nearest one, thus facilitating the study of functional non-coding elements[2]. Then the gene2pathway summarizes gene-level measurements to pathway-level scores, comparing the quantity of significance for gene members within a pathway with those outside a pathway. -

Mir-451A Suppresses Cell Proliferation, Metastasis and EMT Via Targeting YWHAZ in Hepatocellular Carcinoma
European Review for Medical and Pharmacological Sciences 2019; 23: 5158-5167 MiR-451a suppresses cell proliferation, metastasis and EMT via targeting YWHAZ in hepatocellular carcinoma G.-Y. WEI1, M. HU2, L. ZHAO1, W.-S. GUO1 1Department of General Surgery, Henan Traditional Chinese Medicine Hospital, Zhengzhou, China 2Department of Infection Management, Henan Traditional Chinese Medicine Hospital, Zhengzhou, China Abstract. – OBJECTIVE: MicroRNAs (miR- lines. Moreover, miR-451a inhibited the prolif- NAs) have been identified to participate in eration, invasion and migration of HCC cells via the progression of hepatocellular carcinoma targeting YWHAZ. Our findings indicated that (HCC). However, the function of miR-451a in miR-451a could serve as a novel target for HCC HCC remains unknown. The aim of this study diagnosis and biological therapy. was to determine the function of miR-451a by construction of several experiments in HCC tis- Key Words: sues and cells. MiR-451a, Hepatocellular carcinoma (HCC), Prolifer- PATIENTS AND METHODS: The expression ation, Metastasis, YWHAZ. level of miR-451a in 69 paired of HCC and ad- jacent normal tissue samples was detected us- ing quantitative Real-time polymerase chain re- Introduction action (qRT-PCR). MiR-451a expression in HCC derived cell lines was detected as well. By trans- fecting with miR-451a mimics or inhibitor, the Hepatocellular carcinoma (HCC) accounts expression level of miR-451a in HepG2 or Huh-7 for 85-90% of primary liver cancer. HCC is the cells was up- or down-regulated. MTT (3-(4,5-di- fifth most common morbidity and third most methylthiazol-2-yl)-2,5-diphenyl tetrazolium bro- common mortality cancer worldwide1. -

Supplemental Information
Supplemental information Dissection of the genomic structure of the miR-183/96/182 gene. Previously, we showed that the miR-183/96/182 cluster is an intergenic miRNA cluster, located in a ~60-kb interval between the genes encoding nuclear respiratory factor-1 (Nrf1) and ubiquitin-conjugating enzyme E2H (Ube2h) on mouse chr6qA3.3 (1). To start to uncover the genomic structure of the miR- 183/96/182 gene, we first studied genomic features around miR-183/96/182 in the UCSC genome browser (http://genome.UCSC.edu/), and identified two CpG islands 3.4-6.5 kb 5’ of pre-miR-183, the most 5’ miRNA of the cluster (Fig. 1A; Fig. S1 and Seq. S1). A cDNA clone, AK044220, located at 3.2-4.6 kb 5’ to pre-miR-183, encompasses the second CpG island (Fig. 1A; Fig. S1). We hypothesized that this cDNA clone was derived from 5’ exon(s) of the primary transcript of the miR-183/96/182 gene, as CpG islands are often associated with promoters (2). Supporting this hypothesis, multiple expressed sequences detected by gene-trap clones, including clone D016D06 (3, 4), were co-localized with the cDNA clone AK044220 (Fig. 1A; Fig. S1). Clone D016D06, deposited by the German GeneTrap Consortium (GGTC) (http://tikus.gsf.de) (3, 4), was derived from insertion of a retroviral construct, rFlpROSAβgeo in 129S2 ES cells (Fig. 1A and C). The rFlpROSAβgeo construct carries a promoterless reporter gene, the β−geo cassette - an in-frame fusion of the β-galactosidase and neomycin resistance (Neor) gene (5), with a splicing acceptor (SA) immediately upstream, and a polyA signal downstream of the β−geo cassette (Fig. -

Downloaded the “Top Edge” Version
bioRxiv preprint doi: https://doi.org/10.1101/855338; this version posted December 6, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Drosophila models of pathogenic copy-number variant genes show global and 2 non-neuronal defects during development 3 Short title: Non-neuronal defects of fly homologs of CNV genes 4 Tanzeen Yusuff1,4, Matthew Jensen1,4, Sneha Yennawar1,4, Lucilla Pizzo1, Siddharth 5 Karthikeyan1, Dagny J. Gould1, Avik Sarker1, Yurika Matsui1,2, Janani Iyer1, Zhi-Chun Lai1,2, 6 and Santhosh Girirajan1,3* 7 8 1. Department of Biochemistry and Molecular Biology, Pennsylvania State University, 9 University Park, PA 16802 10 2. Department of Biology, Pennsylvania State University, University Park, PA 16802 11 3. Department of Anthropology, Pennsylvania State University, University Park, PA 16802 12 4 contributed equally to work 13 14 *Correspondence: 15 Santhosh Girirajan, MBBS, PhD 16 205A Life Sciences Building 17 Pennsylvania State University 18 University Park, PA 16802 19 E-mail: [email protected] 20 Phone: 814-865-0674 21 1 bioRxiv preprint doi: https://doi.org/10.1101/855338; this version posted December 6, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 22 ABSTRACT 23 While rare pathogenic copy-number variants (CNVs) are associated with both neuronal and non- 24 neuronal phenotypes, functional studies evaluating these regions have focused on the molecular 25 basis of neuronal defects. -
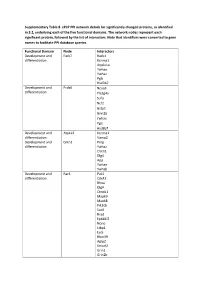
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4 -

Characterization of Genes in the CFTR-Mediated Innate Immune Response
The University of Maine DigitalCommons@UMaine Honors College 5-2012 Characterization of Genes in the CFTR-Mediated Innate Immune Response Eric Peterman Follow this and additional works at: https://digitalcommons.library.umaine.edu/honors Part of the Cell and Developmental Biology Commons, and the Molecular Biology Commons Recommended Citation Peterman, Eric, "Characterization of Genes in the CFTR-Mediated Innate Immune Response" (2012). Honors College. 71. https://digitalcommons.library.umaine.edu/honors/71 This Honors Thesis is brought to you for free and open access by DigitalCommons@UMaine. It has been accepted for inclusion in Honors College by an authorized administrator of DigitalCommons@UMaine. For more information, please contact [email protected]. CHARACTERIZATION OF GENES IN THE CFTR-MEDIATED INNATE IMMUNE RESPONSE by Eric Peterman A Thesis Submitted in Partial Fulfillment of the Requirements for a Degree with Honors (Biochemistry, Molecular and Cellular Biology) The Honors College University of Maine May 2012 Advisory Committee: Carol Kim, Professor, Molecular & Biomedical Sciences Robert Gundersen, Associate Professor, Molecular & Biomedical Sciences John Singer, Professor, Molecular & Biomedical Sciences Julie Gosse, Assistant Professor, Molecular & Biomedical Sciences Keith Hutchison, Professor, Molecular & Biomedical Sciences Mark Haggerty, Lecturer, Rezendes Preceptor for Civic Engagement Abstract: Recently, the Kim Lab has shown that the cystic fibrosis transmembrane conductance regulator (cftr) gene is responsible for mediating resistance to Pseudomonas aeruginosa in a zebrafish infection model. Using the Gene Expression Omnibus, an NCBI functional genomics data repository, it was determined that Smad3, a transcription factor in the TGF-β signaling pathway, is upregulated in the presence of P. aeruginosa. It was found that in our zebrafish model, the Smad3 paralogs Smad3a and Smad3b are upregulated following microinjection of a cftr antisense morpholino oligomer. -

Evolution of Gremlin 2 in Cetartiodactyl Mammals: Gene Loss Coincides with Lack of Upper Jaw Incisors in Ruminants
Evolution of gremlin 2 in cetartiodactyl mammals: gene loss coincides with lack of upper jaw incisors in ruminants Juan C. Opazo1, Kattina Zavala1, Paola Krall2 and Rodrigo A. Arias3 1 Instituto de Ciencias Ambientales y Evolutivas, Universidad Austral de Chile, Valdivia, Chile 2 Unidad de Nefrología, Universidad Austral de Chile, Valdivia, Chile 3 Instituto de Producción Animal, Universidad Austral de Chile, Valdivia, Chile ABSTRACT Understanding the processes that give rise to genomic variability in extant species is an active area of research within evolutionary biology. With the availability of whole genome sequences, it is possible to quantify different forms of variability such as variation in gene copy number, which has been described as an important source of genetic variability and in consequence of phenotypic variability. Most of the research on this topic has been focused on understanding the biological significance of gene duplication, and less attention has been given to the evolutionary role of gene loss. Gremlin 2 is a member of the DAN gene family and plays a significant role in tooth development by blocking the ligand-signaling pathway of BMP2 and BMP4. The goal of this study was to investigate the evolutionary history of gremlin 2 in cetartiodactyl mammals, a group that possesses highly divergent teeth morphology. Results from our analyses indicate that gremlin 2 has experienced a mixture of gene loss, gene duplication, and rate acceleration. Although the last common ancestor of cetartiodactyls possessed a single gene copy, pigs and camels are the only cetartiodactyl groups that have retained gremlin 2. According to the phyletic distribution of this gene and synteny analyses, we propose that gremlin 2 was lost in the common ancestor of ruminants and cetaceans between 56.3 and 63.5 million years ago as a product of a chromosomal rearrangement. -

14-3-3Ζ, a Novel Androgen-Responsive Gene, Is Upregulated in Prostate Cancer And
Author Manuscript Published OnlineFirst on August 17, 2012; DOI: 10.1158/1078-0432.CCR-12-0281 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. 14-3-3ζ, a novel androgen-responsive gene, is upregulated in prostate cancer and promotes prostate cancer cell proliferation and survival Taro Murata1,2 Ϯ, Ken-ichi Takayama1,3,4 Ϯ, Tomohiko Urano1,3,4, Tetsuya Fujimura2, Daisaku Ashikari1,5, Daisuke Obinata1,5, Kuniko Horie-Inoue4, Satoru Takahashi5, Yasuyoshi Ouchi3, Yukio Homma2, Satoshi Inoue1,3,4 Ϯ: These authors contributed equally to this work. 1 Department of Anti-Aging Medicine, Graduate School of Medicine, The University of Tokyo, Tokyo, Japan, 2 Department of Urology, Graduate School of Medicine, The University of Tokyo, Tokyo, Japan, 3 Department of Geriatric Medicine, Graduate School of Medicine, The University of Tokyo, Tokyo, Japan, 4 Division of Gene Regulation and Signal Transduction, Research Center for Genomic Medicine, Saitama Medical University, Saitama, Japan, 5 Department of Urology, School of Medicine, Nihon University, Tokyo, Japan Running Title: Role of 14-3-3ζ in prostate cancer Key Words: Androgen; prostate cancer; 14-3-3 protein; androgen receptor Corresponding Author: Satoshi Inoue, Department of Anti-Aging Medicine, Graduate School of Medicine, University of Tokyo, Bunkyo-ku, Tokyo 113-8655, Japan. Phone: 81-3-5800-8834; Fax: 81-3-5800-9126; E-mail: [email protected]. 1 Downloaded from clincancerres.aacrjournals.org on September 28, 2021. © 2012 American Association for Cancer Research. Author Manuscript Published OnlineFirst on August 17, 2012; DOI: 10.1158/1078-0432.CCR-12-0281 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. -

Akt Is Involved in the Inhibition of Cell Proliferation by EGF
EXPERIMENTAL and MOLECULAR MEDICINE, Vol. 39, No. 4, 491-498, August 2007 Akt is involved in the inhibition of cell proliferation by EGF Soung Hoo Jeon1, Woo-Jeong Jeong1, proliferation and embryonic axis formation (Zeng et Jae-Young Cho1, Kee-Ho Lee2 al., 1997). Axin is a multi-domain scaffold protein and Kang-Yell Choi1,3 that associates directly with β-catenin, GSK3β, and phosphatase PP2A (Hsu et al., 1999). Axin is 1 implicated in down-regulation of Wnt/β-catenin Department of Biotechnology signaling (Ikeda et al., 1998; Itoh et al., 1998; Yonsei University Sakanaka et al., 1998; Kikuchi et al., 2006). There Seoul 120-752, Korea 2 are two vertebrate Axins (Axin 1 and Axin 2). Axin1 Laboratory of Molecular Oncology is constitutively expressed, but Axin 2 is induced Korea Institute of Radiological and Medical Sciences by active Wnt signaling and acts therefore in a Seoul 139-706, Korea negative feedback loop (Yan et al., 2001; Jho et 3 Corresponding author: Tel, 82-2-2123-2887; al., 2002; Lustig et al., 2002). Overexpressed Axin Fax, 82-2-362-7265; E-mail, [email protected] destabilizes β-catenin (Behrens et al., 1998; Hart et al., 1998; Ikeda et al.,1998; Kishida et al., 1998; Accepted 28 May 2007 Nakamura et al., 1998; Sakanaka et al., 1998; Yamamoto et al., 1998). Loss of Axin results in nu- Abbreviations: APC, adenomatos polyposis coli; BrdU, bromode- clear accumulation of β-catenin followed by forma- oxyuridine; DAPI, 4'6-diamidino-2-phenylindole; PI3K, phosphatidyl tion of the β-catenin-TCF transcriptional complex inositol 3-kinase involving transcriptional activation of its target genes (Sakanaka et al., 1998). -

Dedifferentiation, Transdifferentiation, and Reprogramming: Future Directions in Regenerative Medicine
82 Dedifferentiation, Transdifferentiation, and Reprogramming: Future Directions in Regenerative Medicine Cristina Eguizabal, PhD1 Nuria Montserrat, PhD1 Anna Veiga, PhD1,2 Juan Carlos Izpisua Belmonte, PhD1,3 1 Center for Regenerative Medicine in Barcelona Address for correspondence and reprint requests Juan Carlos Izpisua 2 Reproductive Medicine Service, Institut Universitari Dexeus, Belmonte, PhD, Gene Expression Laboratory, The Salk Institute for Barcelona, Spain Biological Studies, 10010 North Torrey Pines Road, La Jolla, CA 93027 3 Gene Expression Laboratory, The Salk Institute for Biological Studies, (e-mail: [email protected]). La Jolla, California Semin Reprod Med 2013;31:82–94 Abstract The main goal of regenerative medicine is to replace damaged tissue. To do this it is Keywords necessary to understand in detail the whole regeneration process including differenti- ► regenerative ated cells that can be converted into progenitor cells (dedifferentiation), cells that can medicine switch into another cell type (transdifferentiation), and somatic cells that can be ► stem cells induced to become pluripotent cells (reprogramming). By studying the regenerative ► dedifferentiation processes in both nonmammal and mammal models, natural or artificial processes ► transdifferentiation could underscore the molecular and cellular mechanisms behind these phenomena and ► reprogramming be used to create future regenerative strategies for humans. To understand any regenerative system, it is crucial to find the potency and differentiate and how they can revert to pluri- cellular origins of renewed tissues. Using techniques like potency (reprogramming) or switch lineages (dedifferentia- genetic lineage tracing and single-cell transplantation helps tion and transdifferentiation). to identify the route of regenerative sources. These tools were We synthesize the studies of different model systems to developed first in nonmammal models (flies, amphibians, and highlight recent insights that are integrating the field. -

The Utility of Resolving Asthma Molecular Signatures Using Tissue-Specific Transcriptome Data 11 12 13 Debajyoti Ghosh1, Lili Ding2, Jonathan A
G3: Genes|Genomes|Genetics Early Online, published on September 8, 2020 as doi:10.1534/g3.120.401718 1 2 3 4 5 6 7 8 9 10 The utility of resolving asthma molecular signatures using tissue-specific transcriptome data 11 12 13 Debajyoti Ghosh1, Lili Ding2, Jonathan A. Bernstein1, and Tesfaye B. Mersha3* 14 15 1Immunology and Allergy, Department of Internal Medicine, University of Cincinnati, Cincinnati, OH, 16 United States of America 17 2Division of Biostatistics and Epidemiology, Cincinnati Children’s Hospital Medical Center, Department 18 of Pediatrics, University of Cincinnati, Cincinnati, OH, United States of America 19 3Division of Asthma Research, Cincinnati Children’s Hospital Medical Center, Department of Pediatrics, 20 University of Cincinnati, Cincinnati, OH, United States of America 21 22 23 24 25 Corresponding author: 26 27 Tesfaye B. Mersha, Ph.D. 28 Associate Professor 29 Cincinnati Children's Hospital Medical Center 30 Department of Pediatrics 31 University of Cincinnati 32 3333 Burnet Avenue 33 MLC 7037 34 Cincinnati, OH 45229-3026 35 Phone: (513) 803-2766 36 Fax: (513) 636-1657 37 Email: [email protected] 38 39 40 41 1 © The Author(s) 2020. Published by the Genetics Society of America. 42 Graphical Abstract 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 2 61 ABSTRACT 62 An integrative analysis focused on multi-tissue transcriptomics has not been done for asthma. Tissue- 63 specific DEGs remain undetected in many multi-tissue analyses, which influences identification of 64 disease-relevant pathways and potential drug candidates.