Characterization of the Skin Cultivable Microbiota Composition of the Frog Pelophylax Perezi Inhabiting Different Environments
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Epidemiological and Molecular Analysis of Virulence and Antibiotic Resistance in Acinetobacter Baumannii
UNIVERSIDAD COMPLUTENSE DE MADRID FACULTAD DE VETERINARIA DEPARTAMENTO DE BIOQUÍMICA Y BIOLOGÍA MOLECULAR IV TESIS DOCTORAL Epidemiological and Molecular Analysis of Virulence and Antibiotic Resistance in Acinetobacter baumannii Análisis Epidemiológico y Molecular de la Virulencia y la Antibiorresistencia en Acinetobacter baumannii MEMORIA PARA OPTAR AL GRADO DE DOCTOR PRESENTADA POR Elias Dahdouh DIRECTORA Mónica Suárez Rodríguez Madrid, 2017 © Elias Dahdouh, 2016 UNIVERSIDAD COMPLUTENSE DE MADRID FACULTAD DE VETERINARIA DEPARTAMENTO DE BIOQUIMICA Y BIOLOGIA MOLECULAR IV TESIS DOCTORAL Análisis Epidemiológico y Molecular de la Virulencia y la Antibiorresistencia en Acinetobacter baumannii Epidemiological and Molecular Analysis of Virulence and Antibiotic Resistance in Acinetobacter baumannii MEMORIA PARA OPTAR AL GRADO DE DOCTOR PRESENTADA POR Elias Dahdouh Directora Mónica Suárez Rodríguez Madrid, 2016 UNIVERSIDAD COMPLUTENSE DE MADRID FACULTAD DE VETERINARIA Departamento de Bioquímica y Biología Molecular IV ANALYSIS EPIDEMIOLOGICO Y MOLECULAR DE LA VIRULENCIA Y LA ANTIBIORRESISTENCIA EN Acinetobacter baumannii EPIDEMIOLOGICAL AND MOLECULAR ANALYSIS OF VIRULENCE AND ANTIBIOTIC RESISTANCE IN Acinetobacter baumannii MEMORIA PARA OPTAR AL GRADO DE DOCTOR PRESENTADA POR Elias Dahdouh Bajo la dirección de la doctora Mónica Suárez Rodríguez Madrid, Diciembre de 2016 First and foremost, I would like to thank God for the continued strength and determination that He has given me. I would also like to thank my father Abdo, my brother Charbel, my fiancée, Marisa, and all my friends for their endless support and for standing by me at all times. Moreover, I would like to thank Dra. Monica Suarez Rodriguez and Dr. Ziad Daoud for giving me the opportunity to complete this doctoral study and for their guidance, encouragement, and friendship. -

Gut Microbiome Alterations in Ulcerative Colitis and After Moxibustion Intervention
Gut Microbiome Alterations In Ulcerative Colitis And After Moxibustion Intervention Qin Qi Shanghai University of Traditional Chinese Medicine Ya-Nan Liu Shanghai University of Traditional Chinese Medicine Si-Yi Lv Shanghai University of Traditional Chinese Medicine Huan-Gan Wu Shanghai University of Traditional Chinese Medicine Lin-Shuang Zhang Zhejiang Institute for Food and Drug Control Zhan Cao Tongji University School of Medicine Hui-Rong Liu Shanghai University of Traditional Chinese Medicine Xiao-Mei Wang ( [email protected] ) Shanghai University of Traditional Chinese Medicine Lu-Yi Wu Shanghai University of Traditional Chinese Medicine Research Article Keywords: Ulcerative colitis, Moxibustion, Gut microbiota, Metagenomic Posted Date: August 11th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-789670/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/22 Abstract Background: Recent studies have shown that the pathogenesis of ulcerative colitis (UC) is closely related to the gut microbiota. Moxibustion, a common treatment in traditional Chinese medicine, is the burning of the herb moxa over acupuncture points. Moxibustion has been used to improve the inammation and gastrointestinal dysfunctions in gastrointestinal disorders such as UC. In this study, we investigated whether moxibustion could improve the gut microbial dysbiosis induced by dextran sulphate sodium (DSS). Methods: Twenty-ve male rats were randomly assigned into ve groups: normal (NG), UC model (UC), moxibustion (UC+MOX), mesalazine (UC+MES), and normal rats with moxibustion (NG+MOX). The UC rat model was established by administering DSS solution. The rats in the UC+MOX and NG+MOX groups were treated with moxibustion at Tianshu (bilateral, ST25) points once daily for 7 consecutive days, and the UC+MES group rats were treated with mesalazine once daily for 7 consecutive days. -

The Microbiota Continuum Along the Female Reproductive Tract and Its Relation to Uterine-Related Diseases
ARTICLE DOI: 10.1038/s41467-017-00901-0 OPEN The microbiota continuum along the female reproductive tract and its relation to uterine-related diseases Chen Chen1,2, Xiaolei Song1,3, Weixia Wei4,5, Huanzi Zhong 1,2,6, Juanjuan Dai4,5, Zhou Lan1, Fei Li1,2,3, Xinlei Yu1,2, Qiang Feng1,7, Zirong Wang1, Hailiang Xie1, Xiaomin Chen1, Chunwei Zeng1, Bo Wen1,2, Liping Zeng4,5, Hui Du4,5, Huiru Tang4,5, Changlu Xu1,8, Yan Xia1,3, Huihua Xia1,2,9, Huanming Yang1,10, Jian Wang1,10, Jun Wang1,11, Lise Madsen 1,6,12, Susanne Brix 13, Karsten Kristiansen1,6, Xun Xu1,2, Junhua Li 1,2,9,14, Ruifang Wu4,5 & Huijue Jia 1,2,9,11 Reports on bacteria detected in maternal fluids during pregnancy are typically associated with adverse consequences, and whether the female reproductive tract harbours distinct microbial communities beyond the vagina has been a matter of debate. Here we systematically sample the microbiota within the female reproductive tract in 110 women of reproductive age, and examine the nature of colonisation by 16S rRNA gene amplicon sequencing and cultivation. We find distinct microbial communities in cervical canal, uterus, fallopian tubes and perito- neal fluid, differing from that of the vagina. The results reflect a microbiota continuum along the female reproductive tract, indicative of a non-sterile environment. We also identify microbial taxa and potential functions that correlate with the menstrual cycle or are over- represented in subjects with adenomyosis or infertility due to endometriosis. The study provides insight into the nature of the vagino-uterine microbiome, and suggests that sur- veying the vaginal or cervical microbiota might be useful for detection of common diseases in the upper reproductive tract. -

Exploring the Microbiome: Diversity of the Microbial Community of Three Foam Nesting Frogs, Genus: Polypedates, Across a Developmental Gradient Sarah Mcgrath
James Madison University JMU Scholarly Commons Masters Theses The Graduate School Summer 2018 Exploring the microbiome: diversity of the microbial community of three foam nesting frogs, Genus: Polypedates, across a developmental gradient Sarah McGrath Follow this and additional works at: https://commons.lib.jmu.edu/master201019 Part of the Environmental Microbiology and Microbial Ecology Commons, and the Other Microbiology Commons Recommended Citation McGrath, Sarah, "Exploring the microbiome: diversity of the microbial community of three foam nesting frogs, Genus: Polypedates, across a developmental gradient" (2018). Masters Theses. 575. https://commons.lib.jmu.edu/master201019/575 This Thesis is brought to you for free and open access by the The Graduate School at JMU Scholarly Commons. It has been accepted for inclusion in Masters Theses by an authorized administrator of JMU Scholarly Commons. For more information, please contact [email protected]. Exploring the microbiome: diversity of the microbial community of three foam nesting frogs, Genus: Polypedates, across a developmental gradient Sarah McGrath A thesis submitted to the Graduate Faculty of JAMES MADISON UNIVERSITY In Partial Fulfillment of the Requirements for the degree of Master of Science Department of Biology August 2018 FACULTY COMMITTEE: Committee Chair: Dr. David S. McLeod Committee Members/Readers: Dr. Morgan Steffen Dr. Reid Harris Dedication This work is dedicated to my father, John Robinson McGrath, for instilling in me a love of nature; my mother, Susanna Terrell McGrath, for continuously demonstrating that hard work pays off; Nicholas Blaser, for being an unwavering source of support; and Jennifer A. Sheridan, for providing me with my first opportunity to fully understand the demand and intricacy of tropical fieldwork. -

Identification, Molecular Epidemiology, and Antibiotic Resistance Characterization of Acinetobacter Spp
FACULTY OF HEALTH SCIENCES DEPARTMENT OF MEDICAL BIOLOGY UNIVERSITY HOSPITAL OF NORTH NORWAY DEPARTMENT OF MICROBIOLOGY AND INFECTION CONTROL REFERENCE CENTRE FOR DETECTION OF ANTIMICROBIAL RESISTANCE Identification, molecular epidemiology, and antibiotic resistance characterization of Acinetobacter spp. clinical isolates Nabil Karah A dissertation for the degree of Philosophiae Doctor June 2011 Acknowledgments The work presented in this thesis has been carried out between January 2009 and September 2011 at the Reference Centre for Detection of Antimicrobial Resistance (K-res), Department of Microbiology and Infection Control, University Hospital of North Norway (UNN); and the Research Group for Host–Microbe Interactions, Department of Medical Biology, Faculty of Health Sciences, University of Tromsø (UIT), Tromsø, Norway. I would like to express my deep and truthful acknowledgment to my main supervisor Ørjan Samuelsen. His understanding and encouraging supervision played a major role in the success of every experiment of my PhD project. Dear Ørjan, I am certainly very thankful for your indispensible contribution in all the four manuscripts. I am also very grateful to your comments, suggestions, and corrections on the present thesis. I am sincerely grateful to my co-supervisor Arnfinn Sundsfjord for his important contribution not only in my MSc study and my PhD study but also in my entire career as a “Medical Microbiologist”. I would also thank you Arnfinn for your nonstop support during my stay in Tromsø at a personal level. My sincere thanks are due to co-supervisors Kristin Hegstad and Gunnar Skov Simonsen for the valuable advice, productive comments, and friendly support. I would like to thank co-authors Christian G. -

A Comparison of Genospecies of Clinical Isolates in the Acinetobacter Spp. Complex Obtained from Hospitalized Patients in Busan, Korea
Biomedical Science Letters 2019, 25(1): 40 ~53 Original Article https://doi.org/10.15616/BSL.2019.25.1.40 eISSN : 2288-7415 A Comparison of Genospecies of Clinical Isolates in the Acinetobacter spp. Complex Obtained from Hospitalized Patients in Busan, Korea Gyu-Nam Park 1,*, Hye-Sook Kang 2,** , Hye-Ran Kim 3,** *, Bo-Kyung Jung 1,*, Do-Hee Kim 4,* and Kyung-Soo Chang 1, †,*** 1Department of Clinical Laboratory Science, College of Health Sciences, Catholic University of Pusan, Busan 46252, Korea 2Department of Laboratory Medicine, Maryknoll Medical Center, Busan 48972, Korea 3Department of Clinical Laboratory Science, College of Health and Therapy, Daegu Haany University, Gyeongsangbuk-Do 38610, Korea 4Department of Laboratory Medicine, Busan Veterans Hospital, Busan 46996, Korea Of the Acinetobacter spp., A. baumannii (genospecies 2) is the most clinically significant in terms of hospital-acquired infections worldwide. It is difficult to perform Acinetobacter -related taxonomy using phenotypic characteristics and routine laboratory methods owing to clusters of closely related species. The ability to accurately identify Acinetobacter spp. is clinically important because antimicrobial susceptibility and clinical relevance differs significantly among the different genospecies. Based on the medical importance of pathogenic Acinetobacter spp., the distribution and characterization of Acinetobacter spp. isolates from 123 clinical samples was determined in the current study using four typically applied bacterial identification methods; partial rpoB gene sequencing, amplified rRNA gene restriction analysis (ARDRA) of the intergenic transcribed spacer (ITS) region of the 16 ~23S rRNA, the VITEK ® 2 system (an automated microbial identification system) and matrix-assisted laser desorption/ionization-time of flight mass spectrometry (MALDI-TOF MS). -

Detection of Acinetobacter Spp. in Blood Cultures by an Improved Fluorescent in Situ Hybridization Assay
Polish Journal of Microbiology 2018, Vol. 67, No 1, 3–10 ORIGINAL PAPER Detection of Acinetobacter spp. in Blood Cultures by an Improved Fluorescent in Situ Hybridization Assay HANIEH ASAADI1, 2, BEHROUZ NAEIMI1, 3, SOMAYYEH GHARIBI4, ABDALNASER KHOSRAVI1, SINA DOBARADARAN5, REZA TAHERKHANI1, 3 and SAEED TAJBAKHSH1, 3* 1 Department of Microbiology and Parasitology, Faculty of Medicine, Bushehr University of Medical Sciences, Bushehr, Iran 2 Student Research Committee, Bushehr University of Medical Sciences, Bushehr, Iran 3 The Persian Gulf Tropical Medicine Research Center, Bushehr University of Medical Sciences, Bushehr, Iran 4 Department of Microbiology, Faculty of Biological Sciences, Alzahra University, Tehran, Iran 5 Department of Environmental Health Engineering, Faculty of Health, Bushehr University of Medical Sciences, Bushehr, Iran Submitted 25 May 2017, revised 1 October 2017, accepted 28 November 2017 Abstract Fluorescent in situ hybridization (FISH) allows rapid detection of microorganisms. We aimed (i) to evaluate the sensitivity and specific- ity of FISH for the detection of Acinetobacter spp. in blood culture specimens and (ii) to test the simultaneous application of two genus- specific probes labeled with the same fluorochrome to increase the fluorescent signal intensity and improve the detection of Acinetobacter spp. Three hundred and twenty blood culture specimens were testedvia both the conventional laboratory methods and FISH to detect Acinetobacter spp. The specimens were examined separately with each genus-specific probe Aci and ACA, and also using a mixture of the both probes Aci and ACA. In all examinations, probe EUB338 was used accompanied by Aci and ACA. The specificity of FISH was 100% (97.5% confidence interval [CI] = 98.7% – 100%). -

The Genetic Analysis of an Acinetobacter Johnsonii Clinical Strain Evidenced the Presence of Horizontal Genetic Transfer
RESEARCH ARTICLE The Genetic Analysis of an Acinetobacter johnsonii Clinical Strain Evidenced the Presence of Horizontal Genetic Transfer Sabrina Montaña1, Sareda T. J. Schramm2, German Matías Traglia1, Kevin Chiem1,2, Gisela Parmeciano Di Noto1, Marisa Almuzara3, Claudia Barberis3, Carlos Vay3, Cecilia Quiroga1, Marcelo E. Tolmasky2, Andrés Iriarte4, María Soledad Ramírez1,2* 1 Instituto de Investigaciones en Microbiología y Parasitología Médica (IMPaM, UBA-CONICET), Buenos Aires, Argentina, 2 Department of Biological Science, California State University Fullerton, Fullerton, CA, a11111 United States of America, 3 Laboratorio de Bacteriología Clínica, Departamento de Bioquímica Clínica, Hospital de Clínicas José de San Martín, Facultad de Farmacia y Bioquímica, Buenos Aires, Argentina, 4 Departamento de Desarrollo Biotecnológico, Instituto de Higiene, Facultad de Medicina, UdelaR, Montevideo, Uruguay * [email protected] OPEN ACCESS Abstract Citation: Montaña S, Schramm STJ, Traglia GM, Chiem K, Parmeciano Di Noto G, Almuzara M, et al. Acinetobacter johnsonii rarely causes human infections. While most A. johnsonii isolates are (2016) The Genetic Analysis of an Acinetobacter β johnsonii Clinical Strain Evidenced the Presence of susceptible to virtually all antibiotics, strains harboring a variety of -lactamases have Horizontal Genetic Transfer. PLoS ONE 11(8): recently been described. An A. johnsonii Aj2199 clinical strain recovered from a hospital in e0161528. doi:10.1371/journal.pone.0161528 Buenos Aires produces PER-2 and OXA-58. We decided to delve into its genome by obtain- Editor: Ruth Hall, University of Sydney, AUSTRALIA ing the whole genome sequence of the Aj2199 strain. Genome comparison studies on Received: March 23, 2016 Aj2199 revealed 240 unique genes and a close relation to strain WJ10621, isolated from the urine of a patient in China. -

The Environmental Acinetobacter Baumannii Isolate DSM30011
GBE The Environmental Acinetobacter baumannii Isolate DSM30011 Reveals Clues into the Preantibiotic Era Genome Diversity, Virulence Potential, and Niche Range of a Predominant Nosocomial Pathogen Guillermo D. Repizo1,2,*, Alejandro M. Viale2,Vıtor Borges3,Marıa M. Cameranesi2, Najwa Taib4, Martın Espariz2,Ce´lineBrochier-Armanet4,Joao~ Paulo Gomes3, and Suzana P. Salcedo1 1Laboratory of Molecular Microbiology and Structural Biochemistry, CNRS UMR5086, University of Lyon, France 2Departamento de Microbiologia, Instituto de Biologia Molecular y Celular de Rosario (IBR, CONICET), Facultad de Ciencias Bioquimicas y Farmaceuticas, Universidad Nacional de Rosario, Argentina 3Bioinformatics Unit, Department of Infectious Diseases, National Institute of Health, Lisbon, Portugal 4Laboratoire de Biome´ trie et Biologie Evolutive, Univ. Lyon, Universite´ Lyon 1, CNRS, UMR5558, Villeurbanne, France *Corresponding author: E-mail: [email protected]. Accepted: August 24, 2017 Data deposition: The whole genome shotgun project for A. baumannii DSM30011 has been deposited at DDBJ/ENA/GenBank under the accession JJOC00000000. The version described in this paper is version JJOC02000000. Abstract Acinetobacter baumannii represents nowadays an important nosocomial opportunistic pathogen whose reservoirs outside the clinical setting are obscure. Here, we traced the origins of the collection strain A. baumannii DSM30011 to an isolate first reported in 1944, obtained from the enriched microbiota responsible of the aerobic decomposition of the resinous desert shrub guayule. Whole-genome sequencing and phylogenetic analysis based on core genes confirmed DSM30011 affiliation to A. baumannii. Comparative studies with 32 complete A. baumannii genomes revealed the presence of 12 unique accessory chromosomal regions in DSM30011 including five encompassing phage-related genes, five containing toxin genes of the type-6 secretion system, and one with an atypical CRISPRs/cas cluster. -
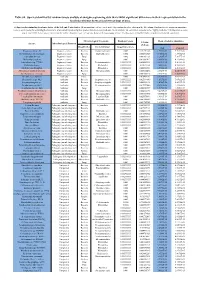
Table S8. Species Identified by Random Forests Analysis of Shotgun Sequencing Data That Exhibit Significant Differences In
Table S8. Species identified by random forests analysis of shotgun sequencing data that exhibit significant differences in their representation in the fecal microbiomes between each two groups of mice. (a) Species discriminating fecal microbiota of the Soil and Control mice. Mean importance of species identified by random forest are shown in the 5th column. Random forests assigns an importance score to each species by estimating the increase in error caused by removing that species from the set of predictors. In our analysis, we considered a species to be “highly predictive” if its importance score was at least 0.001. T-test was performed for the relative abundances of each species between the two groups of mice. P-values were at least 0.05 to be considered statistically significant. Microbiological Taxonomy Random Forests Mean of relative abundance P-Value Species Microbiological Function (T-Test) Classification Bacterial Order Importance Score Soil Control Rhodococcus sp. 2G Engineered strain Bacteria Corynebacteriales 0.002 5.73791E-05 1.9325E-05 9.3737E-06 Herminiimonas arsenitoxidans Engineered strain Bacteria Burkholderiales 0.002 0.005112829 7.1580E-05 1.3995E-05 Aspergillus ibericus Engineered strain Fungi 0.002 0.001061181 9.2368E-05 7.3057E-05 Dichomitus squalens Engineered strain Fungi 0.002 0.018887472 8.0887E-05 4.1254E-05 Acinetobacter sp. TTH0-4 Engineered strain Bacteria Pseudomonadales 0.001333333 0.025523638 2.2311E-05 8.2612E-06 Rhizobium tropici Engineered strain Bacteria Rhizobiales 0.001333333 0.02079554 7.0081E-05 4.2000E-05 Methylocystis bryophila Engineered strain Bacteria Rhizobiales 0.001333333 0.006513543 3.5401E-05 2.2044E-05 Alteromonas naphthalenivorans Engineered strain Bacteria Alteromonadales 0.001 0.000660472 2.0747E-05 4.6463E-05 Saccharomyces cerevisiae Engineered strain Fungi 0.001 0.002980726 3.9901E-05 7.3043E-05 Bacillus phage Belinda Antibiotic Phage 0.002 0.016409765 6.8789E-07 6.0681E-08 Streptomyces sp. -

Investigating Two Component Regulatory Systems for The
Investigating Two Component Regulatory Systems for the Determination of Adaptive Responses in A. baumannii. By LAURA PAULINE EVANS A thesis submitted to the University of Birmingham for the degree of MASTER OF PHILOSOPHY. Antimicrobial Agents Research Group School of Immunity and Infection College of Medical and Dental Sciences University Of Birmingham October 2012 University of Birmingham Research Archive e-theses repository This unpublished thesis/dissertation is copyright of the author and/or third parties. The intellectual property rights of the author or third parties in respect of this work are as defined by The Copyright Designs and Patents Act 1988 or as modified by any successor legislation. Any use made of information contained in this thesis/dissertation must be in accordance with that legislation and must be properly acknowledged. Further distribution or reproduction in any format is prohibited without the permission of the copyright holder. Abstract To investigate the role of the two component systems AdeRS and PmrAB in adaptation to the presence of antimicrobials, adeRS and pmrAB were deleted in multi-drug resistant Acinetobacter baumannii strain AYE. The effect of deleting these genes on antimicrobial susceptibility, growth, accumulation, virulence and ability to form a biofilm was investigated. The deletion of adeRS and pmrAB had no effect on bacterial growth or the accumulation of Hoechst 33342 (bis-benzimide). AYEΔpmrAB, but not AYEΔadeRS, accumulated significantly more norfloxacin than AYE. All strains accumulated more norfloxacin in the presence of the efflux inhibitor carbonyl cyanide m-chlorophenylhydrazone. AYE∆adeRS and AYE∆pmrAB, but not AYE, accumulated more norfloxacin in the presence of verapamil. -

Exploring Coral Microbiome Assemblages in the South China
www.nature.com/scientificreports OPEN Exploring coral microbiome assemblages in the South China Sea Lin Cai1, Ren-Mao Tian1, Guowei Zhou1,2, Haoya Tong1, Yue Him Wong1, Weipeng Zhang1, Apple Pui Yi Chui3, James Y. Xie4, Jian-Wen Qiu 4, Put O. Ang3, Sheng Liu2, Hui Huang2 & 1 Received: 21 July 2017 Pei-Yuan Qian Accepted: 18 January 2018 Coral reefs are signifcant ecosystems. The ecological success of coral reefs relies on not only coral-algal Published: xx xx xxxx symbiosis but also coral-microbial partnership. However, microbiome assemblages in the South China Sea corals remain largely unexplored. Here, we compared the microbiome assemblages of reef-building corals Galaxea (G. fascicularis) and Montipora (M. venosa, M. peltiformis, M. monasteriata) collected from fve diferent locations in the South China Sea using massively-parallel sequencing of 16S rRNA gene and multivariate analysis. The results indicated that microbiome assemblages for each coral species were unique regardless of location and were diferent from the corresponding seawater. Host type appeared to drive the coral microbiome assemblages rather than location and seawater. Network analysis was employed to explore coral microbiome co-occurrence patterns, which revealed 61 and 80 co-occurring microbial species assembling the Galaxea and Montipora microbiomes, respectively. Most of these co-occurring microbial species were commonly found in corals and were inferred to play potential roles in host nutrient metabolism; carbon, nitrogen, sulfur cycles; host detoxifcation; and climate change. These fndings suggest that the co-occurring microbial species explored might be essential to maintain the critical coral-microbial partnership. The present study provides new insights into coral microbiome assemblages in the South China Sea.