ATCC® Food & Water Testing Reference Strains Guide
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

NCTC) Bacterial Strain Equivalents to American Type Culture Collection (ATCC) Bacterial Strains
This list shows National Collection of Type Cultures (NCTC) bacterial strain equivalents to American Type Culture Collection (ATCC) bacterial strains. NCTC Number CurrentName ATCC Number NCTC 7212 Acetobacter pasteurianus ATCC 23761 NCTC 10138 Acholeplasma axanthum ATCC 25176 NCTC 10171 Acholeplasma equifetale ATCC 29724 NCTC 10128 Acholeplasma granularum ATCC 19168 NCTC 10172 Acholeplasma hippikon ATCC 29725 NCTC 10116 Acholeplasma laidlawii ATCC 23206 NCTC 10134 Acholeplasma modicum ATCC 29102 NCTC 10188 Acholeplasma morum ATCC 33211 NCTC 10150 Acholeplasma oculi ATCC 27350 NCTC 10198 Acholeplasma parvum ATCC 29892 NCTC 8582 Achromobacter denitrificans ATCC 15173 NCTC 10309 Achromobacter metalcaligenes ATCC 17910 NCTC 10807 Achromobacter xylosoxidans subsp. xylosoxidans ATCC 27061 NCTC 10808 Achromobacter xylosoxidans subsp. xylosoxidans ATCC 17062 NCTC 10809 Achromobacter xylosoxidans subsp. xylosoxidans ATCC 27063 NCTC 12156 Acinetobacter baumannii ATCC 19606 NCTC 10303 Acinetobacter baumannii ATCC 17904 NCTC 7844 Acinetobacter calcoaceticus ATCC 15308 NCTC 12983 Acinetobacter calcoaceticus ATCC 23055 NCTC 8102 acinetobacter dna group 13 ATCC 17903 NCTC 10304 Acinetobacter genospecies 13 ATCC 17905 NCTC 10306 Acinetobacter haemolyticus ATCC 17907 NCTC 10305 Acinetobacter haemolyticus subsp haemolyticus ATCC 17906 NCTC 10308 Acinetobacter johnsonii ATCC 17909 NCTC 10307 Acinetobacter junii ATCC 17908 NCTC 5866 Acinetobacter lwoffii ATCC 15309 NCTC 12870 Actinobacillus delphinicola ATCC 700179 NCTC 8529 Actinobacillus equuli ATCC 19392 -

Legionella Shows a Diverse Secondary Metabolism Dependent on a Broad Spectrum Sfp-Type Phosphopantetheinyl Transferase
Legionella shows a diverse secondary metabolism dependent on a broad spectrum Sfp-type phosphopantetheinyl transferase Nicholas J. Tobias1, Tilman Ahrendt1, Ursula Schell2, Melissa Miltenberger1, Hubert Hilbi2,3 and Helge B. Bode1,4 1 Fachbereich Biowissenschaften, Merck Stiftungsprofessur fu¨r Molekulare Biotechnologie, Goethe Universita¨t, Frankfurt am Main, Germany 2 Max von Pettenkofer Institute, Ludwig-Maximilians-Universita¨tMu¨nchen, Munich, Germany 3 Institute of Medical Microbiology, University of Zu¨rich, Zu¨rich, Switzerland 4 Buchmann Institute for Molecular Life Sciences, Goethe Universita¨t, Frankfurt am Main, Germany ABSTRACT Several members of the genus Legionella cause Legionnaires’ disease, a potentially debilitating form of pneumonia. Studies frequently focus on the abundant number of virulence factors present in this genus. However, what is often overlooked is the role of secondary metabolites from Legionella. Following whole genome sequencing, we assembled and annotated the Legionella parisiensis DSM 19216 genome. Together with 14 other members of the Legionella, we performed comparative genomics and analysed the secondary metabolite potential of each strain. We found that Legionella contains a huge variety of biosynthetic gene clusters (BGCs) that are potentially making a significant number of novel natural products with undefined function. Surprisingly, only a single Sfp-like phosphopantetheinyl transferase is found in all Legionella strains analyzed that might be responsible for the activation of all carrier proteins in primary (fatty acid biosynthesis) and secondary metabolism (polyketide and non-ribosomal peptide synthesis). Using conserved active site motifs, we predict Submitted 29 June 2016 some novel compounds that are probably involved in cell-cell communication, Accepted 25 October 2016 Published 24 November 2016 differing to known communication systems. -

Legionella Survey in the Plumbing System of a Sparse Academic Campus: a Case Study at the University of Perugia
Article Legionella Survey in the Plumbing System of a Sparse Academic Campus: A Case Study at the University of Perugia Ermanno Federici 1,*, Silvia Meniconi 2, Elisa Ceci 1, Elisa Mazzetti 2, Chiara Casagrande 1, Elena Montalbani 1, Stefania Businelli 3, Tatiana Mariani 3, Paolo Mugnaioli 3, Giovanni Cenci 1 and Bruno Brunone 2 1 Dipartimento di Chimica, Biologia e Biotecnologie, University of Perugia, 06123 Perugia, Italy; [email protected] (E.C.); [email protected] (C.C.); [email protected] (E.M.); [email protected] (G.C.) 2 Dipartimento di Ingegneria Civile ed Ambientale, University of Perugia, 06123 Perugia, Italy; [email protected] (S.M.); [email protected] (E.M.); [email protected] (B.B.) 3 Ripartizione Tecnica, University of Perugia, 06123 Perugia, Italy; [email protected] (S.B.); [email protected] (T.M.); [email protected] (P.M.) * Correspondence: [email protected]; Tel.: +39-075-5857330 Received: 30 June 2017; Accepted: 29 August 2017; Published: 1 September 2017 Abstract: We have monitored the presence of bacteria belonging to the genus Legionella in the plumbing of buildings at the University of Perugia (Italy). More than 300 water samples were collected from 156 control-point taps in 41 buildings comprised in the eight campuses of the University. Legionella was absent in most samples, while it was found in only 12 buildings (29% of the total). Molecular analysis indicated the presence of L. pneumophila (serogroups 1, 8 and 6–10), L. taurinensis and L. anisa. In only three cases contamination levels were above the limit at which remedial actions are required, according to international guidelines. -

Metatranscriptomic Analysis of Community Structure And
School of Environmental Sciences Metatranscriptomic analysis of community structure and metabolism of the rhizosphere microbiome by Thomas Richard Turner Submitted in partial fulfilment of the requirement for the degree of Doctor of Philosophy, September 2013 This copy of the thesis has been supplied on condition that anyone who consults it is understood to recognise that its copyright rests with the author and that use of any information derived there from must be in accordance with current UK Copyright Law. In addition, any quotation or extract must include full attribution. i Declaration I declare that this is an account of my own research and has not been submitted for a degree at any other university. The use of material from other sources has been properly and fully acknowledged, where appropriate. Thomas Richard Turner ii Acknowledgements I would like to thank my supervisors, Phil Poole and Alastair Grant, for their continued support and guidance over the past four years. I’m grateful to all members of my lab, both past and present, for advice and friendship. Graham Hood, I don’t know how we put up with each other, but I don’t think I could have done this without you. Cheers Salt! KK, thank you for all your help in the lab, and for Uma’s biryanis! Andrzej Tkatcz, thanks for the useful discussions about our projects. Alison East, thank you for all your support, particularly ensuring Graham and I did not kill each other. I’m grateful to Allan Downie and Colin Murrell for advice. For sequencing support, I’d like to thank TGAC, particularly Darren Heavens, Sophie Janacek, Kirsten McKlay and Melanie Febrer, as well as John Walshaw, Mark Alston and David Swarbreck for bioinformatic support. -

The Risk to Human Health from Free-Living Amoebae Interaction with Legionella in Drinking and Recycled Water Systems
THE RISK TO HUMAN HEALTH FROM FREE-LIVING AMOEBAE INTERACTION WITH LEGIONELLA IN DRINKING AND RECYCLED WATER SYSTEMS Dissertation submitted by JACQUELINE MARIE THOMAS BACHELOR OF SCIENCE (HONOURS) AND BACHELOR OF ARTS, UNSW In partial fulfillment of the requirements for the award of DOCTOR OF PHILOSOPHY in ENVIRONMENTAL ENGINEERING SCHOOL OF CIVIL AND ENVIRONMENTAL ENGINEERING FACULTY OF ENGINEERING MAY 2012 SUPERVISORS Professor Nicholas Ashbolt Office of Research and Development United States Environmental Protection Agency Cincinnati, Ohio USA and School of Civil and Environmental Engineering Faculty of Engineering The University of New South Wales Sydney, Australia Professor Richard Stuetz School of Civil and Environmental Engineering Faculty of Engineering The University of New South Wales Sydney, Australia Doctor Torsten Thomas School of Biotechnology and Biomolecular Sciences Faculty of Science The University of New South Wales Sydney, Australia ORIGINALITY STATEMENT '1 hereby declare that this submission is my own work and to the best of my knowledge it contains no materials previously published or written by another person, or substantial proportions of material which have been accepted for the award of any other degree or diploma at UNSW or any other educational institution, except where due acknowledgement is made in the thesis. Any contribution made to the research by others, with whom 1 have worked at UNSW or elsewhere, is explicitly acknowledged in the thesis. I also declare that the intellectual content of this thesis is the product of my own work, except to the extent that assistance from others in the project's design and conception or in style, presentation and linguistic expression is acknowledged.' Signed ~ ............................ -

Subcellular Location of Piscirickettsia Salmonis Heat Shock Protein 60 (Hsp60) Chaperone by Using Immunogold Labeling and Proteomic Analysis
microorganisms Article Subcellular Location of Piscirickettsia salmonis Heat Shock Protein 60 (Hsp60) Chaperone by Using Immunogold Labeling and Proteomic Analysis 1, 2,3, 4 5 Cristian Oliver y, Patricio Sánchez y , Karla Valenzuela , Mauricio Hernández , Juan Pablo Pontigo 3, Maria C. Rauch 3, Rafael A. Garduño 4,6 , Ruben Avendaño-Herrera 2,7,* and Alejandro J. Yáñez 2,8,* 1 Laboratorio de Inmunología y Estrés de Organismos Acuáticos, Instituto de Patología Animal, Facultad de Ciencias Veterinarias, Universidad Austral de Chile, Valdivia 5090000, Chile; [email protected] 2 Interdisciplinary Center for Aquaculture Research, (INCAR), Concepción 4070386, Chile; [email protected] 3 Instituto de Bioquímica y Microbiología, Facultad de Ciencias, Universidad Austral de Chile, Valdivia 5090000, Chile; [email protected] (J.P.P.); [email protected] (M.C.R.) 4 Microbiology and Immunology Department, Dalhousie University, Halifax, NS B3H 4R2, Canada; [email protected] (K.V.); [email protected] (R.A.G.) 5 Austral-OMICS, Faculty of Sciences, Universidad Austral de Chile, Valdivia 5090000, Chile; [email protected] 6 Canadian Food Inspection Agency, Dartmouth Laboratory, Dartmouth, NS B3B 1Y9, Canada 7 Universidad Andrés Bello, Laboratorio de Patología de Organismos Acuáticos y Biotecnología Acuícola, Facultad Ciencias de la Vida, Viña del Mar 2531015, Chile 8 Facultad de Ciencias, Universidad Austral de Chile, Valdivia 5090000, Chile * Correspondence: [email protected] (R.A.-H.); [email protected] (A.J.Y.) These authors contributed equally to this work. y Received: 12 November 2019; Accepted: 31 December 2019; Published: 15 January 2020 Abstract: Piscirickettsia salmonis is the causative bacterial agent of piscirickettsiosis, a systemic fish disease that significantly impacts the Chilean salmon industry. -

Genetic and Functional Studies of the Mip Protein of Legionella
t1.ì. Genetic and Functional Studies of the Mip Protein of Legionella Rodney Mark Ratcliff, BSc (Hons)' MASM Infectious Diseases Laboratories Institute of Medical and Veterinary Science and Department of Microbiology and Immunology UniversitY of Adelaide. Adelaide, South Australia A thesis submitted to the University of Adelaide for the degree of I)octor of Philosophy 15'h March 2000 amended 14th June 2000 Colonies of several Legionella strains on charcoal yeast extract agar (CYE) after 4 days incubation at 37"C in air. Various magnifications show typical ground-glass opalescent appearance. Some pure strains exhibit pleomorphic growth or colour. The top two photographs demonstrate typical red (LH) and blue-white (RH) fluorescence exhibited by some species when illuminated by a Woods (IJV) Lamp. * t Table of Contents .1 Chapter One: Introduction .1 Background .............'. .2 Morphology and TaxonomY J Legionellosis ............. 5 Mode of transmission "..'....'. 7 Environmental habitat 8 Interactions between Legionella and phagocytic hosts 9 Attachment 11 Engulfment and internalisation.'.. 13 Intracellular processing 13 Intracellular replication of legionellae .. " "' " "' 15 Host cell death and bacterial release 18 Virulence (the Genetic factors involved with intracellular multiplication and host cell killing .20 icm/dot system) Legiolysin .25 Msp (Znn* metaloprotease) ...'..... .25 .28 Lipopolysaccharide .29 The association of flagella with disease.. .30 Type IV fimbriae.... .31 Major outer membrane proteins....'.......'. JJ Heat shock proteins'.'. .34 Macrophage infectivity potentiator (Mip) protein Virulenceiraits of Legionella species other than L. pneumophila..........' .39 phylogeny .41 Chapter One (continued): Introduction to bacterial classification and .41 Identificati on of Legionella...'.,..'.. .46 Phylogeny .52 Methods of phylogenetic analysis' .53 Parsimony methods.'.. .55 Distance methods UPGMA cluster analYsis.'.'... -

Electronic Laboratory Reporting Use Case January 2011
ILLINOIS HEALTH INFORMATION EXCHANGE Electronic Laboratory Reporting Use Case Electronic Laboratory Reporting and Health Information Exchange Illinois Health Information Exchange Public Health Work Group January 2011 Electronic Laboratory Reporting Use Case January 2011 Table of Contents 1.0 Executive Summary……………………………………………………………………….3 2.0 Introduction…………………………………………………………………...……………..5 3.0 Scope……………………………………..………………………………………………………5 4.0 Use Case Stakeholders…………………………………………………………….….....6 5.0 Issues and Obstacles……………………………………………………………………...8 6.0 Use Case Pre-Conditions .………………….…………………………………………...8 7.0 Use Case Post-Conditions.……………………………………………………………...9 8.0 Detailed Scenarios/Technical Specifications.………………………………10 9.0 Validation and Certification………………………………………………………...12 Appendix ………………………………………………………………………………………….....13 Page 2 Electronic Laboratory Reporting Use Case January 2011 1.0 Executive Summary This Use Case is a product of the Public Health Work Group (PHWG) of the Illinois Health Information Exchange (HIE) Advisory Committee. The Illinois HIE Advisory Committee was constituted as the diverse public healthcare stakeholder body providing input and recommendations on the creation of the Illinois Health Information Exchange Authority (“the Authority”) as the Illinois vehicle for designing and implementing electronic health information exchange in Illinois. The establishment of the Authority marks the formal transition of the work of the HIE Advisory Committee and the Work Groups into alignment with the provisions of Illinois -

WO 2016/188962 Al 1 December 2016 (01.12.2016) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2016/188962 Al 1 December 2016 (01.12.2016) P O P C T (51) International Patent Classification: (74) Agents: GOODFELLOW, Hugh Robin et al; Carpmaels C12Q 1/68 (2006.01) & Ransford LLP, One Southampton Row, London WC1B 5HA (GB). (21) International Application Number: PCT/EP2016/061599 (81) Designated States (unless otherwise indicated, for every kind of national protection available): AE, AG, AL, AM, (22) Date: International Filing AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, 23 May 20 16 (23.05.2016) BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, (25) Filing Language: English DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IR, IS, JP, KE, KG, KN, KP, KR, (26) Publication Language: English KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, MG, (30) Priority Data: MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, 1508860.2 22 May 2015 (22.05.2015) GB PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, (71) Applicant: NATIONAL UNIVERSITY OF IRELAND, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. GALWAY [IE/IE]; University Road, Galway (IE). (84) Designated States (unless otherwise indicated, for every (72) Inventors: REDDINGTON, Kate Mary; Deerpack East, kind of regional protection available): ARIPO (BW, GH, Newport Road, Westport, Co. -

Legionella Pneumophila
Open Research Online The Open University’s repository of research publications and other research outputs Environmental regulation of the growth, physiology and virulence of Legionella pneumophila Thesis How to cite: Mauchline, William Stuart (1996). Environmental regulation of the growth, physiology and virulence of Legionella pneumophila. PhD thesis The Open University. For guidance on citations see FAQs. c 1995 The Author https://creativecommons.org/licenses/by-nc-nd/4.0/ Version: Version of Record Link(s) to article on publisher’s website: http://dx.doi.org/doi:10.21954/ou.ro.0000e0d1 Copyright and Moral Rights for the articles on this site are retained by the individual authors and/or other copyright owners. For more information on Open Research Online’s data policy on reuse of materials please consult the policies page. oro.open.ac.uk ENVIRONMENTAL REGULATION OF THE GROWTH, PHYSIOLOGY AND VIRULENCE OF LEGIONELLA PNEUMOPHILA. WILLIAM STUART MAUCHLINE A thesis submitted in partial fulfilment of the requirements of the Open University for the Degree of Doctor of Philosophy. The research presented in this thesis was undertaken at the Centre for Applied Microbiology and Research, Porton Down, Salisbury. Thames Water Utilities plc was the collaborating establishment. December 1995 APPENDECIES NOT SCANNED AT THE REQUEST OF THE AWARDING UNIVERSITY ABSTRACT Members of the Legionellaceae cause respiratory infections in man; the most severe, pneumonic form is known as Legionnaires' disease. Of the 39 species described to date 16 have been associated with human disease, however the majority of reported cases of legionellosis are caused by Legionella pneumophila serogroup 1. A number of pathogenic bacteria regulate their virulence gene expression in response to environmental stimuli. -
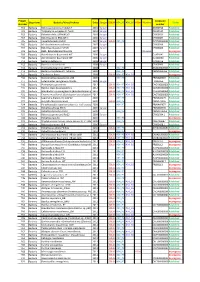
Project Number Organisms Bacteria/Virus/Archaea Date
Project_ Accession Organisms Bacteria/Virus/Archaea Date Sanger SOLiD 454_PE 454_SG PGM Illumina Status Number number P01 Bacteria Rickettsia conorii str.Malish 7 2001 Sanger AE006914 Published P02 Bacteria Tropheryma whipplei str.Twist 2003 Sanger AE014184 Published P03 Bacteria Rickettsia felis URRWXCal2 2005 Sanger CP000053 Published P04 Bacteria Rickettsia bellii RML369-C 2006 Sanger CP000087 Published P05 Bacteria Coxiella burnetii CB109 2007 Sanger SOLiD 454_PE AKYP00000000 Published P06 Bacteria Minibacterium massiliensis 2007 Sanger CP000269 Published P07 Bacteria Rickettsia massiliae MTU5 2007 Sanger CP000683 Published P08 Bacteria BaBL=Bête à Bernard Lascola 2007 Illumina In progress P09 Bacteria Acinetobacter baumannii AYE 2006 Sanger CU459141 Published P10 Bacteria Acinetobacter baumannii SDF 2006 Sanger CU468230 Published P11 Bacteria Borrelia duttonii Ly 2008 Sanger CP000976 Published P12 Bacteria Borrelia recurrentis A1 2008 Sanger CP000993 Published P13 Bacteria Francisella tularensis URFT1 2008 454_PE ABAZ00000000Published P14 Bacteria Borrelia crocidurae str. Achema 2009 454_PE PRJNA162335 Published P15 Bacteria Citrobacter koseri 2009 SOLiD 454_PE 454_SG In progress P16 Bacteria Diplorickettsia massiliensis 20B 2009 454_PE PRJNA86907 Published P17 Bacteria Enterobacter aerogenes EA1509E 2009 Sanger FO203355 Published P18 Bacteria Actinomyces grossensis 2012 SOLiD 454_PE 454_SG CAGY00000000Published P19 Bacteria Bacillus massiliosenegalensis 2012 SOLiD 454_PE 454_SG CAHJ00000000 Published P20 Bacteria Brevibacterium senegalensis -

Aquascreen® Legionella Species Qpcr Detection Kit
AquaScreen® Legionella species qPCR Detection Kit INSTRUCTIONS FOR USE FOR USE IN RESEARCH AND QUALITY CONTROL Symbols Lot No. Cat. No. Expiry date Storage temperature Number of reactions Manufacturer INDICATION The AquaScreen® Legionella species qPCR Detection kit is specifically designed for the quantitative detection of several Legionella species in water samples prepared with the AquaScreen® FastExt- ract kit. Its design complies with the requirements of AFNOR T90-471 and ISO/TS 12869:2012. Legionella are ubiquitous bacteria in surface water and moist soil, where they parasitize protozoa. The optimal growth temperature lies between +15 and +45 °C, whereas these gram-negative bacteria are dormant below 20 °C and do not survive above 60 °C. Importantly, Legionella are well-known as opportunistic intracellular human pathogens causing Legionnaires’ disease and Pontiac fever. The transmission occurs through inhalation of contami- nated aerosols generated by an infected source (e.g. human-made water systems like shower- heads, sink faucets, heaters, cooling towers, and many more). In order to efficiently prevent Legionella outbreaks, water safety control measures need syste- matic application but also reliable validation by fast Legionella testing. TEST PRINCIPLE The AquaScreen® Legionella species Kit uses qPCR for quantitative detection of legionella in wa- ter samples. In contrast to more time-consuming culture-based methods, AquaScreen® assays need less than six hours including sample preparation and qPCR to reliably detect Legionella. Moreover, the AquaScreen® qPCR assay has proven excellent performance in terms of specificity and sensitivity: other bacterial genera remain undetected whereas linear quantification is obtai- ned up to 1 x 106 particles per sample, therefore requiring no material dilution.