EIF4E Pser209 Rabbit Polyclonal Antibody – AP12890PU-N | Origene
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

EIF4E Antibody Cat
EIF4E Antibody Cat. No.: 60-600 EIF4E Antibody Formalin-fixed and paraffin-embedded human breast carcinoma reacted with EIF4E antibody (N-term), which was peroxidase- conjugated to the secondary antibody, followed by DAB staining. Specifications HOST SPECIES: Rabbit SPECIES REACTIVITY: Human Predicted species reactivity based on immunogen sequence: Zebrafish, Bovine, Mouse, HOMOLOGY: Rabbit, Rat, Xenopus This EIF4E antibody is generated from rabbits immunized with a KLH conjugated synthetic IMMUNOGEN: peptide between 32-61 amino acids from the N-terminal region of human EIF4E. TESTED APPLICATIONS: IHC-P, WB For WB starting dilution is: 1:1000 APPLICATIONS: For IHC-P starting dilution is: 1:50~100 September 27, 2021 1 https://www.prosci-inc.com/eif4e-antibody-60-600.html PREDICTED MOLECULAR 25 kDa WEIGHT: Properties This antibody is prepared by Saturated Ammonium Sulfate (SAS) precipitation followed by PURIFICATION: dialysis CLONALITY: Polyclonal ISOTYPE: Rabbit Ig CONJUGATE: Unconjugated PHYSICAL STATE: Liquid BUFFER: Supplied in PBS with 0.09% (W/V) sodium azide. CONCENTRATION: batch dependent Store at 4˚C for three months and -20˚C, stable for up to one year. As with all antibodies STORAGE CONDITIONS: care should be taken to avoid repeated freeze thaw cycles. Antibodies should not be exposed to prolonged high temperatures. Additional Info OFFICIAL SYMBOL: EIF4E Eukaryotic translation initiation factor 4E, eIF-4E, eIF4E, eIF-4F 25 kDa subunit, mRNA cap- ALTERNATE NAMES: binding protein, EIF4E, EIF4EL1, EIF4F ACCESSION NO.: P06730 PROTEIN GI NO.: 1352435 GENE ID: 1977 USER NOTE: Optimal dilutions for each application to be determined by the researcher. Background and References eIF4F is a multi-subunit complex, the composition of which varies with external and internal environmental conditions. -

Download Product Insert (PDF)
PRODUCT INFORMATION eIF4E (human recombinant) Item No. 15137 Overview and Properties Synonyms: eIF-4F 25 kDa subunit, Eukaryotic Translation Initiation Factor 4E, mRNA Cap-binding Protein Source: Recombinant protein expressed in E. coli Amino Acids: 2-217 (full length) Uniprot No.: P06730 Molecular Weight: 25.2 kDa Storage: -80°C (as supplied) Stability: ≥1 year Purity: batch specific (≥55% estimated by SDS-PAGE) Supplied in: 20 mM HEPES, pH 7.5, containing 100 mM potassium chloride, 2 mM DTT, and 10% glycerol Protein Concentration: batch specific mg/ml Image 1 2 3 4 250 kDa · · · · · · · 150 kDa · · · · · · · 100 kDa · · · · · · · 75 kDa · · · · · · · 50 kDa · · · · · · · 37 kDa · · · · · · · 25 kDa · · · · · · · 20 kDa · · · · · · · 15 kDa · · · · · · · Lane 1: MW Markers Lane 2: eIF4E (human recombinant) (4 µg) Lane 3: eIF4E (human recombinant) (2 µg) Lane 4: eIF4E (human recombinant) (1 µg) Representaõve gel image shown; actual purity may vary between each batch. WARNING CAYMAN CHEMICAL THIS PRODUCT IS FOR RESEARCH ONLY - NOT FOR HUMAN OR VETERINARY DIAGNOSTIC OR THERAPEUTIC USE. 1180 EAST ELLSWORTH RD SAFETY DATA ANN ARBOR, MI 48108 · USA This material should be considered hazardous until further information becomes available. Do not ingest, inhale, get in eyes, on skin, or on clothing. Wash thoroughly after handling. Before use, the user must review the complete Safety Data Sheet, which has been sent via email to your institution. PHONE: [800] 364-9897 WARRANTY AND LIMITATION OF REMEDY [734] 971-3335 Buyer agrees to purchase the material subject to Cayman’s Terms and Conditions. Complete Terms and Conditions including Warranty and Limitation of Liability information can be found on our website. -

Polyubiquitin Gene Ubb Is Required for Upregulation of Piwi Protein Level During Mouse Testis Development
www.nature.com/cddiscovery ARTICLE OPEN Polyubiquitin gene Ubb is required for upregulation of Piwi protein level during mouse testis development 1,4 2,4 2 1 1 2 ✉ Bitnara Han , Byung-Kwon✉ Jung , So-Hyun Park , Kyu Jin Song , Muhammad Ayaz Anwar , Kwon-Yul Ryu and Kwang Pyo Kim 1,3 © The Author(s) 2021 Testis development, including early embryonic gonad formation and late postnatal spermatogenesis, is essential for the reproduction of higher metazoans to generate fertile gametes, called sperm. We have previously reported that the polyubiquitin gene Ubb is required for fertility in both male and female mice. In particular, the Ubb-null male mice showed an azoospermia phenotype due to arrest of spermatogenesis at the pachytene stage. Here, we analyzed the whole testis proteome at postnatal day 20 to define the molecular mediators of the male-infertility phenotype caused by Ubb knockout. From the identified proteome, 564 proteins were significantly and differentially expressed in Ubb-knockout testes and, among these, 36 downregulated proteins were involved at different stages of spermatogenesis. We also found that levels of piRNA metabolic process-related proteins, including Piwil2 and Tdrd1, were downregulated in Ubb-null testes through functional gene ontology analysis. Further, protein–protein interaction mapping revealed that 24 testis development-related proteins, including Hsp90aa1, Eef1a1, and Pabpc1, were directly influenced by the depletion of ubiquitin. In addition, the reduced mRNA levels of these proteins were observed in Ubb-knockout testes, which closely resembled the global downregulation of piRNA-metabolic gene expression at the transcriptional and post- transcriptional levels. Together with proteomic and transcriptional analyses, our data suggest that Ubb expression is essential for the maintenance of testicular RNA-binding regulators and piRNA-metabolic proteins to complete spermatogenesis in mice. -
Beyond Traditional Morphological Characterization of Lung
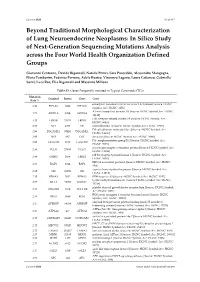
Cancers 2020 S1 of S15 Beyond Traditional Morphological Characterization of Lung Neuroendocrine Neoplasms: In Silico Study of Next-Generation Sequencing Mutations Analysis across the Four World Health Organization Defined Groups Giovanni Centonze, Davide Biganzoli, Natalie Prinzi, Sara Pusceddu, Alessandro Mangogna, Elena Tamborini, Federica Perrone, Adele Busico, Vincenzo Lagano, Laura Cattaneo, Gabriella Sozzi, Luca Roz, Elia Biganzoli and Massimo Milione Table S1. Genes Frequently mutated in Typical Carcinoids (TCs). Mutation Original Entrez Gene Gene Rate % eukaryotic translation initiation factor 1A X-linked [Source: HGNC 4.84 EIF1AX 1964 EIF1AX Symbol; Acc: HGNC: 3250] AT-rich interaction domain 1A [Source: HGNC Symbol;Acc: HGNC: 4.71 ARID1A 8289 ARID1A 11110] LDL receptor related protein 1B [Source: HGNC Symbol; Acc: 4.35 LRP1B 53353 LRP1B HGNC: 6693] 3.53 NF1 4763 NF1 neurofibromin 1 [Source: HGNC Symbol;Acc: HGNC: 7765] DS cell adhesion molecule like 1 [Source: HGNC Symbol; Acc: 2.90 DSCAML1 57453 DSCAML1 HGNC: 14656] 2.90 DST 667 DST dystonin [Source: HGNC Symbol;Acc: HGNC: 1090] FA complementation group D2 [Source: HGNC Symbol; Acc: 2.90 FANCD2 2177 FANCD2 HGNC: 3585] piccolo presynaptic cytomatrix protein [Source: HGNC Symbol; Acc: 2.90 PCLO 27445 PCLO HGNC: 13406] erb-b2 receptor tyrosine kinase 2 [Source: HGNC Symbol; Acc: 2.44 ERBB2 2064 ERBB2 HGNC: 3430] BRCA1 associated protein 1 [Source: HGNC Symbol; Acc: HGNC: 2.35 BAP1 8314 BAP1 950] capicua transcriptional repressor [Source: HGNC Symbol; Acc: 2.35 CIC 23152 CIC HGNC: -
Drosophila and Human Transcriptomic Data Mining Provides Evidence for Therapeutic
Drosophila and human transcriptomic data mining provides evidence for therapeutic mechanism of pentylenetetrazole in Down syndrome Author Abhay Sharma Institute of Genomics and Integrative Biology Council of Scientific and Industrial Research Delhi University Campus, Mall Road Delhi 110007, India Tel: +91-11-27666156, Fax: +91-11-27662407 Email: [email protected] Nature Precedings : hdl:10101/npre.2010.4330.1 Posted 5 Apr 2010 Running head: Pentylenetetrazole mechanism in Down syndrome 1 Abstract Pentylenetetrazole (PTZ) has recently been found to ameliorate cognitive impairment in rodent models of Down syndrome (DS). The mechanism underlying PTZ’s therapeutic effect is however not clear. Microarray profiling has previously reported differential expression of genes in DS. No mammalian transcriptomic data on PTZ treatment however exists. Nevertheless, a Drosophila model inspired by rodent models of PTZ induced kindling plasticity has recently been described. Microarray profiling has shown PTZ’s downregulatory effect on gene expression in fly heads. In a comparative transcriptomics approach, I have analyzed the available microarray data in order to identify potential mechanism of PTZ action in DS. I find that transcriptomic correlates of chronic PTZ in Drosophila and DS counteract each other. A significant enrichment is observed between PTZ downregulated and DS upregulated genes, and a significant depletion between PTZ downregulated and DS dowwnregulated genes. Further, the common genes in PTZ Nature Precedings : hdl:10101/npre.2010.4330.1 Posted 5 Apr 2010 downregulated and DS upregulated sets show enrichment for MAP kinase pathway. My analysis suggests that downregulation of MAP kinase pathway may mediate therapeutic effect of PTZ in DS. Existing evidence implicating MAP kinase pathway in DS supports this observation. -

Relevance of Translation Initiation in Diffuse Glioma Biology and Its
cells Review Relevance of Translation Initiation in Diffuse Glioma Biology and its Therapeutic Potential Digregorio Marina 1, Lombard Arnaud 1,2, Lumapat Paul Noel 1, Scholtes Felix 1,2, Rogister Bernard 1,3 and Coppieters Natacha 1,* 1 Laboratory of Nervous System Disorders and Therapy, GIGA-Neurosciences Research Centre, University of Liège, 4000 Liège, Belgium; [email protected] (D.M.); [email protected] (L.A.); [email protected] (L.P.N.); [email protected] (S.F.); [email protected] (R.B.) 2 Department of Neurosurgery, CHU of Liège, 4000 Liège, Belgium 3 Department of Neurology, CHU of Liège, 4000 Liège, Belgium * Correspondence: [email protected] Received: 18 October 2019; Accepted: 26 November 2019; Published: 29 November 2019 Abstract: Cancer cells are continually exposed to environmental stressors forcing them to adapt their protein production to survive. The translational machinery can be recruited by malignant cells to synthesize proteins required to promote their survival, even in times of high physiological and pathological stress. This phenomenon has been described in several cancers including in gliomas. Abnormal regulation of translation has encouraged the development of new therapeutics targeting the protein synthesis pathway. This approach could be meaningful for glioma given the fact that the median survival following diagnosis of the highest grade of glioma remains short despite current therapy. The identification of new targets for the development of novel therapeutics is therefore needed in order to improve this devastating overall survival rate. This review discusses current literature on translation in gliomas with a focus on the initiation step covering both the cap-dependent and cap-independent modes of initiation. -

Duke University Dissertation Template
Regulation of Eukaryotic Translation Initiation by Signal Transduction by Mayya Shveygert Department of Molecular Genetics and Microbiology Duke University Date:_______________________ Approved: ___________________________ Matthias Gromeier, Supervisor ___________________________ Christopher Nicchitta ___________________________ Jeffrey Rathmell ___________________________ Micah Luftig Dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Molecular Genetics and Microbiology in the Graduate School of Duke University 2012 i v ABSTRACT Regulation of Eukaryotic Translation Initiation by Signal Transduction by Mayya Shveygert Department of Molecular Genetics and Microbiology Duke University Date:_______________________ Approved: ___________________________ Matthias Gromeier, Supervisor ___________________________ Christopher Nicchitta ___________________________ Jeffrey Rathmell ___________________________ Micah Luftig An abstract of a dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Molecular Genetics and Microbiology in the Graduate School of Duke University 2012 Copyright by Mayya Shveygert 2012 Abstract Eukaryotic translation initiation is a rate-limiting step of protein synthesis and is controlled by signal transduction in response to various extracellular cues and stresses. This thesis is focused on the eukaryotic initiation factor 4G (eIF4G), which participates in multiple steps of initiation: (i) eIF4G interaction with polyA-binding protein (PABP) links the 5’ pre-initiation complex to the 3’ poly(A) tail of mRNAs; (ii) eIF4G recruits 40S ribosomal subunit to mRNAs through interactions with the m7-G cap binding protein eIF4E; (iii) eIF4G binds to the eIF4E kinase, mitogen activated protein kinase (MAPK) interacting kinase 1 (Mnk1), modulating eIF4E phosphorylation. I studied how eIF4G function is affected by viral infection, mitogenic stimulation and during mitosis. -

DEAD-Box Protein Family Member DDX28 Is a Negative Regulator Of
bioRxiv preprint doi: https://doi.org/10.1101/632331; this version posted May 9, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. DDX28 negatively regulates HIF-2α 1 DEAD-box protein family member DDX28 is a negative regulator 2 of HIF-2α and eIF4E2-directed hypoxic translation 3 Sonia L. Evagelou1, Olivia Bebenek1, Erin J. Specker1, James Uniacke1, * 4 1Department of Molecular and Cellular Biology, University of Guelph, 50 Stone Road East, 5 Guelph, Ontario, Canada, N1G 2W1 6 *Corresponding author: 7 James Uniacke: Department of Molecular and Cellular Biology, University of Guelph, 50 8 Stone Road East, Guelph, Ontario, Canada N1G 2W1; [email protected]; Tel.: 519- 9 824-4120 ext. 54739 10 RUNNING TITLE: DDX28 negatively regulates HIF-2α 11 Abbreviations: BrdU, bromodeoxyuridine; CCRCC, clear cell renal cell carcinoma; DDX28, 12 DEAD box protein 28; EEF2, Eukaryotic Translation Elongation Factor 2; EGFR, Epidermal 13 Growth Factor Receptor; eIF4E, eukaryotic initiation factor 4E; HIF, Hypoxia-inducible factor 14 HSP90ab1, Heat Shock Protein 90 Alpha Family Class B Member 1; IGF1R, Insulin Like 15 Growth Factor 1 Receptor; PEI, polyethylenamine; rHRE, RNA Hypoxia Response Elements 16 Keywords: Translation, hypoxia, HIF, DEAD-box protein, oxygen, eIF4E2 17 Conflict of interest: The authors declare that they have no conflicts of interest with the contents 18 of this article. 1 bioRxiv preprint doi: https://doi.org/10.1101/632331; this version posted May 9, 2019. -
Mrna Expression in Human Leiomyoma and Eker Rats As Measured by Microarray Analysis
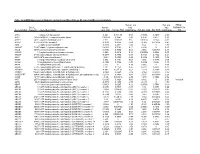
Table 3S: mRNA Expression in Human Leiomyoma and Eker Rats as Measured by Microarray Analysis Human_avg Rat_avg_ PENG_ Entrez. Human_ log2_ log2_ RAPAMYCIN Gene.Symbol Gene.ID Gene Description avg_tstat Human_FDR foldChange Rat_avg_tstat Rat_FDR foldChange _DN A1BG 1 alpha-1-B glycoprotein 4.982 9.52E-05 0.68 -0.8346 0.4639 -0.38 A1CF 29974 APOBEC1 complementation factor -0.08024 0.9541 -0.02 0.9141 0.421 0.10 A2BP1 54715 ataxin 2-binding protein 1 2.811 0.01093 0.65 0.07114 0.954 -0.01 A2LD1 87769 AIG2-like domain 1 -0.3033 0.8056 -0.09 -3.365 0.005704 -0.42 A2M 2 alpha-2-macroglobulin -0.8113 0.4691 -0.03 6.02 0 1.75 A4GALT 53947 alpha 1,4-galactosyltransferase 0.4383 0.7128 0.11 6.304 0 2.30 AACS 65985 acetoacetyl-CoA synthetase 0.3595 0.7664 0.03 3.534 0.00388 0.38 AADAC 13 arylacetamide deacetylase (esterase) 0.569 0.6216 0.16 0.005588 0.9968 0.00 AADAT 51166 aminoadipate aminotransferase -0.9577 0.3876 -0.11 0.8123 0.4752 0.24 AAK1 22848 AP2 associated kinase 1 -1.261 0.2505 -0.25 0.8232 0.4689 0.12 AAMP 14 angio-associated, migratory cell protein 0.873 0.4351 0.07 1.656 0.1476 0.06 AANAT 15 arylalkylamine N-acetyltransferase -0.3998 0.7394 -0.08 0.8486 0.456 0.18 AARS 16 alanyl-tRNA synthetase 5.517 0 0.34 8.616 0 0.69 AARS2 57505 alanyl-tRNA synthetase 2, mitochondrial (putative) 1.701 0.1158 0.35 0.5011 0.6622 0.07 AARSD1 80755 alanyl-tRNA synthetase domain containing 1 4.403 9.52E-05 0.52 1.279 0.2609 0.13 AASDH 132949 aminoadipate-semialdehyde dehydrogenase -0.8921 0.4247 -0.12 -2.564 0.02993 -0.32 AASDHPPT 60496 aminoadipate-semialdehyde -
Integrated Functional Genomic Analyses of Klinefelter and Turner Syndromes Reveal Global Network Effects of Altered X Chromosome Dosage
Integrated functional genomic analyses of Klinefelter and Turner syndromes reveal global network effects of altered X chromosome dosage Xianglong Zhanga,b,1, David Honga,1,2, Shining Mac,d,1, Thomas Warda,b, Marcus Hoa,b, Reenal Pattnia,b, Zhana Durenc,d, Atanas Stankova, Sharon Bade Shresthaa, Joachim Hallmayera, Wing Hung Wongc,d,2, Allan L. Reissa,2, and Alexander E. Urbana,b,2 aDepartment of Psychiatry and Behavioral Sciences, Stanford University School of Medicine, Stanford, CA 94305; bDepartment of Genetics, Stanford University School of Medicine, Stanford, CA 94305; cDepartment of Statistics, Stanford University, Stanford, CA 94305; and dDepartment of Biomedical Data Science, Stanford University, Stanford, CA 94305 Contributed by Wing Hung Wong, January 16, 2020 (sent for review June 20, 2019; reviewed by Carolyn J. Brown and Joseph F. Cubells) In both Turner syndrome (TS) and Klinefelter syndrome (KS) copy with short stature in TS (4) and tall stature in KS (5). X-chromosome number aberrations of the X chromosome lead to various develop- inactivation (XCI) transcriptionally silences one X chromosome mental symptoms. We report a comparative analysis of TS vs. KS in female mammals; as such, most genes on the inactivated X regarding differences at the genomic network level measured in chromosome are silenced and inactive (hereinafter referred to as primary samples by analyzing gene expression, DNA methylation, inactive genes). However, some genes escape XCI (hereinafter and chromatin conformation. X-chromosome inactivation (XCI) referred to as escape genes) and have also been hypothesized to silences transcription from one X chromosome in female mammals, contribute to the phenotypes of SCAs (6, 7). -
Driver-Map™ Immunenet™ Panel 1,741 Genes
Cellecta, Inc. 320 Logue Ave Mountain View, CA 94043 www.cellecta.com 650-938-3910 Driver-Map™ ImmuneNet™ Panel 1,741 Genes A2M APOE BCL3 CASP9 CD38 CHD4 CSNK2A2 DHX35 ETS1 FRG1 GZMM IFIT1 AADAT APOL2 BCL6 CAST CD3D CHD6 CSRP1 DHX58 ETV3 FRK HAL IFIT2 ABCB1 APOL3 BCL7A CAT CD3E CHD8 CST7 DHX8 EVI2B FRMD4A HAMP IFITM1 ABCB4 APOL6 BCLAF1 CBFB CD3EAP CHD9 CTAG1B DHX9 EWSR1 FRMD8 HAVCR2 IFITM2 ABCB9 APP BCOR CBL CD3G CHEK2 CTAGE1 DICER1 EZH2 FSHR HCFC1 IFNA1 ABCF1 AQP7 BEND5 CBLB CD4 CHI3L1 CTCF DIS3 F12 FUBP1 HCK IFNA10 ABL1 AQP9 BFSP1 CCAR1 CD40 CHI3L2 CTCFL DLG1 F13A1 FUT5 HCLS1 IFNA16 ABL2 AQR BID CCDC69 CD40LG CHIT1 CTLA4 DMAP1 F2RL1 FUT7 HDAC1 IFNA2 ACACA AR BIN2 CCL1 CD44 CHST15 CTNNB1 DMBT1 F8 FXR1 HDAC3 IFNA21 ACAD8 ARAP3 BIRC3 CCL11 CD46 CHST7 CTNND1 DNM1 FADD FYB HDAC9 IFNA5 ACAP1 AREG BIRC5 CCL13 CD47 CHUK CTSA DNMT3A FAF1 FYN HDC IFNA8 ACD ARF6 BLK CCL14 CD48 CIC CTSB DOCK2 FAIM3 FZD2 HECA IFNAR1 ACHE ARFGAP1 BLM CCL15 CD5 CIITA CTSC DOCK9 FAM124B FZD3 HESX1 IFNAR2 ACKR2 ARFGAP3 BLNK CCL16 CD52 CKLF CTSG DPEP2 FAM174B FZD9 HFE IFNB1 ACKR4 ARFGEF1 BMI1 CCL17 CD53 CLASP1 CTSH DPP4 FAM198B FZR1 HGF IFNGR1 ACO1 ARFGEF2 BMP2K CCL19 CD55 CLASP2 CTSL DSC3 FAM212B G3BP1 HHEX IFNGR2 ACP1 ARG1 BMPR1A CCL2 CD58 CLC CTSS DUSP2 FAM26F G3BP2 HIC1 IFNK ACSL3 ARG2 BMPR2 CCL20 CD59 CLCC1 CTSW DUSP4 FAM46C G6PD HIF1A IFNL2 ACSL6 ARHGAP15 BNC2 CCL21 CD6 CLCF1 CTSZ DUSP6 FAM65B GABBR1 HIST1H2AE IFNW1 ACTG2 ARHGAP22 BNIP3 CCL22 CD63 CLEC10A CTTN DZIP1 FANCI GAGE1 HIST1H2BG IGF1R ACVR1B ARHGAP26 BPI CCL23 CD68 CLEC18C CUL1 EBI3 FAS GAL3ST4 HK3 IGF2BP1