DEAD-Box Protein Family Member DDX28 Is a Negative Regulator Of
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Cancer Cells Exploit Eif4e2-Directed Translation to Enhance Their Proliferation, Migration and Invasion
Cancer cells exploit eIF4E2-directed translation to enhance their proliferation, migration and invasion by Joseph F. Varga A Thesis presented to The University of Guelph In partial fulfilment of requirements for the degree of Master of Science in Molecular and Cellular Biology Guelph, Ontario, Canada © Joseph F. Varga, August, 2016 ABSTRACT Cancer cells exploit eIF4E2-directed translation to enhance their proliferation, migration and invasion Joseph F. Varga Advisor: University of Guelph, 2016 Professor James Uniacke Despite the diversity found in the genetic makeup of cancer, many cancers share the same tumor microenvironment. Hypoxia, an aspect of the tumor microenvironment, causes the suppression of the primary translational machinery. Hypoxic cells switch from using the eukaryotic initiation factor 4E (eIF4E) to using a homologue of eIF4E (eIF4E2), in order to initiate the translation of select mRNAs. This thesis investigates the role of eIF4E2-directed translation in a panel of cancer cell lines during autonomous proliferation, migration and invasion. In this thesis, we show that silencing eIF4E2 abrogates the autonomous proliferation of colon carcinoma. Silencing eIF4E2 in glioblastoma cells resulted in decreased migration and invasion. Furthermore, we link eIF4E2-directed translation of cadherin 22 with the hypoxic migration of glioblastoma. These findings answer questions regarding the biology of cancer and expand the current knowledge of genes exploited during tumor progression. This data also highlights eIF4E2 as a potential therapeutic target. Acknowledgements I would like to express my sincere gratitude to Dr. James Uniacke for taking me on as a graduate student in his laboratory and providing me with the opportunity to contribute to scientific research. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

WO 2019/079361 Al 25 April 2019 (25.04.2019) W 1P O PCT
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date WO 2019/079361 Al 25 April 2019 (25.04.2019) W 1P O PCT (51) International Patent Classification: CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, C12Q 1/68 (2018.01) A61P 31/18 (2006.01) DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, C12Q 1/70 (2006.01) HR, HU, ID, IL, IN, IR, IS, JO, JP, KE, KG, KH, KN, KP, KR, KW, KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, (21) International Application Number: MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, PCT/US2018/056167 OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, (22) International Filing Date: SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, 16 October 2018 (16. 10.2018) TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (25) Filing Language: English (84) Designated States (unless otherwise indicated, for every kind of regional protection available): ARIPO (BW, GH, (26) Publication Language: English GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, ST, SZ, TZ, (30) Priority Data: UG, ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, RU, TJ, 62/573,025 16 October 2017 (16. 10.2017) US TM), European (AL, AT, BE, BG, CH, CY, CZ, DE, DK, EE, ES, FI, FR, GB, GR, HR, HU, ΓΕ , IS, IT, LT, LU, LV, (71) Applicant: MASSACHUSETTS INSTITUTE OF MC, MK, MT, NL, NO, PL, PT, RO, RS, SE, SI, SK, SM, TECHNOLOGY [US/US]; 77 Massachusetts Avenue, TR), OAPI (BF, BJ, CF, CG, CI, CM, GA, GN, GQ, GW, Cambridge, Massachusetts 02139 (US). -

EIF4E Antibody Cat
EIF4E Antibody Cat. No.: 60-600 EIF4E Antibody Formalin-fixed and paraffin-embedded human breast carcinoma reacted with EIF4E antibody (N-term), which was peroxidase- conjugated to the secondary antibody, followed by DAB staining. Specifications HOST SPECIES: Rabbit SPECIES REACTIVITY: Human Predicted species reactivity based on immunogen sequence: Zebrafish, Bovine, Mouse, HOMOLOGY: Rabbit, Rat, Xenopus This EIF4E antibody is generated from rabbits immunized with a KLH conjugated synthetic IMMUNOGEN: peptide between 32-61 amino acids from the N-terminal region of human EIF4E. TESTED APPLICATIONS: IHC-P, WB For WB starting dilution is: 1:1000 APPLICATIONS: For IHC-P starting dilution is: 1:50~100 September 27, 2021 1 https://www.prosci-inc.com/eif4e-antibody-60-600.html PREDICTED MOLECULAR 25 kDa WEIGHT: Properties This antibody is prepared by Saturated Ammonium Sulfate (SAS) precipitation followed by PURIFICATION: dialysis CLONALITY: Polyclonal ISOTYPE: Rabbit Ig CONJUGATE: Unconjugated PHYSICAL STATE: Liquid BUFFER: Supplied in PBS with 0.09% (W/V) sodium azide. CONCENTRATION: batch dependent Store at 4˚C for three months and -20˚C, stable for up to one year. As with all antibodies STORAGE CONDITIONS: care should be taken to avoid repeated freeze thaw cycles. Antibodies should not be exposed to prolonged high temperatures. Additional Info OFFICIAL SYMBOL: EIF4E Eukaryotic translation initiation factor 4E, eIF-4E, eIF4E, eIF-4F 25 kDa subunit, mRNA cap- ALTERNATE NAMES: binding protein, EIF4E, EIF4EL1, EIF4F ACCESSION NO.: P06730 PROTEIN GI NO.: 1352435 GENE ID: 1977 USER NOTE: Optimal dilutions for each application to be determined by the researcher. Background and References eIF4F is a multi-subunit complex, the composition of which varies with external and internal environmental conditions. -

Gene Expression Profiling Analysis Contributes to Understanding the Association Between Non-Syndromic Cleft Lip and Palate, and Cancer
2110 MOLECULAR MEDICINE REPORTS 13: 2110-2116, 2016 Gene expression profiling analysis contributes to understanding the association between non-syndromic cleft lip and palate, and cancer HONGYI WANG, TAO QIU, JIE SHI, JIULONG LIANG, YANG WANG, LIANGLIANG QUAN, YU ZHANG, QIAN ZHANG and KAI TAO Department of Plastic Surgery, General Hospital of Shenyang Military Area Command, PLA, Shenyang, Liaoning 110016, P.R. China Received March 10, 2015; Accepted December 18, 2015 DOI: 10.3892/mmr.2016.4802 Abstract. The present study aimed to investigate the for NSCL/P were implicated predominantly in the TGF-β molecular mechanisms underlying non-syndromic cleft lip, signaling pathway, the cell cycle and in viral carcinogenesis. with or without cleft palate (NSCL/P), and the association The TP53, CDK1, SMAD3, PIK3R1 and CASP3 genes were between this disease and cancer. The GSE42589 data set found to be associated, not only with NSCL/P, but also with was downloaded from the Gene Expression Omnibus data- cancer. These results may contribute to a better understanding base, and contained seven dental pulp stem cell samples of the molecular mechanisms of NSCL/P. from children with NSCL/P in the exfoliation period, and six controls. Differentially expressed genes (DEGs) were Introduction screened using the RankProd method, and their potential functions were revealed by pathway enrichment analysis and Non-syndromic cleft lip, with or without cleft palate (NSCL/P) construction of a pathway interaction network. Subsequently, is one of the most common types of congenital defect and cancer genes were obtained from six cancer databases, and affects 3.4-22.9/10,000 individuals worldwide (1). -

Apoptotic Genes As Potential Markers of Metastatic Phenotype in Human Osteosarcoma Cell Lines
17-31 10/12/07 14:53 Page 17 INTERNATIONAL JOURNAL OF ONCOLOGY 32: 17-31, 2008 17 Apoptotic genes as potential markers of metastatic phenotype in human osteosarcoma cell lines CINZIA ZUCCHINI1, ANNA ROCCHI2, MARIA CRISTINA MANARA2, PAOLA DE SANCTIS1, CRISTINA CAPANNI3, MICHELE BIANCHINI1, PAOLO CARINCI1, KATIA SCOTLANDI2 and LUISA VALVASSORI1 1Dipartimento di Istologia, Embriologia e Biologia Applicata, Università di Bologna, Via Belmeloro 8, 40126 Bologna; 2Laboratorio di Ricerca Oncologica, Istituti Ortopedici Rizzoli; 3IGM-CNR, Unit of Bologna, c/o Istituti Ortopedici Rizzoli, Via di Barbiano 1/10, 40136 Bologna, Italy Received May 29, 2007; Accepted July 19, 2007 Abstract. Metastasis is the most frequent cause of death among malignant primitive bone tumor, usually developing in children patients with osteosarcoma. We have previously demonstrated and adolescents, with a high tendency to metastasize (2). in independent experiments that the forced expression of Metastases in osteosarcoma patients spread through peripheral L/B/K ALP and CD99 in U-2 OS osteosarcoma cell lines blood very early and colonize primarily the lung, and later markedly reduces the metastatic ability of these cancer cells. other skeleton districts (3). Since disseminated hidden micro- This behavior makes these cell lines a useful model to assess metastases are present in 80-90% of OS patients at the time the intersection of multiple and independent gene expression of diagnosis, the identification of markers of invasiveness signatures concerning the biological problem of dissemination. and metastasis forms a target of paramount importance in With the aim to characterize a common transcriptional profile planning the treatment of osteosarcoma lesions and enhancing reflecting the essential features of metastatic behavior, we the prognosis. -

GPRC5A Suppresses Protein Synthesis at the Endoplasmic Reticulum to Prevent Radiation-Induced Lung Tumorigenesis
ARTICLE Received 8 Mar 2016 | Accepted 28 Apr 2016 | Published 8 Jun 2016 DOI: 10.1038/ncomms11795 OPEN GPRC5A suppresses protein synthesis at the endoplasmic reticulum to prevent radiation-induced lung tumorigenesis Jian Wang1, Alton B. Farris2, Kaiming Xu1, Ping Wang1, Xiangming Zhang1, Duc M. Duong3, Hong Yi4, Hui-Kuo Shu1, Shi-Yong Sun5 & Ya Wang1 GPRC5A functions as a lung tumour suppressor to prevent spontaneous and environmentally induced lung carcinogenesis; however, the underlying mechanism remains unclear. Here we reveal that GPRC5A at the endoplasmic reticulum (ER) membrane suppresses synthesis of the secreted or membrane-bound proteins including a number of oncogenes, the most important one being Egfr. The ER-located GPRC5A disturbs the assembly of the eIF4F-mediated translation initiation complex on the mRNA cap through directly binding to the eIF4F complex with its two middle extracellular loops. Particularly, suppression of EGFR by GPRC5A contributes significantly to preventing ionizing radiation (IR)-induced lung tumorigenesis. Thus, GPRC5A deletion enhances IR-promoted EGFR expression through an increased translation rate, thereby significantly increasing lung tumour incidence in Gprc5a À / À mice. Our findings indicate that under-expressed GPRC5A during lung tumor- igenesis enhances any transcriptional stimulation through an active translational status, which can be used to control oncogene expression and potentially the resulting related disease. 1 Department of Radiation Oncology, Emory University School of Medicine and the Winship Cancer Institute, Emory University, Atlanta, Georgia 30322, USA. 2 Department of Pathology, Emory University School of Medicine and the Winship Cancer Institute, Emory University, Atlanta, Georgia 30322, USA. 3 Emory Integrated Proteomics Core and Biochemistry Department, Atlanta, Georgia 30322, USA. -

Download Product Insert (PDF)
PRODUCT INFORMATION eIF4E (human recombinant) Item No. 15137 Overview and Properties Synonyms: eIF-4F 25 kDa subunit, Eukaryotic Translation Initiation Factor 4E, mRNA Cap-binding Protein Source: Recombinant protein expressed in E. coli Amino Acids: 2-217 (full length) Uniprot No.: P06730 Molecular Weight: 25.2 kDa Storage: -80°C (as supplied) Stability: ≥1 year Purity: batch specific (≥55% estimated by SDS-PAGE) Supplied in: 20 mM HEPES, pH 7.5, containing 100 mM potassium chloride, 2 mM DTT, and 10% glycerol Protein Concentration: batch specific mg/ml Image 1 2 3 4 250 kDa · · · · · · · 150 kDa · · · · · · · 100 kDa · · · · · · · 75 kDa · · · · · · · 50 kDa · · · · · · · 37 kDa · · · · · · · 25 kDa · · · · · · · 20 kDa · · · · · · · 15 kDa · · · · · · · Lane 1: MW Markers Lane 2: eIF4E (human recombinant) (4 µg) Lane 3: eIF4E (human recombinant) (2 µg) Lane 4: eIF4E (human recombinant) (1 µg) Representaõve gel image shown; actual purity may vary between each batch. WARNING CAYMAN CHEMICAL THIS PRODUCT IS FOR RESEARCH ONLY - NOT FOR HUMAN OR VETERINARY DIAGNOSTIC OR THERAPEUTIC USE. 1180 EAST ELLSWORTH RD SAFETY DATA ANN ARBOR, MI 48108 · USA This material should be considered hazardous until further information becomes available. Do not ingest, inhale, get in eyes, on skin, or on clothing. Wash thoroughly after handling. Before use, the user must review the complete Safety Data Sheet, which has been sent via email to your institution. PHONE: [800] 364-9897 WARRANTY AND LIMITATION OF REMEDY [734] 971-3335 Buyer agrees to purchase the material subject to Cayman’s Terms and Conditions. Complete Terms and Conditions including Warranty and Limitation of Liability information can be found on our website. -

Polyubiquitin Gene Ubb Is Required for Upregulation of Piwi Protein Level During Mouse Testis Development
www.nature.com/cddiscovery ARTICLE OPEN Polyubiquitin gene Ubb is required for upregulation of Piwi protein level during mouse testis development 1,4 2,4 2 1 1 2 ✉ Bitnara Han , Byung-Kwon✉ Jung , So-Hyun Park , Kyu Jin Song , Muhammad Ayaz Anwar , Kwon-Yul Ryu and Kwang Pyo Kim 1,3 © The Author(s) 2021 Testis development, including early embryonic gonad formation and late postnatal spermatogenesis, is essential for the reproduction of higher metazoans to generate fertile gametes, called sperm. We have previously reported that the polyubiquitin gene Ubb is required for fertility in both male and female mice. In particular, the Ubb-null male mice showed an azoospermia phenotype due to arrest of spermatogenesis at the pachytene stage. Here, we analyzed the whole testis proteome at postnatal day 20 to define the molecular mediators of the male-infertility phenotype caused by Ubb knockout. From the identified proteome, 564 proteins were significantly and differentially expressed in Ubb-knockout testes and, among these, 36 downregulated proteins were involved at different stages of spermatogenesis. We also found that levels of piRNA metabolic process-related proteins, including Piwil2 and Tdrd1, were downregulated in Ubb-null testes through functional gene ontology analysis. Further, protein–protein interaction mapping revealed that 24 testis development-related proteins, including Hsp90aa1, Eef1a1, and Pabpc1, were directly influenced by the depletion of ubiquitin. In addition, the reduced mRNA levels of these proteins were observed in Ubb-knockout testes, which closely resembled the global downregulation of piRNA-metabolic gene expression at the transcriptional and post- transcriptional levels. Together with proteomic and transcriptional analyses, our data suggest that Ubb expression is essential for the maintenance of testicular RNA-binding regulators and piRNA-metabolic proteins to complete spermatogenesis in mice. -
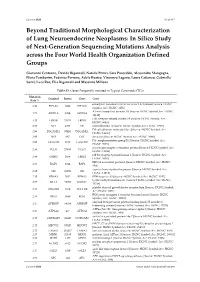
Beyond Traditional Morphological Characterization of Lung
Cancers 2020 S1 of S15 Beyond Traditional Morphological Characterization of Lung Neuroendocrine Neoplasms: In Silico Study of Next-Generation Sequencing Mutations Analysis across the Four World Health Organization Defined Groups Giovanni Centonze, Davide Biganzoli, Natalie Prinzi, Sara Pusceddu, Alessandro Mangogna, Elena Tamborini, Federica Perrone, Adele Busico, Vincenzo Lagano, Laura Cattaneo, Gabriella Sozzi, Luca Roz, Elia Biganzoli and Massimo Milione Table S1. Genes Frequently mutated in Typical Carcinoids (TCs). Mutation Original Entrez Gene Gene Rate % eukaryotic translation initiation factor 1A X-linked [Source: HGNC 4.84 EIF1AX 1964 EIF1AX Symbol; Acc: HGNC: 3250] AT-rich interaction domain 1A [Source: HGNC Symbol;Acc: HGNC: 4.71 ARID1A 8289 ARID1A 11110] LDL receptor related protein 1B [Source: HGNC Symbol; Acc: 4.35 LRP1B 53353 LRP1B HGNC: 6693] 3.53 NF1 4763 NF1 neurofibromin 1 [Source: HGNC Symbol;Acc: HGNC: 7765] DS cell adhesion molecule like 1 [Source: HGNC Symbol; Acc: 2.90 DSCAML1 57453 DSCAML1 HGNC: 14656] 2.90 DST 667 DST dystonin [Source: HGNC Symbol;Acc: HGNC: 1090] FA complementation group D2 [Source: HGNC Symbol; Acc: 2.90 FANCD2 2177 FANCD2 HGNC: 3585] piccolo presynaptic cytomatrix protein [Source: HGNC Symbol; Acc: 2.90 PCLO 27445 PCLO HGNC: 13406] erb-b2 receptor tyrosine kinase 2 [Source: HGNC Symbol; Acc: 2.44 ERBB2 2064 ERBB2 HGNC: 3430] BRCA1 associated protein 1 [Source: HGNC Symbol; Acc: HGNC: 2.35 BAP1 8314 BAP1 950] capicua transcriptional repressor [Source: HGNC Symbol; Acc: 2.35 CIC 23152 CIC HGNC: -
Drosophila and Human Transcriptomic Data Mining Provides Evidence for Therapeutic
Drosophila and human transcriptomic data mining provides evidence for therapeutic mechanism of pentylenetetrazole in Down syndrome Author Abhay Sharma Institute of Genomics and Integrative Biology Council of Scientific and Industrial Research Delhi University Campus, Mall Road Delhi 110007, India Tel: +91-11-27666156, Fax: +91-11-27662407 Email: [email protected] Nature Precedings : hdl:10101/npre.2010.4330.1 Posted 5 Apr 2010 Running head: Pentylenetetrazole mechanism in Down syndrome 1 Abstract Pentylenetetrazole (PTZ) has recently been found to ameliorate cognitive impairment in rodent models of Down syndrome (DS). The mechanism underlying PTZ’s therapeutic effect is however not clear. Microarray profiling has previously reported differential expression of genes in DS. No mammalian transcriptomic data on PTZ treatment however exists. Nevertheless, a Drosophila model inspired by rodent models of PTZ induced kindling plasticity has recently been described. Microarray profiling has shown PTZ’s downregulatory effect on gene expression in fly heads. In a comparative transcriptomics approach, I have analyzed the available microarray data in order to identify potential mechanism of PTZ action in DS. I find that transcriptomic correlates of chronic PTZ in Drosophila and DS counteract each other. A significant enrichment is observed between PTZ downregulated and DS upregulated genes, and a significant depletion between PTZ downregulated and DS dowwnregulated genes. Further, the common genes in PTZ Nature Precedings : hdl:10101/npre.2010.4330.1 Posted 5 Apr 2010 downregulated and DS upregulated sets show enrichment for MAP kinase pathway. My analysis suggests that downregulation of MAP kinase pathway may mediate therapeutic effect of PTZ in DS. Existing evidence implicating MAP kinase pathway in DS supports this observation. -

Relevance of Translation Initiation in Diffuse Glioma Biology and Its
cells Review Relevance of Translation Initiation in Diffuse Glioma Biology and its Therapeutic Potential Digregorio Marina 1, Lombard Arnaud 1,2, Lumapat Paul Noel 1, Scholtes Felix 1,2, Rogister Bernard 1,3 and Coppieters Natacha 1,* 1 Laboratory of Nervous System Disorders and Therapy, GIGA-Neurosciences Research Centre, University of Liège, 4000 Liège, Belgium; [email protected] (D.M.); [email protected] (L.A.); [email protected] (L.P.N.); [email protected] (S.F.); [email protected] (R.B.) 2 Department of Neurosurgery, CHU of Liège, 4000 Liège, Belgium 3 Department of Neurology, CHU of Liège, 4000 Liège, Belgium * Correspondence: [email protected] Received: 18 October 2019; Accepted: 26 November 2019; Published: 29 November 2019 Abstract: Cancer cells are continually exposed to environmental stressors forcing them to adapt their protein production to survive. The translational machinery can be recruited by malignant cells to synthesize proteins required to promote their survival, even in times of high physiological and pathological stress. This phenomenon has been described in several cancers including in gliomas. Abnormal regulation of translation has encouraged the development of new therapeutics targeting the protein synthesis pathway. This approach could be meaningful for glioma given the fact that the median survival following diagnosis of the highest grade of glioma remains short despite current therapy. The identification of new targets for the development of novel therapeutics is therefore needed in order to improve this devastating overall survival rate. This review discusses current literature on translation in gliomas with a focus on the initiation step covering both the cap-dependent and cap-independent modes of initiation.