In Embryonic Development and Gene-Silencing Control
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

EBP1/PA2G4 Rabbit Pab
Leader in Biomolecular Solutions for Life Science EBP1/PA2G4 Rabbit pAb Catalog No.: A5376 Basic Information Background Catalog No. This gene encodes an RNA-binding protein that is involved in growth regulation. This A5376 protein is present in pre-ribosomal ribonucleoprotein complexes and may be involved in ribosome assembly and the regulation of intermediate and late steps of rRNA Observed MW processing. This protein can interact with the cytoplasmic domain of the ErbB3 receptor 44kDa and may contribute to transducing growth regulatory signals. This protein is also a transcriptional co-repressor of androgen receptor-regulated genes and other cell cycle Calculated MW regulatory genes through its interactions with histone deacetylases. This protein has 38kDa/43kDa been implicated in growth inhibition and the induction of differentiation of human cancer cells. Six pseudogenes, located on chromosomes 3, 6, 9, 18, 20 and X, have been Category identified. Primary antibody Applications WB, IHC Cross-Reactivity Human, Mouse, Rat Recommended Dilutions Immunogen Information WB 1:500 - 1:2000 Gene ID Swiss Prot 5036 Q9UQ80 IHC 1:50 - 1:200 Immunogen Recombinant fusion protein containing a sequence corresponding to amino acids 1-394 of human EBP1/PA2G4 (NP_006182.2). Synonyms PA2G4;EBP1;HG4-1;p38-2G4 Contact Product Information www.abclonal.com Source Isotype Purification Rabbit IgG Affinity purification Storage Store at -20℃. Avoid freeze / thaw cycles. Buffer: PBS with 0.02% sodium azide,50% glycerol,pH7.3. Validation Data Western blot analysis of extracts of various cell lines, using EBP1/PA2G4 antibody (A5376) at 1:1000 dilution. Secondary antibody: HRP Goat Anti-Rabbit IgG (H+L) (AS014) at 1:10000 dilution. -

Genome-Wide Analysis of Androgen Receptor Binding and Gene Regulation in Two CWR22-Derived Prostate Cancer Cell Lines
Endocrine-Related Cancer (2010) 17 857–873 Genome-wide analysis of androgen receptor binding and gene regulation in two CWR22-derived prostate cancer cell lines Honglin Chen1, Stephen J Libertini1,4, Michael George1, Satya Dandekar1, Clifford G Tepper 2, Bushra Al-Bataina1, Hsing-Jien Kung2,3, Paramita M Ghosh2,3 and Maria Mudryj1,4 1Department of Medical Microbiology and Immunology, University of California Davis, 3147 Tupper Hall, Davis, California 95616, USA 2Division of Basic Sciences, Department of Biochemistry and Molecular Medicine, Cancer Center and 3Department of Urology, University of California Davis, Sacramento, California 95817, USA 4Veterans Affairs Northern California Health Care System, Mather, California 95655, USA (Correspondence should be addressed to M Mudryj at Department of Medical Microbiology and Immunology, University of California, Davis; Email: [email protected]) Abstract Prostate carcinoma (CaP) is a heterogeneous multifocal disease where gene expression and regulation are altered not only with disease progression but also between metastatic lesions. The androgen receptor (AR) regulates the growth of metastatic CaPs; however, sensitivity to androgen ablation is short lived, yielding to emergence of castrate-resistant CaP (CRCaP). CRCaP prostate cancers continue to express the AR, a pivotal prostate regulator, but it is not known whether the AR targets similar or different genes in different castrate-resistant cells. In this study, we investigated AR binding and AR-dependent transcription in two related castrate-resistant cell lines derived from androgen-dependent CWR22-relapsed tumors: CWR22Rv1 (Rv1) and CWR-R1 (R1). Expression microarray analysis revealed that R1 and Rv1 cells had significantly different gene expression profiles individually and in response to androgen. -

A Simple Differentiation Protocol for Generation of Induced
1 Article, Special Issue "Induced Pluripotent Stem Cells in Neurodegenerative Diseases: Application for Therapy and 2 Disease Modeling" 3 A simple differentiation protocol for generation of 4 induced pluripotent stem cell-derived basal forebrain 5 cholinergic neurons for Alzheimer’s disease and 6 frontotemporal dementia disease modeling 7 Supplemental information 8 Method 1. Reprogramming and characterisation of MBE2960 healthy control iPSC line 9 The iPSCs were generated using skin fibroblasts obtained from subjects over the age of 18 years 10 by episomal method as described [40]. Briefly, reprogramming was performed on passage 8-10 11 fibroblasts by nucleofection (Lonza Amaxa Nucleofector) with episomal vectors expressing 12 OCT4, SOX2, KLF4, L-MYC, LIN28 and shRNA against p53 [41] in feeder- and serum- free 13 conditions using TeSR-E7 medium (Stemcell Technologies). Subsequently, reprogrammed 14 colonies were manually dissected to establish clonal cell lines [42]. Three clones were assessed 15 for pluripotency markers via immunocytochemistry (Figure S1A). The iPSC line was expanded 16 and characterised. Embryoid bodies were obtained as described [43] and using tri-lineage 17 differentiation kit (Stemcell Technologies). Germ layer differentiation was assessed by 18 immunochemistry (Figure S1B). Copy number variation (CNV) analysis of original fibroblasts 19 and iPSCs from MBE2960 (Figure S1C) was performed using Illumina HumanCore Beadchip 20 arrays as we described [40]. CNV analyses were performed using PennCNV and QuantiSNP with 21 default parameter settings [44,45]. Chromosomal aberrations were deemed to involve at least 10 22 contiguous single nucleotide polymorphisms (SNPs) or a genomic region spanning at least 1MB 23 [44,45]. The B allele frequency (BAF) and the log R ratio (LRR) were extracted from 24 GenomeStudio (Illumina) for representation (Figure S1D). -

Diagnosis of FOXG1 Syndrome Caused by Recurrent Balanced Chromosomal Rearrangements: Case Study and Literature Review Connor P
Craig et al. Mol Cytogenet (2020) 13:40 https://doi.org/10.1186/s13039-020-00506-1 CASE REPORT Open Access Diagnosis of FOXG1 syndrome caused by recurrent balanced chromosomal rearrangements: case study and literature review Connor P. Craig1,2, Emily Calamaro3, Chin‑To Fong3, Anwar M. Iqbal1, Alexander R. Paciorkowski3,4,5,6 and Bin Zhang1,3* Abstract Background: The FOXG1 gene plays a vital role in mammalian brain diferentiation and development. Intra‑ and intergenic mutations resulting in loss of function or altered expression of the FOXG1 gene cause FOXG1 syndrome. The hallmarks of this syndrome are severe developmental delay with absent verbal language, post‑natal growth restric‑ tion, post‑natal microcephaly, and a recognizable movement disorder characterized by chorea and dystonia. Case presentation: Here we describe a case of a 7‑year‑old male patient found to have a de novo balanced translo‑ cation between chromosome 3 at band 3q14.1 and chromosome 14 at band 14q12 via G‑banding chromosome and Fluorescence In Situ Hybridization (FISH) analyses. This rearrangement disrupts the proximity of FOXG1 to a previously described smallest region of deletion overlap (SRO), likely resulting in haploinsufciency. Conclusions: This case adds to the growing body of literature implicating chromosomal structural variants in the manifestation of this disorder and highlights the vital role of cis‑acting regulatory elements in the normal expression of this gene. Finally, we propose a protocol for refex FISH analysis to improve diagnostic efciency for patients with suspected FOXG1 syndrome. Keywords: FOXG1, Haploinsufciency, Postnatal microcephaly, FISH, Enhancer, Chromosomal rearrangement, Diagnosis Introduction brain development, with high levels of expression in the Te Forkhead Box G1 (FOXG1) gene [OMIM: 164874], developing fetal telencephalon [1–4]. -
Notch Signal Controls Several Steps of Inner Ear Development Norio Yamamoto and Matthew W
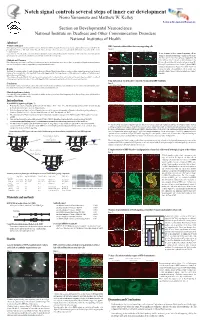
NIDCD Notch signal controls several steps of inner ear development Norio Yamamoto and Matthew W. Kelley Section on Developmental Neuroscience Section on Developmental Neuroscience National Institute on Deafness and Other Communication Disorders National Institutes of Health Abstract Problem addressed RBP-J mutant cochleae did not have any supporting cells Notch signaling has been reported to contribute to inner ear development, however, its specific functions remain unclear, partly because of discrepancies between the phenotypes of mutant mice with single deletion of specific Notch related genes. These discrepancies are probably due to functional compensation by other Notch Figure 4 receptors or ligands. Foxg1 Cre;RBP-J floxed/+ Foxg1 Cre;RBP-J floxed/floxed A B C D To determine the effects of complete elimination of Notch signaling, we used a conditional knockout of the Rbpsuh gene. RBP-J protein is a critical transcriptional p27 Phalloidin p27 Phalloidin To test if mutant cochleae contained supporting cells we co-activator for all Notch molecules and thus deletion of this protein inhibits all Notch signaling. examined expression of supporting cell markers such as p27 and Prox1. On E17.5 p27 was expressed in supporting cells Methods and Measures ***** ***** under or between hair cells such as inner pharyngeal cells, Floxed Rbpsuh mice were crossed with Foxg1-Cre knock-in mice to delete the Rbpsuh gene in the inner ear. Inner ear phenotypes in Rbpsuh conditional knockout Deiter's cells and pillar cells (asterisks in figure 4 A and B). mice were determined at various developmental stages using immunohistochemistry. But no p27 expression was detected in those regions of RBP-J E Prox1 F Phalloidin G Prox1 H Phalloidin conditional knockout mice (Figure 4 C and D). -

PA2G4 Antibody (F54781)
PA2G4 Antibody / EBP1 / ErbB3-binding protein 1 (F54781) Catalog No. Formulation Size F54781-0.4ML In 1X PBS, pH 7.4, with 0.09% sodium azide 0.4 ml F54781-0.08ML In 1X PBS, pH 7.4, with 0.09% sodium azide 0.08 ml Bulk quote request Availability 1-3 business days Species Reactivity Human Format Purified Clonality Polyclonal (rabbit origin) Isotype Rabbit Ig Purity Purified UniProt Q9UQ80 Localization Cytoplasmic, nuclear Applications Immunofluorescence : 1:25 Flow cytometry : 1:25 (1x10e6 cells) Immunohistochemistry (FFPE) : 1:25 Western blot : 1:500-1:1000 Limitations This PA2G4 antibody is available for research use only. IHC testing of FFPE human lung carcinoma tissue with PA2G4 antibody. HIER: steam section in pH6 citrate buffer for 20 min and allow to cool prior to staining. Immunofluorescent staining of fixed and permeabilized human U-251 cells with PA2G4 antibody (green), DAPI nuclear stain (blue) and anti-Actin (red). Western blot testing of human 1) A431 and 2) Jurkat cell lysate with PA2G4 antibody. Predicted molecular weight ~44 kDa. Flow cytometry testing of human A2058 cells with PA2G4 antibody; Blue=isotype control, Green= PA2G4 antibody. Description PA2G4 is an RNA-binding protein that is involved in growth regulation. This protein is present in pre-ribosomal ribonucleoprotein complexes and may be involved in ribosome assembly and the regulation of intermediate and late steps of rRNA processing. The protein can interact with the cytoplasmic domain of the ErbB3 receptor and may contribute to transducing growth regulatory signals. It is also a transcriptional co-repressor of androgen receptor-regulated genes and other cell cycle regulatory genes through its interactions with histone deacetylases. -

A Dissertation Entitled the Androgen Receptor
A Dissertation entitled The Androgen Receptor as a Transcriptional Co-activator: Implications in the Growth and Progression of Prostate Cancer By Mesfin Gonit Submitted to the Graduate Faculty as partial fulfillment of the requirements for the PhD Degree in Biomedical science Dr. Manohar Ratnam, Committee Chair Dr. Lirim Shemshedini, Committee Member Dr. Robert Trumbly, Committee Member Dr. Edwin Sanchez, Committee Member Dr. Beata Lecka -Czernik, Committee Member Dr. Patricia R. Komuniecki, Dean College of Graduate Studies The University of Toledo August 2011 Copyright 2011, Mesfin Gonit This document is copyrighted material. Under copyright law, no parts of this document may be reproduced without the expressed permission of the author. An Abstract of The Androgen Receptor as a Transcriptional Co-activator: Implications in the Growth and Progression of Prostate Cancer By Mesfin Gonit As partial fulfillment of the requirements for the PhD Degree in Biomedical science The University of Toledo August 2011 Prostate cancer depends on the androgen receptor (AR) for growth and survival even in the absence of androgen. In the classical models of gene activation by AR, ligand activated AR signals through binding to the androgen response elements (AREs) in the target gene promoter/enhancer. In the present study the role of AREs in the androgen- independent transcriptional signaling was investigated using LP50 cells, derived from parental LNCaP cells through extended passage in vitro. LP50 cells reflected the signature gene overexpression profile of advanced clinical prostate tumors. The growth of LP50 cells was profoundly dependent on nuclear localized AR but was independent of androgen. Nevertheless, in these cells AR was unable to bind to AREs in the absence of androgen. -

Downloaded from Bioscientifica.Com at 09/26/2021 08:38:14PM Via Free Access
1 185 L C Gregory, P Gergics, OTX2 mutations in congenital 185:1 121–135 Clinical Study M Nakaguma and others hypopituitarism The phenotypic spectrum associated with OTX2 mutations in humans Louise C Gregory 1,*, Peter Gergics2,* , Marilena Nakaguma3,* , Hironori Bando2 , Giuseppa Patti1,4,5, Mark J McCabe1, Qing Fang 2, Qianyi Ma2 , Ayse Bilge Ozel2 , Jun Z Li2 , Michele Moreira Poina3, Alexander A L Jorge 3, Anna F Figueredo Benedetti3, Antonio M Lerario3, Ivo J P Arnhold3 , Berenice B Mendonca3 , Mohamad Maghnie4,5, Sally A Camper2 , Luciani R S Carvalho3 and Mehul T Dattani1 1Section of Molecular Basis of Rare Disease, Genetics and Genomic Medicine Research & Teaching Department, UCL Great Ormond Street Institute of Child Health, London, UK, 2Department of Human Genetics, University of Michigan, Correspondence Ann Arbor, Michigan, USA, 3Developmental Endocrinology Unit, Hospital das Clinicas da Faculdade de Medicina da should be addressed Universidade de São Paulo, São Paulo, Brazil, 4Department of Pediatrics, IRCCS Istituto Giannina Gaslini and to S A Camper or M T 5Department of Neuroscience, Rehabilitation, Ophthalmology, Genetics, Maternal and Child Health, University of Dattani Genova, Genova, Italy Email *(L C Gregory, P Gergics and M Nakaguma contributed equally to this work) [email protected] or [email protected] Abstract Objective: The transcription factor OTX2 is implicated in ocular, craniofacial, and pituitary development. Design: We aimed to establish the contribution of OTX2 mutations in congenital hypopituitarism patients with/without eye abnormalities, study functional consequences, and establish OTX2 expression in the human brain, with a view to investigate the mechanism of action. Methods: We screened patients from the UK (n = 103), international centres (n = 24), and Brazil (n = 282); 145 were within the septo-optic dysplasia spectrum, and 264 had no eye phenotype. -
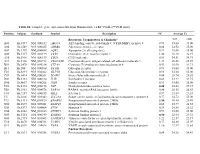
Table S1. Complete Gene Expression Data from Human Diabetes RT² Profiler™ PCR Array Receptors, Transporters & Channels* A
Table S1. Complete gene expression data from Human Diabetes RT² Profiler™ PCR Array Position Unigene GenBank Symbol Description FC Average Ct Receptors, Transporters & Channels* NGT GDM A01 Hs,5447 NM_000352 ABCC8 ATP-binding cassette, sub-family C (CFTR/MRP), member 8 0.93 35.00 35.00 A04 0Hs,2549 NM_000025 ADRB3 Adrenergic, beta-3-, receptor 0.88 34.92 35.00 A07 Hs,1307 NM_000486 AQP2 Aquaporin 2 (collecting duct) 0.93 35.00 35.00 A09 30Hs,5117 NM_001123 CCR2 Chemokine (C-C motif) receptor 2 1.00 26.28 26.17 A10 94Hs,5916 396NM_006139 CD28 CD28 molecule 0.81 34.51 34.71 A11 29Hs,5126 NM_001712 CEACAM1 Carcinoembryonic antigen-related cell adhesion molecule 1 1.31 26.08 25.59 B01 82Hs,2478 NM_005214 CTLA4 (biliaryCytotoxic glycoprotein) T-lymphocyte -associated protein 4 0.53 30.90 31.71 B11 24Hs,208 NM_000160 GCGR Glucagon receptor 0.93 35.00 35.00 C01 Hs,3891 NM_002062 GLP1R Glucagon-like peptide 1 receptor 0.93 35.00 35.00 C07 03Hs,6434 NM_000201 ICAM1 Intercellular adhesion molecule 1 0.84 28.74 28.89 D02 47Hs,5134 NM_000418 IL4R Interleukin 4 receptor 0.64 34.22 34.75 D06 57Hs,4657 NM_000208 INSR Insulin receptor 0.93 35.00 35.00 E05 44Hs,4312 NM_006178 NSF N-ethylmaleimide-sensitive factor 0.48 28.42 29.37 F08 79Hs,2961 NM_004578 RAB4A RAB4A, member RAS oncogene family 0.88 20.55 20.63 F10 69Hs,7287 NM_000655 SELL Selectin L 0.97 23.89 23.83 F11 56Hs,3806 NM_001042 SLC2A4 Solute carrier family 2 (facilitated glucose transporter), member 4 0.77 34.72 35.00 F12 91Hs,5111 NM_003825 SNAP23 Synaptosomal-associated protein, 23kDa 3.90 -

Open Data for Differential Network Analysis in Glioma
International Journal of Molecular Sciences Article Open Data for Differential Network Analysis in Glioma , Claire Jean-Quartier * y , Fleur Jeanquartier y and Andreas Holzinger Holzinger Group HCI-KDD, Institute for Medical Informatics, Statistics and Documentation, Medical University Graz, Auenbruggerplatz 2/V, 8036 Graz, Austria; [email protected] (F.J.); [email protected] (A.H.) * Correspondence: [email protected] These authors contributed equally to this work. y Received: 27 October 2019; Accepted: 3 January 2020; Published: 15 January 2020 Abstract: The complexity of cancer diseases demands bioinformatic techniques and translational research based on big data and personalized medicine. Open data enables researchers to accelerate cancer studies, save resources and foster collaboration. Several tools and programming approaches are available for analyzing data, including annotation, clustering, comparison and extrapolation, merging, enrichment, functional association and statistics. We exploit openly available data via cancer gene expression analysis, we apply refinement as well as enrichment analysis via gene ontology and conclude with graph-based visualization of involved protein interaction networks as a basis for signaling. The different databases allowed for the construction of huge networks or specified ones consisting of high-confidence interactions only. Several genes associated to glioma were isolated via a network analysis from top hub nodes as well as from an outlier analysis. The latter approach highlights a mitogen-activated protein kinase next to a member of histondeacetylases and a protein phosphatase as genes uncommonly associated with glioma. Cluster analysis from top hub nodes lists several identified glioma-associated gene products to function within protein complexes, including epidermal growth factors as well as cell cycle proteins or RAS proto-oncogenes. -

14Q12q13.3 Deletion Diagnosed Using Chromo Somal Microarray
Case Report Neonatal Med 2020 November;27(4):207-213 https://doi.org/10.5385/nm.2020.27.4.207 pISSN 2287-9412 . eISSN 2287-9803 14q12q13.3 Deletion Diagnosed Using Chromo somal Microarray Analysis in an Infant Showing Seizures, Hypoplasia of the Corpus Callosum, and Developmental Delay Jae Hyuk Kwon, MD1, Young Hwa Song, MD1, Jung Min Yoon, MD1, Eun Jung Cheon, MD1, Kyung Ok Ko, MD1, Jae Woo Lim, MD1, and Hyon J. Kim, MD, FACMG2 Departments of 1Pediatrics and 2Medical Genetics, Konyang University College of Medicine, Daejeon, Korea ABSTRACT Received: 18 August 2020 Revised: 25 September 2020 14q12q13.3 Deletion is a rare microdeletion syndrome associated with neurodevelop Accepted: 6 October 2020 mental delay, failure to thrive, seizures, and abnormal brain development. Symptoms Correspondence to: Hyon J. Kim, MD vary depending on the sites of gene deletion, and establishing the diagnosis is often Department of Medical Genetics, difficult, as the condition cannot be detected with routine chromosome analysis. In Konyang University College of Medi this report, we present a patient with intrauterine growth retardation, microcephaly, cine, 158 Gwanjeodongro, Seogu, muscle weakness, seizures, and hypoplasia of the corpus callosum who underwent Daejeon 35365, Korea diagnostic tests, including karyotyping in the neonatal period without leading to a Tel: +82426009230 specific diagnosis. The patient was confirmed with a serious developmental disorder, Fax: +82426009090 and a chromosomal microarray analysis was performed at 8 months of age, revealing Email: [email protected] a 14q12q13.3 deletion. In this case, the condition was diagnosed in early infancy, in contrast to previously reported cases, and the patient had diverse and severe symptoms. -

PA2G4 Antibody (Center R243) Blocking Peptide Synthetic Peptide Catalog # Bp2848d
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 PA2G4 Antibody (Center R243) Blocking Peptide Synthetic peptide Catalog # BP2848d Specification PA2G4 Antibody (Center R243) Blocking PA2G4 Antibody (Center R243) Blocking Peptide - Peptide - Background Product Information PA2G4 is an RNA-binding protein that is Primary Accession Q9UQ80 involved in growth regulation. This protein is present in pre-ribosomal ribonucleoprotein complexes and may be involved in ribosome PA2G4 Antibody (Center R243) Blocking Peptide - Additional Information assembly and the regulation of intermediate and late steps of rRNA processing. The protein can interact with the cytoplasmic domain of Gene ID 5036 the ErbB3 receptor and may contribute to transducing growth regulatory signals. It is also Other Names a transcriptional co-repressor of androgen Proliferation-associated protein 2G4, Cell receptor-regulated genes and other cell cycle cycle protein p38-2G4 homolog, hG4-1, regulatory genes through its interactions with ErbB3-binding protein 1, PA2G4, EBP1 histone deacetylases. It has been implicated in Target/Specificity growth inhibition and the induction of The synthetic peptide sequence used to differentiation of human cancer cells. generate the antibody <a href=/products/AP2848d>AP2848d</a> PA2G4 Antibody (Center R243) Blocking was selected from the Center region of Peptide - References human PA2G4. A 10 to 100 fold molar excess to antibody is recommended. Zhang,Y., Mol. Cancer Ther. 7 (10), 3176-3186 Precise conditions should be optimized for a (2008)Zhang,Y., Cancer Lett. 265 (2), 298-306 particular assay. (2008)Okada,M., J. Biol. Chem. 282 (50), 36744-36754 (2007) Format Peptides are lyophilized in a solid powder format.