Improving Tissue Differentiation in Cerebral Organoids
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

BR27069-Stemdiff™ Cerebral Organoid
CEREBRAL ORGANOIDS STEMdiff™ Cerebral Organoid Kit Scientists Helping Scientists™ | WWW.STEMCELL.COM TABLE OF CONTENTS 3 Introduction 4 Cerebral Organoid Formation 5 Cerebral Organoid Characterization 7 Performance of STEMdiffTM Cerebral Organoid Kit vs. Unqualified Reagents 7 Product Information 7 References Introduction For the Culture and Maturation of Why Use the STEMdiff™ Cerebral Organoid Kit? Human Cerebral Organoids PHYSIOLOGICAL. Three-dimensional in vitro system The metazoan brain is a highly complex and organized structure. recapitulates the developmental processes and Two-dimensional (2D) neural cultures derived from human organization of the developing brain. pluripotent stem cells (hPSCs) are useful models to study the nervous INNOVATIVE. Serum-free human pluripotent stem system, but they are limited in their capacity to recapitulate the cell-based model enables the study of development and complex organization of brain tissues. disease processes. hPSC-derived cerebral organoids are three-dimensional (3D) in vitro OPTIMIZED. Formulation is optimized for increased culture systems that recapitulate the developmental processes and efficiency of organoid formation. organization of the developing human brain. They provide a physiologically relevant in vitro model for the RELIABLE. Rigorous raw material screening and study of neurological development and disease processes that extensive quality control testing ensure reproducibility are unique to the human nervous system. Cerebral organoids and minimal lot-to-lot variability. have important applications in studying human brain development SIMPLE. Convenient format and easy-to-use protocol. and neurological disorders such as autism, schizophrenia or brain defects caused by Zika virus infection. The STEMdiff™ Cerebral Organoid Kit is a serum-free culture system that is designed to generate cerebral organoids from human embryonic stem (ES) cells observed during in vivo human brain development. -

A Simple Differentiation Protocol for Generation of Induced
1 Article, Special Issue "Induced Pluripotent Stem Cells in Neurodegenerative Diseases: Application for Therapy and 2 Disease Modeling" 3 A simple differentiation protocol for generation of 4 induced pluripotent stem cell-derived basal forebrain 5 cholinergic neurons for Alzheimer’s disease and 6 frontotemporal dementia disease modeling 7 Supplemental information 8 Method 1. Reprogramming and characterisation of MBE2960 healthy control iPSC line 9 The iPSCs were generated using skin fibroblasts obtained from subjects over the age of 18 years 10 by episomal method as described [40]. Briefly, reprogramming was performed on passage 8-10 11 fibroblasts by nucleofection (Lonza Amaxa Nucleofector) with episomal vectors expressing 12 OCT4, SOX2, KLF4, L-MYC, LIN28 and shRNA against p53 [41] in feeder- and serum- free 13 conditions using TeSR-E7 medium (Stemcell Technologies). Subsequently, reprogrammed 14 colonies were manually dissected to establish clonal cell lines [42]. Three clones were assessed 15 for pluripotency markers via immunocytochemistry (Figure S1A). The iPSC line was expanded 16 and characterised. Embryoid bodies were obtained as described [43] and using tri-lineage 17 differentiation kit (Stemcell Technologies). Germ layer differentiation was assessed by 18 immunochemistry (Figure S1B). Copy number variation (CNV) analysis of original fibroblasts 19 and iPSCs from MBE2960 (Figure S1C) was performed using Illumina HumanCore Beadchip 20 arrays as we described [40]. CNV analyses were performed using PennCNV and QuantiSNP with 21 default parameter settings [44,45]. Chromosomal aberrations were deemed to involve at least 10 22 contiguous single nucleotide polymorphisms (SNPs) or a genomic region spanning at least 1MB 23 [44,45]. The B allele frequency (BAF) and the log R ratio (LRR) were extracted from 24 GenomeStudio (Illumina) for representation (Figure S1D). -

Diagnosis of FOXG1 Syndrome Caused by Recurrent Balanced Chromosomal Rearrangements: Case Study and Literature Review Connor P
Craig et al. Mol Cytogenet (2020) 13:40 https://doi.org/10.1186/s13039-020-00506-1 CASE REPORT Open Access Diagnosis of FOXG1 syndrome caused by recurrent balanced chromosomal rearrangements: case study and literature review Connor P. Craig1,2, Emily Calamaro3, Chin‑To Fong3, Anwar M. Iqbal1, Alexander R. Paciorkowski3,4,5,6 and Bin Zhang1,3* Abstract Background: The FOXG1 gene plays a vital role in mammalian brain diferentiation and development. Intra‑ and intergenic mutations resulting in loss of function or altered expression of the FOXG1 gene cause FOXG1 syndrome. The hallmarks of this syndrome are severe developmental delay with absent verbal language, post‑natal growth restric‑ tion, post‑natal microcephaly, and a recognizable movement disorder characterized by chorea and dystonia. Case presentation: Here we describe a case of a 7‑year‑old male patient found to have a de novo balanced translo‑ cation between chromosome 3 at band 3q14.1 and chromosome 14 at band 14q12 via G‑banding chromosome and Fluorescence In Situ Hybridization (FISH) analyses. This rearrangement disrupts the proximity of FOXG1 to a previously described smallest region of deletion overlap (SRO), likely resulting in haploinsufciency. Conclusions: This case adds to the growing body of literature implicating chromosomal structural variants in the manifestation of this disorder and highlights the vital role of cis‑acting regulatory elements in the normal expression of this gene. Finally, we propose a protocol for refex FISH analysis to improve diagnostic efciency for patients with suspected FOXG1 syndrome. Keywords: FOXG1, Haploinsufciency, Postnatal microcephaly, FISH, Enhancer, Chromosomal rearrangement, Diagnosis Introduction brain development, with high levels of expression in the Te Forkhead Box G1 (FOXG1) gene [OMIM: 164874], developing fetal telencephalon [1–4]. -

Optimization of Cerebral Organoid Culture from Embr
CALIFORNIA STATE UNIVERSITY, NORTHRIDGE Growing Brains to Study Development: Optimization of Cerebral Organoid Culture from Embryonic Stem Cells A thesis submitted in partial fulfillment of the requirements For the degree of Master of Science in Biology By Jessie Erin Buth December 2015 The thesis of Jessie Erin Buth is approved: __________________________________ ____________________________________ Dr. Aida Metzenberg Date __________________________________ ____________________________________ Dr. Randy Cohen Date __________________________________ ____________________________________ Dr. Cindy Malone, Chair Date California State University, Northridge ii Table of Contents Signature Page ii List of Figures iii Abstract vi Chapter 1: Introduction 1 Chapter 2: Methods 15 Chapter 3: Results 23 Chapter 4: Discussion 50 References 55 Appendix: Supplemental Methods 59 List of Figures Figure 1: Human organoid protocol (Cell line H9) and morphology at various time points. 23 Figure 2: Review of cortex development. 24 Figure 3: V-bottomed 96-well plates improve aggregate formation and generates a thick neuroepithelial layer on day 18 compared to U-bottomed 96-well plates. 25 Figure 4: Plating at 9,000 cells per well on day 0 generates thickest continuous neuroepithelial layer compared to other cell numbers. 26 Figure 5: Addition of the BMP inhibitor (LDN193189) does not improve the efficiency of producing FOXG1+ cortical progenitors and inhibits formation of continuous N-cadherin+ apical membrane. 27 Figure 6: Fold change in gene expression of cortical markers increases as stem cell markers decrease in human cortical organoids over time by qPCR. 28 Figure 7: Human embryonic stem cells efficiently form cortical progenitors with 81.7% of total live cells per organoid positive for cortical marker FOXG1 at day 18. -
Notch Signal Controls Several Steps of Inner Ear Development Norio Yamamoto and Matthew W
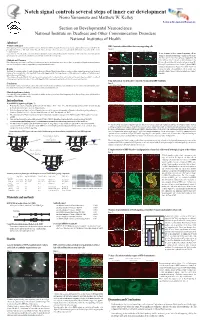
NIDCD Notch signal controls several steps of inner ear development Norio Yamamoto and Matthew W. Kelley Section on Developmental Neuroscience Section on Developmental Neuroscience National Institute on Deafness and Other Communication Disorders National Institutes of Health Abstract Problem addressed RBP-J mutant cochleae did not have any supporting cells Notch signaling has been reported to contribute to inner ear development, however, its specific functions remain unclear, partly because of discrepancies between the phenotypes of mutant mice with single deletion of specific Notch related genes. These discrepancies are probably due to functional compensation by other Notch Figure 4 receptors or ligands. Foxg1 Cre;RBP-J floxed/+ Foxg1 Cre;RBP-J floxed/floxed A B C D To determine the effects of complete elimination of Notch signaling, we used a conditional knockout of the Rbpsuh gene. RBP-J protein is a critical transcriptional p27 Phalloidin p27 Phalloidin To test if mutant cochleae contained supporting cells we co-activator for all Notch molecules and thus deletion of this protein inhibits all Notch signaling. examined expression of supporting cell markers such as p27 and Prox1. On E17.5 p27 was expressed in supporting cells Methods and Measures ***** ***** under or between hair cells such as inner pharyngeal cells, Floxed Rbpsuh mice were crossed with Foxg1-Cre knock-in mice to delete the Rbpsuh gene in the inner ear. Inner ear phenotypes in Rbpsuh conditional knockout Deiter's cells and pillar cells (asterisks in figure 4 A and B). mice were determined at various developmental stages using immunohistochemistry. But no p27 expression was detected in those regions of RBP-J E Prox1 F Phalloidin G Prox1 H Phalloidin conditional knockout mice (Figure 4 C and D). -

Downloaded from Bioscientifica.Com at 09/26/2021 08:38:14PM Via Free Access
1 185 L C Gregory, P Gergics, OTX2 mutations in congenital 185:1 121–135 Clinical Study M Nakaguma and others hypopituitarism The phenotypic spectrum associated with OTX2 mutations in humans Louise C Gregory 1,*, Peter Gergics2,* , Marilena Nakaguma3,* , Hironori Bando2 , Giuseppa Patti1,4,5, Mark J McCabe1, Qing Fang 2, Qianyi Ma2 , Ayse Bilge Ozel2 , Jun Z Li2 , Michele Moreira Poina3, Alexander A L Jorge 3, Anna F Figueredo Benedetti3, Antonio M Lerario3, Ivo J P Arnhold3 , Berenice B Mendonca3 , Mohamad Maghnie4,5, Sally A Camper2 , Luciani R S Carvalho3 and Mehul T Dattani1 1Section of Molecular Basis of Rare Disease, Genetics and Genomic Medicine Research & Teaching Department, UCL Great Ormond Street Institute of Child Health, London, UK, 2Department of Human Genetics, University of Michigan, Correspondence Ann Arbor, Michigan, USA, 3Developmental Endocrinology Unit, Hospital das Clinicas da Faculdade de Medicina da should be addressed Universidade de São Paulo, São Paulo, Brazil, 4Department of Pediatrics, IRCCS Istituto Giannina Gaslini and to S A Camper or M T 5Department of Neuroscience, Rehabilitation, Ophthalmology, Genetics, Maternal and Child Health, University of Dattani Genova, Genova, Italy Email *(L C Gregory, P Gergics and M Nakaguma contributed equally to this work) [email protected] or [email protected] Abstract Objective: The transcription factor OTX2 is implicated in ocular, craniofacial, and pituitary development. Design: We aimed to establish the contribution of OTX2 mutations in congenital hypopituitarism patients with/without eye abnormalities, study functional consequences, and establish OTX2 expression in the human brain, with a view to investigate the mechanism of action. Methods: We screened patients from the UK (n = 103), international centres (n = 24), and Brazil (n = 282); 145 were within the septo-optic dysplasia spectrum, and 264 had no eye phenotype. -
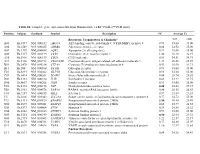
Table S1. Complete Gene Expression Data from Human Diabetes RT² Profiler™ PCR Array Receptors, Transporters & Channels* A
Table S1. Complete gene expression data from Human Diabetes RT² Profiler™ PCR Array Position Unigene GenBank Symbol Description FC Average Ct Receptors, Transporters & Channels* NGT GDM A01 Hs,5447 NM_000352 ABCC8 ATP-binding cassette, sub-family C (CFTR/MRP), member 8 0.93 35.00 35.00 A04 0Hs,2549 NM_000025 ADRB3 Adrenergic, beta-3-, receptor 0.88 34.92 35.00 A07 Hs,1307 NM_000486 AQP2 Aquaporin 2 (collecting duct) 0.93 35.00 35.00 A09 30Hs,5117 NM_001123 CCR2 Chemokine (C-C motif) receptor 2 1.00 26.28 26.17 A10 94Hs,5916 396NM_006139 CD28 CD28 molecule 0.81 34.51 34.71 A11 29Hs,5126 NM_001712 CEACAM1 Carcinoembryonic antigen-related cell adhesion molecule 1 1.31 26.08 25.59 B01 82Hs,2478 NM_005214 CTLA4 (biliaryCytotoxic glycoprotein) T-lymphocyte -associated protein 4 0.53 30.90 31.71 B11 24Hs,208 NM_000160 GCGR Glucagon receptor 0.93 35.00 35.00 C01 Hs,3891 NM_002062 GLP1R Glucagon-like peptide 1 receptor 0.93 35.00 35.00 C07 03Hs,6434 NM_000201 ICAM1 Intercellular adhesion molecule 1 0.84 28.74 28.89 D02 47Hs,5134 NM_000418 IL4R Interleukin 4 receptor 0.64 34.22 34.75 D06 57Hs,4657 NM_000208 INSR Insulin receptor 0.93 35.00 35.00 E05 44Hs,4312 NM_006178 NSF N-ethylmaleimide-sensitive factor 0.48 28.42 29.37 F08 79Hs,2961 NM_004578 RAB4A RAB4A, member RAS oncogene family 0.88 20.55 20.63 F10 69Hs,7287 NM_000655 SELL Selectin L 0.97 23.89 23.83 F11 56Hs,3806 NM_001042 SLC2A4 Solute carrier family 2 (facilitated glucose transporter), member 4 0.77 34.72 35.00 F12 91Hs,5111 NM_003825 SNAP23 Synaptosomal-associated protein, 23kDa 3.90 -

Modelling Heme-Mediated Brain Injury Associated with Cerebral Malaria In
www.nature.com/scientificreports OPEN Modelling heme-mediated brain injury associated with cerebral malaria in human brain cortical organoids Adriana Harbuzariu1*, Sidney Pitts1, Juan Carlos Cespedes1, Keri Oxendine Harp1, Annette Nti1, Andrew P. Shaw2, Mingli Liu1 & Jonathan K. Stiles1* Human cerebral malaria (HCM), a severe encephalopathy associated with Plasmodium falciparum infection, has a 20–30% mortality rate and predominantly afects African children. The mechanisms mediating HCM-associated brain injury are difcult to study in human subjects, highlighting the urgent need for non-invasive ex vivo human models. HCM elevates the systemic levels of free heme, which damages the blood-brain barrier and neurons in distinct regions of the brain. We determined the efects of heme on induced pluripotent stem cells (iPSCs) and a three-dimensional cortical organoid system and assessed apoptosis and diferentiation. We evaluated biomarkers associated with heme-induced brain injury, including a pro-infammatory chemokine, CXCL-10, and its receptor, CXCR3, brain-derived neurotrophic factor (BDNF) and a receptor tyrosine-protein kinase, ERBB4, in the organoids. We then tested the neuroprotective efect of neuregulin-1 (NRG-1) against heme treatment in organoids. Neural stem and mature cells diferentially expressed CXCL-10, CXCR3, BDNF and ERBB4 in the developing organoids and in response to heme-induced neuronal injury. The organoids underwent apoptosis and structural changes that were attenuated by NRG-1. Thus, cortical organoids can be used to model heme-induced cortical brain injury associated with HCM pathogenesis as well as for testing agents that reduce brain injury and neurological sequelae. Brain organoids are self-assembled three-dimensional (3D) aggregates derived from pluripotent stem cells (iPSCs) with cell types and formations that mimic the embryonic human brain1–7. -

Stem Cells, Genome Editing and the Path to Translational Medicine
Stem Cells, Genome Editing, and the Path to Translational Medicine The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation Soldner, Frank and Rudolf Jaenisch. "Stem Cells, Genome Editing, and the Path to Translational Medicine." Cell 175, 3 (October 2018): P615-632 © 2018 Elsevier Inc As Published http://dx.doi.org/10.1016/j.cell.2018.09.010 Publisher Elsevier BV Version Author's final manuscript Citable link https://hdl.handle.net/1721.1/125973 Terms of Use Creative Commons Attribution-NonCommercial-NoDerivs License Detailed Terms http://creativecommons.org/licenses/by-nc-nd/4.0/ HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Cell. Author Manuscript Author manuscript; Manuscript Author available in PMC 2019 October 18. Published in final edited form as: Cell. 2018 October 18; 175(3): 615–632. doi:10.1016/j.cell.2018.09.010. Stem cells, genome editing and the path to translational medicine Frank Soldner1 and Rudolf Jaenisch1,2,3 1The Whitehead Institute, 455 Main Street, Cambridge, MA 02142, USA 2Department of Biology, Massachusetts Institute of Technology, 31 Ames Street, Cambridge, MA 02139, USA Summary The derivation of human embryonic stem cells (hESCs) and the stunning discovery that somatic cells can be reprogrammed into human induced pluripotent stem cells (hiPSCs) holds the promise to revolutionize biomedical research and regenerative medicine. In this review, we focus on disorders of the central nervous system and explore how advances in human pluripotent stem cells (hPSCs) coincide with evolutions in genome engineering and genomic technologies to provide realistic opportunities to tackle some of the most devastating complex disorders. -

Building Three-Dimensional Human Brain Organoids Sergiu P
Building three-dimensional human brain organoids Sergiu P. Paşca The organogenesis of the human central nervous system is an intricately culture methods, to self-organize into brain spheroids or organoids. orchestrated series of events that occurs over several months and These organoid cultures can be derived from any individual, can be ultimately gives rise to the circuits underlying cognition and behavior. guided to resemble specific brain regions, and can be employed to model There is a pressing need for developing reliable, realistic, and complex cell-cell interactions in assembloids and to build human circuits. personalized in vitro models of the human brain to advance our This emerging technology, in combination with bioengineering and other understanding of neural development, evolution, and disease. Pluripotent state-of-the-art methods for probing and manipulating neural tissue, has stem cells have the remarkable ability to dierentiate in vitro into any of the potential to bring insights into human brain organogenesis and the the germ layers and, with the advent of three-dimensional (3D) cell pathogenesis of neurological and psychiatric disorders. Brain organogenesis in vitro and in vivo Brain organogenesis in vitro and in vivo Brain assembloids Methods for generating neural cells in vitro aim to recapitulate Spinal Forebrain Forebrain Blastocyst Cerebral Dorsal Pallial-subpallial assembloid key stages of in vivo brain organogenesis. Folding of the Neural Neural cord Paired recordingOptogenetics Pharmacology fold crest cortex Glutamate uncaging ectoderm-derived neural plate gives rise to the neural tube, Embryonic Yo lk sac Region 1 Region 2Region 3 which becomes enlarged on the anterior side to form the stem cell Neural groove Ventral Inner cell forebrain in the central nervous system (CNS). -

Organoid Based Personalized Medicine: from Bench to Bedside Yaqi Li1,2†, Peiyuan Tang3†, Sanjun Cai1,2, Junjie Peng1,2 and Guoqiang Hua3,4*
Li et al. Cell Regeneration (2020) 9:21 https://doi.org/10.1186/s13619-020-00059-z REVIEW Open Access Organoid based personalized medicine: from bench to bedside Yaqi Li1,2†, Peiyuan Tang3†, Sanjun Cai1,2, Junjie Peng1,2 and Guoqiang Hua3,4* Abstract Three-dimensional cultured organoids have become a powerful in vitro research tool that preserves genetic, phenotypic and behavioral trait of in vivo organs, which can be established from both pluripotent stem cells and adult stem cells. Organoids derived from adult stem cells can be established directly from diseased epithelium and matched normal tissues, and organoids can also be genetically manipulated by CRISPR-Cas9 technology. Applications of organoids in basic research involve the modeling of human development and diseases, including genetic, infectious and malignant diseases. Importantly, accumulating evidence suggests that biobanks of patient- derived organoids for many cancers and cystic fibrosis have great value for drug development and personalized medicine. In addition, organoids hold promise for regenerative medicine. In the present review, we discuss the applications of organoids in the basic and translational research. Keywords: Organoids, Stem cells, Disease modeling, Biobanks, Personalized medicine Background phenocopy tumor heterogeneity is highly needed for re- Two-dimensional (2D) cultured cell lines have been the search on the mechanisms of cancer progression and ac- main in vitro research tool for the past decades. Cell quired drug resistance. In 1953, the first patient-derived lines are relatively cheap, easy to handle and can be ap- xenograft (PDX) models were successfully established plied to multiple experimental techniques. However, the (Toolan 1953). In this model, primary tumor tissue is establishment of a cell line is time-consuming and transplanted into immune-deficient mice, while tumor involves extensive genetic and phenotypic adaption to structure and the relative proportion of tumor cells and culture conditions. -

14Q12q13.3 Deletion Diagnosed Using Chromo Somal Microarray
Case Report Neonatal Med 2020 November;27(4):207-213 https://doi.org/10.5385/nm.2020.27.4.207 pISSN 2287-9412 . eISSN 2287-9803 14q12q13.3 Deletion Diagnosed Using Chromo somal Microarray Analysis in an Infant Showing Seizures, Hypoplasia of the Corpus Callosum, and Developmental Delay Jae Hyuk Kwon, MD1, Young Hwa Song, MD1, Jung Min Yoon, MD1, Eun Jung Cheon, MD1, Kyung Ok Ko, MD1, Jae Woo Lim, MD1, and Hyon J. Kim, MD, FACMG2 Departments of 1Pediatrics and 2Medical Genetics, Konyang University College of Medicine, Daejeon, Korea ABSTRACT Received: 18 August 2020 Revised: 25 September 2020 14q12q13.3 Deletion is a rare microdeletion syndrome associated with neurodevelop Accepted: 6 October 2020 mental delay, failure to thrive, seizures, and abnormal brain development. Symptoms Correspondence to: Hyon J. Kim, MD vary depending on the sites of gene deletion, and establishing the diagnosis is often Department of Medical Genetics, difficult, as the condition cannot be detected with routine chromosome analysis. In Konyang University College of Medi this report, we present a patient with intrauterine growth retardation, microcephaly, cine, 158 Gwanjeodongro, Seogu, muscle weakness, seizures, and hypoplasia of the corpus callosum who underwent Daejeon 35365, Korea diagnostic tests, including karyotyping in the neonatal period without leading to a Tel: +82426009230 specific diagnosis. The patient was confirmed with a serious developmental disorder, Fax: +82426009090 and a chromosomal microarray analysis was performed at 8 months of age, revealing Email: [email protected] a 14q12q13.3 deletion. In this case, the condition was diagnosed in early infancy, in contrast to previously reported cases, and the patient had diverse and severe symptoms.