Selection Signatures in Tropical Cattle Are Enriched for Promoter And
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

1 Evidence for Gliadin Antibodies As Causative Agents in Schizophrenia
1 Evidence for gliadin antibodies as causative agents in schizophrenia. C.J.Carter PolygenicPathways, 20 Upper Maze Hill, Saint-Leonard’s on Sea, East Sussex, TN37 0LG [email protected] Tel: 0044 (0)1424 422201 I have no fax Abstract Antibodies to gliadin, a component of gluten, have frequently been reported in schizophrenia patients, and in some cases remission has been noted following the instigation of a gluten free diet. Gliadin is a highly immunogenic protein, and B cell epitopes along its entire immunogenic length are homologous to the products of numerous proteins relevant to schizophrenia (p = 0.012 to 3e-25). These include members of the DISC1 interactome, of glutamate, dopamine and neuregulin signalling networks, and of pathways involved in plasticity, dendritic growth or myelination. Antibodies to gliadin are likely to cross react with these key proteins, as has already been observed with synapsin 1 and calreticulin. Gliadin may thus be a causative agent in schizophrenia, under certain genetic and immunological conditions, producing its effects via antibody mediated knockdown of multiple proteins relevant to the disease process. Because of such homology, an autoimmune response may be sustained by the human antigens that resemble gliadin itself, a scenario supported by many reports of immune activation both in the brain and in lymphocytes in schizophrenia. Gluten free diets and removal of such antibodies may be of therapeutic benefit in certain cases of schizophrenia. 2 Introduction A number of studies from China, Norway, and the USA have reported the presence of gliadin antibodies in schizophrenia 1-5. Gliadin is a component of gluten, intolerance to which is implicated in coeliac disease 6. -
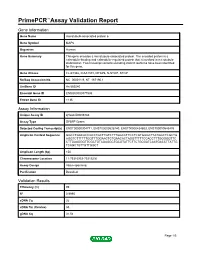
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name microtubule-associated protein 6 Gene Symbol MAP6 Organism Human Gene Summary This gene encodes a microtubule-associated protein. The encoded protein is a calmodulin-binding and calmodulin-regulated protein that is involved in microtubule stabilization. Two transcript variants encoding distinct isoforms have been identified for this gene. Gene Aliases FLJ41346, KIAA1878, MTAP6, N-STOP, STOP RefSeq Accession No. NC_000011.9, NT_167190.1 UniGene ID Hs.585540 Ensembl Gene ID ENSG00000171533 Entrez Gene ID 4135 Assay Information Unique Assay ID qHsaCID0006783 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000304771, ENST00000526740, ENST00000434603, ENST00000545476 Amplicon Context Sequence GGCCTGACACCGCCTGCTTGTCTTTGGCCTTCCTCGTGGGCTTATGGCTCGCTG AGGTCTTTTTTGGTTTGGAACTCTGAACACTAGGTTTTTCCACCTTTGGGGGTTC CTTGAAGGGTTCGCTGTAGAGGCTGCGTATTCTTCTGCGATCAATGACCTTATTG TCAGCTGTTGTTGGCT Amplicon Length (bp) 150 Chromosome Location 11:75316953-75319236 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 99 R2 0.9995 cDNA Cq 26 cDNA Tm (Celsius) 85 gDNA Cq 31.54 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report MAP6, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 3/5 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -

The Role of the C-Terminus Merlin in Its Tumor Suppressor Function Vinay Mandati
The role of the C-terminus Merlin in its tumor suppressor function Vinay Mandati To cite this version: Vinay Mandati. The role of the C-terminus Merlin in its tumor suppressor function. Agricultural sciences. Université Paris Sud - Paris XI, 2013. English. NNT : 2013PA112140. tel-01124131 HAL Id: tel-01124131 https://tel.archives-ouvertes.fr/tel-01124131 Submitted on 19 Mar 2015 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. 1 TABLE OF CONTENTS Abbreviations ……………………………………………………………………………...... 8 Resume …………………………………………………………………………………… 10 Abstract …………………………………………………………………………………….. 11 1. Introduction ………………………………………………………………………………12 1.1 Neurofibromatoses ……………………………………………………………………….14 1.2 NF2 disease ………………………………………………………………………………15 1.3 The NF2 gene …………………………………………………………………………….17 1.4 Mutational spectrum of NF2 gene ………………………………………………………..18 1.5 NF2 in other cancers ……………………………………………………………………...20 2. ERM proteins and Merlin ……………………………………………………………….21 2.1 ERMs ……………………………………………………………………………………..21 2.1.1 Band 4.1 Proteins and ERMs …………………………………………………………...21 2.1.2 ERMs structure ………………………………………………………………………....23 2.1.3 Sub-cellular localization and tissue distribution of ERMs ……………………………..25 2.1.4 ERM proteins and their binding partners ……………………………………………….25 2.1.5 Assimilation of ERMs into signaling pathways ………………………………………...26 2.1.5. A. ERMs and Ras signaling …………………………………………………...26 2.1.5. B. ERMs in membrane transport ………………………………………………29 2.1.6 ERM functions in metastasis …………………………………………………………...30 2.1.7 Regulation of ERM proteins activity …………………………………………………...31 2.1.7. -

And EPB41L3 (Protein 4.1B) Genes
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Genomics 86 (2005) 701 – 707 www.elsevier.com/locate/ygeno Evolutionarily conserved coupling of transcription and alternative splicing in the EPB41 (protein 4.1R) and EPB41L3 (protein 4.1B) genes Jeff S. Tan a, Narla Mohandas b, John G. Conboy a,* a Life Sciences Division and Genomics Department, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA b Red Cell Physiology Group, New York Blood Center, New York, NY 10021, USA Received 20 April 2005; accepted 11 August 2005 Available online 20 October 2005 Abstract Recent studies have shown that transcription and alternative splicing can be mechanistically coupled. In the EPB41 (protein 4.1R) and EPB41L3 (protein 4.1B) genes, we showed previously that promoter/alternative first exon choice is coupled to downstream splicing events in exon 2. Here we demonstrate that this coupling is conserved among several vertebrate classes from fish to mammals. The EPB41 and EPB41L3 genes from fish, bird, amphibian, and mammal genomes exhibit shared features including alternative first exons and differential splice acceptors in exon 2. In all cases, the 5V-most exon (exon 1A) splices exclusively to a weaker internal acceptor site in exon 2, skipping a fragment designated as exon 2V. Conversely, alternative first exons 1B and 1C always splice to the stronger first acceptor site, retaining exon 2V. These correlations are independent of cell type or species of origin. Since exon 2V contains a translation initiation site, splice variants generate protein isoforms with distinct N-termini. -

Microtubule Organization and Microtubule- Associated Proteins (Maps)
Chapter 3 Microtubule Organization and Microtubule- Associated Proteins (MAPs) Elena Tortosa, Lukas C. Kapitein, and Casper C. Hoogenraad Abstract Dendrites have a unique microtubule organization. In vertebrates, den- dritic microtubules are organized in antiparallel bundles, oriented with their plus ends either pointing away or toward the soma. The mixed microtubule arrays control intracellular trafficking and local signaling pathways, and are essential for dendrite development and function. The organization of microtubule arrays largely depends on the combined function of different microtubule regulatory factors or generally named microtubule-associated proteins (MAPs). Classical MAPs, also called structural MAPs, were identified more than 20 years ago based on their ability to bind to and copurify with microtubules. Most classical MAPs bind along the microtubule lattice and regulate microtubule polymerization, bundling, and stabilization. Recent evidences suggest that classical MAPs also guide motor protein transport, interact with the actin cytoskeleton, and act in various neuronal signaling networks. Here, we give an overview of microtubule organization in dendrites and the role of classical MAPs in dendrite development, dendritic spine formation, and synaptic plasticity. Keywords Neuron • Dendrite • Cytoskeleton • Microtubule • Microtubule- associated protein • MAP1 • MAP2 • MAP4 • MAP6 • MAP7 • MAP9 • Tau 3.1 Introduction Microtubules (MTs) are cytoskeletal structures that play essential roles in all eukaryotic cells. MTs are important not only during cell division but also in non-dividing cells, where they are critical structures in numerous cellular processes such as cell motility, migration, differentiation, intracellular transport and organelle positioning. MTs are composed of two proteins, α- and β-tubulin, that form heterodimers and organize themselves in a head-to-tail manner. -

In This Table Protein Name, Uniprot Code, Gene Name P-Value
Supplementary Table S1: In this table protein name, uniprot code, gene name p-value and Fold change (FC) for each comparison are shown, for 299 of the 301 significantly regulated proteins found in both comparisons (p-value<0.01, fold change (FC) >+/-0.37) ALS versus control and FTLD-U versus control. Two uncharacterized proteins have been excluded from this list Protein name Uniprot Gene name p value FC FTLD-U p value FC ALS FTLD-U ALS Cytochrome b-c1 complex P14927 UQCRB 1.534E-03 -1.591E+00 6.005E-04 -1.639E+00 subunit 7 NADH dehydrogenase O95182 NDUFA7 4.127E-04 -9.471E-01 3.467E-05 -1.643E+00 [ubiquinone] 1 alpha subcomplex subunit 7 NADH dehydrogenase O43678 NDUFA2 3.230E-04 -9.145E-01 2.113E-04 -1.450E+00 [ubiquinone] 1 alpha subcomplex subunit 2 NADH dehydrogenase O43920 NDUFS5 1.769E-04 -8.829E-01 3.235E-05 -1.007E+00 [ubiquinone] iron-sulfur protein 5 ARF GTPase-activating A0A0C4DGN6 GIT1 1.306E-03 -8.810E-01 1.115E-03 -7.228E-01 protein GIT1 Methylglutaconyl-CoA Q13825 AUH 6.097E-04 -7.666E-01 5.619E-06 -1.178E+00 hydratase, mitochondrial ADP/ATP translocase 1 P12235 SLC25A4 6.068E-03 -6.095E-01 3.595E-04 -1.011E+00 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 Protein kinase C and casein Q9BY11 PACSIN1 3.837E-03 -5.863E-01 3.680E-06 -1.824E+00 kinase substrate in neurons protein 1 Tubulin polymerization- O94811 TPPP 6.466E-03 -5.755E-01 6.943E-06 -1.169E+00 promoting protein MIC C9JRZ6 CHCHD3 2.912E-02 -6.187E-01 2.195E-03 -9.781E-01 Mitochondrial 2- -

Mutational Mechanisms That Activate Wnt Signaling and Predict Outcomes in Colorectal Cancer Patients William Hankey1, Michael A
Published OnlineFirst December 6, 2017; DOI: 10.1158/0008-5472.CAN-17-1357 Cancer Genome and Epigenome Research Mutational Mechanisms That Activate Wnt Signaling and Predict Outcomes in Colorectal Cancer Patients William Hankey1, Michael A. McIlhatton1, Kenechi Ebede2, Brian Kennedy3, Baris Hancioglu3, Jie Zhang4, Guy N. Brock3, Kun Huang4, and Joanna Groden1 Abstract APC biallelic loss-of-function mutations are the most prevalent also exhibiting unique changes in pathways related to prolifera- genetic changes in colorectal tumors, but it is unknown whether tion, cytoskeletal organization, and apoptosis. Apc-mutant ade- these mutations phenocopy gain-of-function mutations in the nomas were characterized by increased expression of the glial CTNNB1 gene encoding b-catenin that also activate canonical nexin Serpine2, the human ortholog, which was increased in WNT signaling. Here we demonstrate that these two mutational advanced human colorectal tumors. Our results support the mechanisms are not equivalent. Furthermore, we show how hypothesis that APC-mutant colorectal tumors are transcription- differences in gene expression produced by these different ally distinct from APC-wild-type colorectal tumors with canonical mechanisms can stratify outcomes in more advanced human WNT signaling activated by other mechanisms, with possible colorectal cancers. Gene expression profiling in Apc-mutant and implications for stratification and prognosis. Ctnnb1-mutant mouse colon adenomas identified candidate Significance: These findings suggest that colon adenomas genes for subsequent evaluation of human TCGA (The Cancer driven by APC mutations are distinct from those driven by WNT Genome Atlas) data for colorectal cancer outcomes. Transcrip- gain-of-function mutations, with implications for identifying tional patterns exhibited evidence of activated canonical Wnt at-risk patients with advanced disease based on gene expression signaling in both types of adenomas, with Apc-mutant adenomas patterns. -
Inter-Chromosomal Variation in the Pattern of Human Population Genetic Structure Tesfaye M
PRIMARY RESEARCH Inter-chromosomal variation in the pattern of human population genetic structure Tesfaye M. Baye* Cincinnati Children’s Hospital Medical Center, Division of Asthma Research, Department of Pediatrics, University of Cincinnati, 3333 Burnet Avenue, Cincinnati, OH 45229, USA *Correspondence to: Tel: þ1 513 803 2766; Fax: þ1 513 636 1657; E-mail: [email protected] Date received (in revised form): 1st March 2011 Abstract Emerging technologies now make it possible to genotype hundreds of thousands of genetic variations in individuals, across the genome. The study of loci at finer scales will facilitate the understanding of genetic variation at genomic and geographic levels. We examined global and chromosomal variations across HapMap populations using 3.7 million single nucleotide polymorphisms to search for the most stratified genomic regions of human populations and linked these regions to ontological annotation and functional network analysis. To achieve this, we used five complementary statistical and genetic network procedures: principal component (PC), cluster, discriminant, fix- ation index (FST) and network/pathway analyses. At the global level, the first two PC scores were sufficient to account for major population structure; however, chromosomal level analysis detected subtle forms of population structure within continental populations, and as many as 31 PCs were required to classify individuals into homo- geneous groups. Using recommended population ancestry differentiation measures, a total of 126 regions of the genome were catalogued. Gene ontology and networks analyses revealed that these regions included the genes encoding oculocutaneous albinism II (OCA2), hect domain and RLD 2 (HERC2), ectodysplasin A receptor (EDAR) and solute carrier family 45, member 2 (SLC45A2). -

Comprehensive Analysis Reveals Novel Gene Signature in Head and Neck Squamous Cell Carcinoma: Predicting Is Associated with Poor Prognosis in Patients
5892 Original Article Comprehensive analysis reveals novel gene signature in head and neck squamous cell carcinoma: predicting is associated with poor prognosis in patients Yixin Sun1,2#, Quan Zhang1,2#, Lanlin Yao2#, Shuai Wang3, Zhiming Zhang1,2 1Department of Breast Surgery, The First Affiliated Hospital of Xiamen University, School of Medicine, Xiamen University, Xiamen, China; 2School of Medicine, Xiamen University, Xiamen, China; 3State Key Laboratory of Cellular Stress Biology, School of Life Sciences, Xiamen University, Xiamen, China Contributions: (I) Conception and design: Y Sun, Q Zhang; (II) Administrative support: Z Zhang; (III) Provision of study materials or patients: Y Sun, Q Zhang; (IV) Collection and assembly of data: Y Sun, L Yao; (V) Data analysis and interpretation: Y Sun, S Wang; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors. #These authors contributed equally to this work. Correspondence to: Zhiming Zhang. Department of Surgery, The First Affiliated Hospital of Xiamen University, Xiamen, China. Email: [email protected]. Background: Head and neck squamous cell carcinoma (HNSC) remains an important public health problem, with classic risk factors being smoking and excessive alcohol consumption and usually has a poor prognosis. Therefore, it is important to explore the underlying mechanisms of tumorigenesis and screen the genes and pathways identified from such studies and their role in pathogenesis. The purpose of this study was to identify genes or signal pathways associated with the development of HNSC. Methods: In this study, we downloaded gene expression profiles of GSE53819 from the Gene Expression Omnibus (GEO) database, including 18 HNSC tissues and 18 normal tissues. -

Protein 4.1, a Component of the Erythrocyte Membrane Skeleton and Its Related Homologue Proteins Forming the Protein 4.1/FERM Superfamily
FOLIA HISTOCHEMICA Review article ET CYTOBIOLOGICA Vol. 44, No. 4, 2006 pp. 231-248 Protein 4.1, a component of the erythrocyte membrane skeleton and its related homologue proteins forming the protein 4.1/FERM superfamily Witold Diakowski, Micha³ Grzybek and Aleksander F. Sikorski Faculty of Biotechnology, University of Wroc³aw, Wroc³aw, Poland Abstract: The review is focused on the domain structure and function of protein 4.1, one of the proteins belonging to the mem- brane skeleton. The protein 4.1 of the red blood cells (4.1R) is a multifunctional protein that localizes to the membrane skele- ton and stabilizes erythrocyte shape and membrane mechanical properties, such as deformability and stability, via lateral inter- actions with spectrin, actin, glycophorin C and protein p55. Protein 4.1 binding is modulated through the action of kinases and/or calmodulin-Ca2+. Non-erythroid cells express the 4.1R homologues: 4.1G (general type), 4.1B (brain type), and 4.1N (neuron type), and the whole group belongs to the protein 4.1 superfamily, which is characterized by the presence of a highly conserved FERM domain at the N-terminus of the molecule. Proteins 4.1R, 4.1 G, 4.1 N and 4.1 B are encoded by different genes. Most of the 4.1 superfamily proteins also contain an actin-binding domain. To date, more than 40 members have been identified. They can be divided into five groups: protein 4.1 molecules, ERM proteins, talin-related molecules, protein tyrosine phosphatase (PTPH) proteins and NBL4 proteins. We have focused our attention on the main, well known representatives of 4.1 superfamily and tried to choose the proteins which are close to 4.1R or which have distinct functions. -

Novel Protein Pathways in Development and Progression of Pulmonary Sarcoidosis Maneesh Bhargava1*, K
www.nature.com/scientificreports OPEN Novel protein pathways in development and progression of pulmonary sarcoidosis Maneesh Bhargava1*, K. J. Viken1, B. Barkes2, T. J. Grifn3, M. Gillespie2, P. D. Jagtap3, R. Sajulga3, E. J. Peterson4, H. E. Dincer1, L. Li2, C. I. Restrepo2, B. P. O’Connor5, T. E. Fingerlin5, D. M. Perlman1 & L. A. Maier2 Pulmonary involvement occurs in up to 95% of sarcoidosis cases. In this pilot study, we examine lung compartment-specifc protein expression to identify pathways linked to development and progression of pulmonary sarcoidosis. We characterized bronchoalveolar lavage (BAL) cells and fuid (BALF) proteins in recently diagnosed sarcoidosis cases. We identifed 4,306 proteins in BAL cells, of which 272 proteins were diferentially expressed in sarcoidosis compared to controls. These proteins map to novel pathways such as integrin-linked kinase and IL-8 signaling and previously implicated pathways in sarcoidosis, including phagosome maturation, clathrin-mediated endocytic signaling and redox balance. In the BALF, the diferentially expressed proteins map to several pathways identifed in the BAL cells. The diferentially expressed BALF proteins also map to aryl hydrocarbon signaling, communication between innate and adaptive immune response, integrin, PTEN and phospholipase C signaling, serotonin and tryptophan metabolism, autophagy, and B cell receptor signaling. Additional pathways that were diferent between progressive and non-progressive sarcoidosis in the BALF included CD28 signaling and PFKFB4 signaling. Our studies demonstrate the power of contemporary proteomics to reveal novel mechanisms operational in sarcoidosis. Application of our workfows in well-phenotyped large cohorts maybe benefcial to identify biomarkers for diagnosis and prognosis and therapeutically tenable molecular mechanisms. -

Supplementary Figures and Table
SUPPLEMENTARY DATA Supplementary Figure 1. ©2014 American Diabetes Association. Published online at http://diabetes.diabetesjournals.org/lookup/suppl/doi:10.2337/db141 -0066/-/DC1 SUPPLEMENTARY DATA Supplementary Figure 2. ©2014 American Diabetes Association. Published online at http://diabetes.diabetesjournals.org/lookup/suppl/doi:10.2337/db142 -0066/-/DC1 SUPPLEMENTARY DATA -/- Supplementary Table 1. Fold increase of Ser/Thr/Tyr phosphorylation in livers of MKP-3 male mice versus wild type male mice fed on a high fat diet (n=5 for each group). Symbol Name Phosphorylation KO/WT ratio Q Value sites Apoptosis ACIN1 Acin1 protein S64 11.4 0.02 T66 8.3 0.02 API5 Apoptosis inhibitor 5 S461 2.2 0.03 S462 1.8 0.03 AIFM3 Apoptosis-inducing factor 3 S30 7.4 0.03 TP53BP2 Apoptosis-stimulating of p53 protein 2 S479 3.7 0.02 ACIN1 Apoptotic chromatin condensation inducer S64S70 5.7 0.02 1 S208 7.1 0.02 S210 7.0 0.02 S479S482S491 105.7 0.03 S729 2.8 0.02 PEA15 Astrocytic phosphoprotein PEA-15 S116 10.8 0.02 BAG3 BAG family molecular chaperone regulator S179 3.3 0.02 3 S353S357 2.3 0.03 S360 2.3 0.03 S390 8.4 0.02 BNIP2 BCL2/adenovirus E1B 19 kDa-interacting S114 3.9 0.02 protein 2 alpha BNIP3 BCL2/adenovirus E1B 19 kDa protein- S60 19.8 0.03 interacting protein 3 S85T86 14.5 0.02 S88 6.1 0.02 BCL2L13 Bcl-2-like protein 13 S387 4.0 0.02 T389 3.1 0.02 CAAP1 Caspase activity and apoptosis inhibitor S183 2.3 0.03 CARD6 Card6 caspase recruitment domain family, S809 3.6 0.03 member 6 CASP8 Caspase-8 S188 2.2 0.02 DAP Death-associated protein S51 5.4 0.02 DAPK2 Death-associated protein kinase 2 S299 3.8 0.02 S349 3.5 0.02 FAF1 FAS-associated factor 1 S269 17.1 0.04 GAS2 Growth arrest-specific protein 2 T282 5.3 0.02 S283 7.4 0.02 S287 5.3 0.02 S289 7.4 0.02 GCH1 GTP cyclohydrolase 1 S24 3.9 0.02 HTT Huntingtin S398S409S411 9.7 0.02 KRT18 Keratin, type I cytoskeletal 18 T9 2.7 0.02 S31S32S35 2.8 0.02 S43S45 3.1 0.02 PDCD5 MCG128907 S119 10.7 0.02 Y126 4.0 0.02 BNIP3I MCG2480, isoform CRA_b S61S62 12.9 0.03 S63S64 8.1 0.02 ©2014 American Diabetes Association.