Catalytic Properties and Inhibition of Proline-Specific Dipeptidyl Peptidases II, IV and VII
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genetic Polymorphism of the Angiotensin-Converting Enzyme (ACE) in Asthmatic Patients
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector RESPIRATORY MEDICINE (1998) 92, 1305-1310 Original Articles Genetic polymorphism of the angiotensin-converting enzyme (ACE) in asthmatic patients H. TOMITA, S. SATO, R. MATSUDA, N. OGISU, T. MORI, T. NIIMI AND S. SHIMIZU 2nd Department of Intemal Medicine, Nagoya City University Medical School, Nagoya, Japan Angiotensin-converting enzyme (ACE) inactivates bradykinin, substance P and neurokinin A, which are believed to play important roles in the pathogenesis of asthma, especially in neurogenic inflammation. It has recently been shown that an insertion (1)ldeletion (D) polymorphism in the ACE gene accounts for variation in serum ACE levels. There are thus three genotypes (insertion homozygote, II; deletion homozygote, DD; heterozygotes, DI). The serum ACE level with the DD type is reported to be about double that of the II type and intermediate in the DI case. In the present study, we examined whether asthma is linked with this ACE gene polymorphism. Seventy-one patients with asthma (27 males and 44 females) and 142 sex- and age-matched healthy controls were determined for their genotype by the polymerase chain reaction (PCR) method. Twenty-five asthmatics demonstrated the II type (352%) 37 the DI type (52.1%), and nine the DD type (12.7%). There were no significant differences in the distributions of genotypes and serum ACE levels between healthy controls and patients. No significant differences were evident in serum IgE levels, age at onset, proportion of atopic type patients and severity of asthma among the three genotypes. -

B Number Gene Name Mrna Intensity Mrna
sample) total list predicted B number Gene name assignment mRNA present mRNA intensity Gene description Protein detected - Membrane protein membrane sample detected (total list) Proteins detected - Functional category # of tryptic peptides # of tryptic peptides # of tryptic peptides detected (membrane b0002 thrA 13624 P 39 P 18 P(m) 2 aspartokinase I, homoserine dehydrogenase I Metabolism of small molecules b0003 thrB 6781 P 9 P 3 0 homoserine kinase Metabolism of small molecules b0004 thrC 15039 P 18 P 10 0 threonine synthase Metabolism of small molecules b0008 talB 20561 P 20 P 13 0 transaldolase B Metabolism of small molecules chaperone Hsp70; DNA biosynthesis; autoregulated heat shock b0014 dnaK 13283 P 32 P 23 0 proteins Cell processes b0015 dnaJ 4492 P 13 P 4 P(m) 1 chaperone with DnaK; heat shock protein Cell processes b0029 lytB 1331 P 16 P 2 0 control of stringent response; involved in penicillin tolerance Global functions b0032 carA 9312 P 14 P 8 0 carbamoyl-phosphate synthetase, glutamine (small) subunit Metabolism of small molecules b0033 carB 7656 P 48 P 17 0 carbamoyl-phosphate synthase large subunit Metabolism of small molecules b0048 folA 1588 P 7 P 1 0 dihydrofolate reductase type I; trimethoprim resistance Metabolism of small molecules peptidyl-prolyl cis-trans isomerase (PPIase), involved in maturation of b0053 surA 3825 P 19 P 4 P(m) 1 GenProt outer membrane proteins (1st module) Cell processes b0054 imp 2737 P 42 P 5 P(m) 5 GenProt organic solvent tolerance Cell processes b0071 leuD 4770 P 10 P 9 0 isopropylmalate -
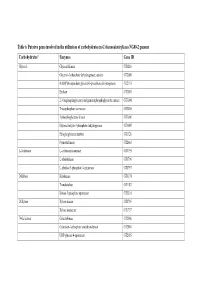
Table 6. Putative Genes Involved in the Utilization of Carbohydrates in G
Table 6. Putative genes involved in the utilization of carbohydrates in G. thermodenitrificans NG80-2 genome Carbohydrates* Enzymes Gene ID Glycerol Glycerol Kinase GT1216 Glycerol-3-phosphate dehydrogenase, aerobic GT2089 NAD(P)H-dependent glycerol-3-phosphate dehydrogenase GT2153 Enolase GT3003 2,3-bisphosphoglycerate-independentphosphoglycerate mutase GT3004 Triosephosphate isomerase GT3005 3-phosphoglycerate kinase GT3006 Glyceraldehyde-3-phosphate dehydrogenase GT3007 Phosphoglycerate mutase GT1326 Pyruvate kinase GT2663 L-Arabinose L-arabinose isomerase GT1795 L-ribulokinase GT1796 L-ribulose 5-phosphate 4-epimerase GT1797 D-Ribose Ribokinase GT3174 Transketolase GT1187 Ribose 5-phosphate epimerase GT3316 D-Xylose Xylose kinase GT1756 Xylose isomerase GT1757 D-Galactose Galactokinase GT2086 Galactose-1-phosphate uridyltransferase GT2084 UDP-glucose 4-epimerase GT2085 Carbohydrates* Enzymes Gene ID D-Fructose 1-phosphofructokinase GT1727 Fructose-1,6-bisphosphate aldolase GT1805 Fructose-1,6-bisphosphate aldolase type II GT3331 Triosephosphate isomerase GT3005 D-Mannose Mannnose-6 phospate isomelase GT3398 6-phospho-1-fructokinase GT2664 D-Mannitol Mannitol-1-phosphate dehydrogenase GT1844 N-Acetylglucosamine N-acetylglucosamine-6-phosphate deacetylase GT2205 N-acetylglucosamine-6-phosphate isomerase GT2204 D-Maltose Alpha-1,4-glucosidase GT0528, GT1643 Sucrose Sucrose phosphorylase GT3215 D-Trehalose Alpha-glucosidase GT1643 Glucose kinase GT2381 Inositol Myo-inositol catabolism protein iolC;5-dehydro-2- GT1807 deoxygluconokinase -

Partial Characterisation of Dipeptidyl Peptidase 4 (DP 4
Partial Characterisation of Dipeptidyl Peptidase 4 (DP 4) for the Treatment of Type 2 Diabetes and its Identification as a novel Pharmaceutical Target for Stress-related Indications Beiträge zur Charakterisierung der Dipeptidylpeptidase 4 (DP 4) für die Behandlung von Typ-2-Diabetes und deren Identifizierung als ein neuartiges pharmazeutisches Ziel für stressvermittelte Indikationen Der Naturwissenschaftlichen Fakultät der Friedrich-Alexander-Universität Erlangen-Nürnberg zur Erlangung des Doktorgrades Dr. rer. nat. vorgelegt von M. Sc. (Biochemie) Leona Wagner aus Rheinfelden (Baden) Als Dissertation genehmigt von der Naturwissenschaftlichen Fakultät der Friedrich-Alexander-Universität Erlangen-Nürnberg Tag der mündlichen Prüfung: 23.10.2012 Vorsitzender der Promotionskommission: Prof. Dr. Rainer Fink Erstberichterstatter: Prof. Dr. med. Stephan von Hörsten Zweitberichterstatter: Prof. Dr. Johann Helmut Brandstätter Publications, Presentations and Posters Part of the present study have been published or submitted in the following journals or presented on the following conferences: Publications: 1. Hinke, S.A., McIntosh, C.H., Hoffmann, T., Kuhn-Wache, K., Wagner, L., Bar, J., Manhart, S., Wermann, M., Pederson, R.A., and Demuth, H.U. (2002) On combination therapy of diabetes with metformin and dipeptidyl peptidase IV inhibitors. Diabetes Care 25, 1490- 1491. 2. Bär, J., Weber, A., Hoffmann, T., Stork, J., Wermann, M., Wagner, L., Aust, S., Gerhartz, B., and Demuth, H.U. (2003) Characterisation of human dipeptidyl peptidase IV expressed in Pichia pastoris. A structural and mechanistic comparison between the recombinant human and the purified porcine enzyme. Biol. Chem. 384, 1553-1563. 3. Engel, M., Hoffmann, T., Wagner, L., Wermann, M., Heiser, U., Kiefersauer, R., Huber, R., Bode, W., Demuth, H.U., and Brandstetter, H. -

Dipeptidase Aminodipeptidase, Quiescent Cell Proline of a Novel
Vesicular Localization and Characterization of a Novel Post-Proline-Cleaving Aminodipeptidase, Quiescent Cell Proline Dipeptidase This information is current as of September 26, 2021. Murali Chiravuri, Fernando Agarraberes, Suzanne L. Mathieu, Henry Lee and Brigitte T. Huber J Immunol 2000; 165:5695-5702; ; doi: 10.4049/jimmunol.165.10.5695 http://www.jimmunol.org/content/165/10/5695 Downloaded from References This article cites 28 articles, 18 of which you can access for free at: http://www.jimmunol.org/content/165/10/5695.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on September 26, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2000 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. Vesicular Localization and Characterization of a Novel Post-Proline-Cleaving Aminodipeptidase, Quiescent Cell Proline Dipeptidase1 Murali Chiravuri,* Fernando Agarraberes,† Suzanne L. Mathieu,* Henry Lee,* and Brigitte T. Huber2* A large number of chemokines, cytokines, and signal peptides share a highly conserved X-Pro motif on the N-terminus. -

Aminopeptidase P1 Recombinant Protein Cat
Aminopeptidase P1 Recombinant Protein Cat. No.: 91-173 Aminopeptidase P1 Recombinant Protein Specifications SPECIES: Human SOURCE SPECIES: E. coli SEQUENCE: Pro2-His623 FUSION TAG: C-6 His tag TESTED APPLICATIONS: APPLICATIONS: This recombinant protein can be used for biological assays. For research use only. PREDICTED MOLECULAR 70.6 kD WEIGHT: Properties Greater than 95% as determined by reducing SDS-PAGE. PURITY: Endotoxin level less than 0.1 ng/ug (1 IEU/ug) as determined by LAL test. PHYSICAL STATE: Liquid Supplied as a 0.2 um filtered solution of 20mM TrisHCl, 10% Glycerol, pH 8.0. It is not BUFFER: recommended to reconstitute to a concentration less than 100 ug/ml. September 23, 2021 1 https://www.prosci-inc.com/aminopeptidase-p1-recombinant-protein-91-173.html Store at -20˚C, stable for 6 months after receipt. STORAGE CONDITIONS: Please aliquot the reconstituted solution to minimize freeze-thaw cycles. Additional Info OFFICIAL SYMBOL: XPNPEP1 Xaa-Pro Aminopeptidase 1, Aminoacylproline Aminopeptidase, Cytosolic Aminopeptidase ALTERNATE NAMES: P, Soluble Aminopeptidase P, sAmp, X-Pro Aminopeptidase 1, X-Prolyl Aminopeptidase 1 Soluble, XPNPEP1, XPNPEPL, XPNPEPL1 ACCESSION NO.: Q9NQW7 GENE ID: 7511 Background and References X-Prolyl Aminopeptidase (XPNPEP1) is a proline-specific metalloaminopeptidase that specifically catalyzes the removal of any unsubstituted N-terminal amino acid that is adjacent to a penultimate proline residue. Because of its specificity toward proline, it has been suggested that X-Prolyl Aminopeptidase is important in the maturation and degradation of peptide hormones, neuropeptides, and tachykinins, as well as in the digestion of otherwise resistant dietary protein fragments, thereby complementing the pancreatic peptidases. -

Brush Border and Cytosol Peptidase Activities of Human Small Intestine in Normal Subjects and Celiac Patients
Pediatr. Res. 14: 8 12-8 18 ( 1980) brush border membrane digestive peptidase: celiac disease small intestine cytosol membrane Brush Border and Cytosol Peptidase Activities of Human Small Intestine in Normal Subjects and Celiac Patients CiLNtKOSO ANIIKIA. SALVIZ'TOKE <'UC'<'lIIAKA. BIZSILIO DL: VIZIA. (;IOKGIO IIL: KITIS. <;ABRIEL.E MAZZA<'<'A. AN11 SALVATORE 1Z~R1~('1110'~" I)c.prrrrn~enrof Pediclrrrc.~ond Depurrmenr of (;ii.crrt~c~nrerologr,I1 I.irc~ttlt~ij .&ledicrne. L'tt~r.i~rvrtrof Ni~pler.Sc~plt,\. Irull.. und ('on.\r,qlro .Vu:ronule dcdle Rrc.erchr. Pro~rirrn(4- Prevc,ttrrve Medic~itrc(Prt~~cc./ I'ermi~rirl ,Wedrc.itlc~. Rontc. Irul, Summary Studies in animals have demonstrated two major subcellular localization of digestive peptidases in the enterocyte. cytosol. and Peptidase activities have been investigated in the brush border brush border, M~~~ of the hydrolyzing dipep- of human proximal jejunum by using dipeptides and tripeptides [ides tripeprides is localized in ,he cytosol (1, 16, 17. 21, 26. and P-naphthylamides of glycyl-I.-proline and amino acids as sub- 32, 35, 42, 46, 49, 53, 54, 60) few except,ons (25, 26. 42. 46. \trates. I'he activities hydrolyzing glycyl-I.-leucine. I.-phenylalanyl- 49, 60): three soluble enzymes dipeptidase and tripeptidase I.-alaninc, and I.-le~cyl~lycylglycinein the brush horder were found have been and (12, 15, 16, 20, 48. to be only 1.5. 15. and 16% of the total peptidase activities present 5 1. 62). Almost all the hydrolyzing the P- in the intestinal mucosa, but the specific activities for the hydrol- naphthylamides ofamino (6. -

Contribution of Endo- and Exopeptidases to Formation of Non-Protein N During Ensiling of Alfalfa
Contribution of endo- and exopeptidases to formation of non-protein N during ensiling of alfalfa Dr. Xusheng Guo,1 W. Cheng,1 L. Tao,2 Yu Zhu,2 F.Y. Yang 2 & H. Zhou2 1.School of Life Science, Lanzhou University 1. International Centre for Tibetan Plateau Ecosystem Management (ICTPEM), Lanzhou University 2. Institute of Grassland Science, China Agricultural University Hämeenlinna, Finland 2-4 July 2012 Introduction Alfalfa (Medicago Sativa L.) is well known for its high nutritive value However, after ensiling: N use efficiency Extensive ppyroteolysis Reduce True protein NPN(Peptide, FAA, NH3-N etc.) Silage DM intake (44-87% of Total N; Muck, 1987) Silage Fermentation Proteolysis in ensiled forage mainly results from plant proteases (Ohshima and McDonald, 1978; McKersie, 1981; Heron et al., 1988). Proteases (peptidases) are divided into 2 classes (NC-IUBMB, 1992): Exopeptidase Endopeptidase Objectives Proteases (peptidases, E.C.3.4) Endopeptidases Exopeptidases ?! Ser ine pep tidase (E .C .3 .4 .21) Aminopeptidase (EC 3.4.11) Carboxypeptidase (EC 3.4.16) Cysteine peptidase (E.C.3.4.22) Dipeptidase (EC 3.4.13) Aspartic Peptidase (E.C.3.4.23) Dipeptidyl-peptidase (EC 3.4.14) Metallopeptidase (E.C.3.4.24) Tripeptidyl-peptidase (EC 3.4.14) Peptidyl-dipeptidase (EC 3.4.15) Aims of our research were: 1. To clarify the classes of exo- and endopeptidases that are involved in proteolysis within ensiled alfalfa. 2. To determine the contribution of these peptidases to the formation of different NPN compounds (peptide-N, FAA-N, and NH3-N) during -

Cathepsin B Carboxydipeptidase Specificity Analysis Using Internally
Biochem. J. (2002) 368, 365–369 (Printed in Great Britain) 365 Cathepsin B carboxydipeptidase specificity analysis using internally quenched fluorescent peptides Maria Helena S. CEZARI, Luciano PUZER, Maria Aparecida JULIANO, Adriana K. CARMONA and Luiz JULIANO1 Department of Biophysics, Escola Paulista de Medicina, Universidade Federal de Sa4 o Paulo, Rua Tre# s de Maio, 100, Sa4 o Paulo 04044-020, Brazil We have examined in detail the specificity of the subsites S",S#, analysis of its specificity. The subsite S" accepted preferentially h h S" and S# for the carboxydipeptidase activity of cathepsin B by basic amino acids for hydrolysis; however, substrates with synthesizing and assaying four series of internally quenched phenylalanine and aliphatic side-chain-containing amino acids at #%& fluorescent peptides based on the sequence Dnp-GFRFW-OH, P" had lower Km values. Despite the presence of Glu at S#, where Dnp (2,4-dinitrophenyl) is the quenching group of the this subsite presented clear preference for aromatic amino acid fluorescence of the tryptophan residue. Each position, except the residues, and the substrate with a lysine residue at P# was h glycine, was substituted with 15 different naturally occurring hydrolysed better than that containing an arginine residue. S" is h amino acids. Based on the results we obtained, we also synthesized essentially a hydrophobic subsite, and S# has particular efficient and sensitive substrates that contained o-aminobenzoic preference for phenylalanine or tryptophan residues. acid and 3-Dnp-(2,3-diaminopropionic acid), or ε-amino-Dnp- Lys, as the fluorescence donor–receptor pair. The higher kinetic parameter values for the carboxydipeptidase compared with the Key words: fluorescent peptide, fluorogenic substrate, proteinase, endopeptidase activity of cathepsin B allowed an accurate thiol protease. -

Human Aminopeptidases: a Review of the Literature
Sanderink et al: Aminopeptidases, a review 795 J. Clin. Chem. Clin. Biochem. Vol. 26, 1988, pp. 795-807 © 1988 Waiter de Gruyter & Co. Berlin · New York Human Aminopeptidases: A Review of the Literature By G.-J. Sanderink, Y. Artur and G. Siesf Laboratoire du Centre de Medecine Preventive, UA CNRS n° 597, Vandoeuvre-les-Nancy, France (Received April 5/August 1, 1988) Summary: The aminopeptidases constitute a group of enzymes with closely related activities. In clinical chemistry the analysis of the aminopeptidases and of their multiple forms in serum has for a long time been hindered by considerable confusion concerning their identification, and by a lack of characterization. This is in part due to the often large, and sometimes overlapping substrate specificities of the aminopeptidases. This paper reviews the biochemical properties of the different aminopeptidases, the specificities of the assays used for their analysis in serum, some aspects of their multiple forms — which are especially known to occur for alanine äminopeptidase (EC 3.4.11.2) — and the importance of the determination of aminopeptidases and their multiple forms in clinical chemistry. Introduction ificities, pH optima, activators, etc. A list of some Aminopeptidases are enzymes which hydrolyse pep- human aminopeptidases acting on polypeptides is tide bonds near the N-terminal end of polypeptides. given in table 1. They can be subdivided into aminopeptidases which — substrate specificities; the different aminopepti- hydrolyse the first peptide bond (aminoacyl-peptide dases have a closely related enzymatic activity with hydrolases and iminoacyl-peptide hydrolases) and sometimes broad specificities. These specificities those which remove dipeptides from polypeptide overlap, so that many natural and synthetic sub- chains (dipeptidyl-peptide hydrolases). -

Production of the Antihypertensive Peptide Tyr-Pro from Milk Using the White-Rot Fungus Peniophora Sp
Article Production of the Antihypertensive Peptide Tyr-Pro from Milk Using the White-Rot Fungus Peniophora sp. in Submerged Fermentation and a Jar Fermentor Kenji Okamoto * , Ryosuke Ito, June Hayashi and Mizuki Tagawa Department of Chemistry and Biotechnology, Graduate School of Engineering, Tottori University, Tottori 680-8552, Japan; [email protected] (R.I.); [email protected] (J.H.); [email protected] (M.T.) * Correspondence: [email protected] Abstract: In order to evaluate the blood pressure-lowering peptide Tyr-Pro (YP) derived from casein, we wanted to develop an efficient fermentation method. Therefore, we chose to use a jar fermentor for this purpose. Strains with an excellent antihypertensive peptide-releasing ability from casein were selected from basidiomycete fungi that grow well in milk under shaking conditions accompanied by physical stimulation. Among them, the white-rot fungus Peniophora sp., which is suited for growth only in cow’s milk or low-fat milk under vigorous shaking conditions, was found to release peptides and amino acids from milk. When comparing the growth in cow’s milk and low-fat milk, there was no particular difference in the growth of mycelia between the two, but this fungus tended to preferentially consume lactose under low-fat conditions. The fermented milk exhibited good production of the target peptide YP. The expression of many genes encoding proteolytic enzymes, such as aminopeptidases and carboxypeptidases, was observed during the milk Citation: Okamoto, K.; Ito, R.; fermentation. Furthermore, this fungus showed good growth in a jar fermentor culture using only Hayashi, J.; Tagawa, M. Production of cow’s milk or low-fat milk, which enabled the efficient production of YP and ACE-inhibitory activity. -

Structures and Mechanism of Dipeptidyl Peptidases 8 and 9
Structures and mechanism of dipeptidyl peptidases PNAS PLUS 8 and 9, important players in cellular homeostasis and cancer Breyan Rossa,b,1, Stephan Krappb, Martin Augustinb, Reiner Kierfersauerb, Marcelino Arciniegac, Ruth Geiss-Friedlanderd, and Robert Hubera,e,f,1 aMax Planck Institut für Biochemie, D-82152 Martinsried, Germany; bProteros Biostructures GmbH, D-82152 Martinsried, Germany; cDepartment of Biochemistry and Structural Biology, Institute of Cellular Physiology, Universidad Nacional Autónoma de México, 04510 Mexico City, Mexico; dAbteilung für Molekularbiologie, Universitätsmedizin Göttingen, D-37073 Göttingen, Germany; eZentrum für Medizinische Biotechnologie, Universität Duisburg-Essen, D-45117 Essen, Germany; and fFakultät für Chemie, Technische Universität München, D-85747 Garching, Germany Contributed by Robert Huber, December 12, 2017 (sent for review October 16, 2017; reviewed by Ingrid De Meester and Guy S. Salvesen) Dipeptidyl peptidases 8 and 9 are intracellular N-terminal dipeptidyl be processed by DPP9, thereby regulating B cell signaling (18). peptidases (preferentially postproline) associated with pathophysi- DPP9 activity is also connected to pathophysiological conditions, ological roles in immune response and cancer biology. While the as promoting tumoregenicity and metastasis in nonsmall cell lung DPP family member DPP4 is extensively characterized in molecular cancer (19). Recently, DPP9 fusion genes were identified in high- terms as a validated therapeutic target of type II diabetes, experimental grade serous