Human Aminopeptidases: a Review of the Literature
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Enzymatic Encoding Methods for Efficient Synthesis Of
(19) TZZ__T (11) EP 1 957 644 B1 (12) EUROPEAN PATENT SPECIFICATION (45) Date of publication and mention (51) Int Cl.: of the grant of the patent: C12N 15/10 (2006.01) C12Q 1/68 (2006.01) 01.12.2010 Bulletin 2010/48 C40B 40/06 (2006.01) C40B 50/06 (2006.01) (21) Application number: 06818144.5 (86) International application number: PCT/DK2006/000685 (22) Date of filing: 01.12.2006 (87) International publication number: WO 2007/062664 (07.06.2007 Gazette 2007/23) (54) ENZYMATIC ENCODING METHODS FOR EFFICIENT SYNTHESIS OF LARGE LIBRARIES ENZYMVERMITTELNDE KODIERUNGSMETHODEN FÜR EINE EFFIZIENTE SYNTHESE VON GROSSEN BIBLIOTHEKEN PROCEDES DE CODAGE ENZYMATIQUE DESTINES A LA SYNTHESE EFFICACE DE BIBLIOTHEQUES IMPORTANTES (84) Designated Contracting States: • GOLDBECH, Anne AT BE BG CH CY CZ DE DK EE ES FI FR GB GR DK-2200 Copenhagen N (DK) HU IE IS IT LI LT LU LV MC NL PL PT RO SE SI • DE LEON, Daen SK TR DK-2300 Copenhagen S (DK) Designated Extension States: • KALDOR, Ditte Kievsmose AL BA HR MK RS DK-2880 Bagsvaerd (DK) • SLØK, Frank Abilgaard (30) Priority: 01.12.2005 DK 200501704 DK-3450 Allerød (DK) 02.12.2005 US 741490 P • HUSEMOEN, Birgitte Nystrup DK-2500 Valby (DK) (43) Date of publication of application: • DOLBERG, Johannes 20.08.2008 Bulletin 2008/34 DK-1674 Copenhagen V (DK) • JENSEN, Kim Birkebæk (73) Proprietor: Nuevolution A/S DK-2610 Rødovre (DK) 2100 Copenhagen 0 (DK) • PETERSEN, Lene DK-2100 Copenhagen Ø (DK) (72) Inventors: • NØRREGAARD-MADSEN, Mads • FRANCH, Thomas DK-3460 Birkerød (DK) DK-3070 Snekkersten (DK) • GODSKESEN, -

PROTEOMIC ANALYSIS of HUMAN URINARY EXOSOMES. Patricia
ABSTRACT Title of Document: PROTEOMIC ANALYSIS OF HUMAN URINARY EXOSOMES. Patricia Amalia Gonzales Mancilla, Ph.D., 2009 Directed By: Associate Professor Nam Sun Wang, Department of Chemical and Biomolecular Engineering Exosomes originate as the internal vesicles of multivesicular bodies (MVBs) in cells. These small vesicles (40-100 nm) have been shown to be secreted by most cell types throughout the body. In the kidney, urinary exosomes are released to the urine by fusion of the outer membrane of the MVBs with the apical plasma membrane of renal tubular epithelia. Exosomes contain apical membrane and cytosolic proteins and can be isolated using differential centrifugation. The analysis of urinary exosomes provides a non- invasive means of acquiring information about the physiological or pathophysiological state of renal cells. The overall objective of this research was to develop methods and knowledge infrastructure for urinary proteomics. We proposed to conduct a proteomic analysis of human urinary exosomes. The first objective was to profile the proteome of human urinary exosomes using liquid chromatography-tandem spectrometry (LC- MS/MS) and specialized software for identification of peptide sequences from fragmentation spectra. We unambiguously identified 1132 proteins. In addition, the phosphoproteome of human urinary exosomes was profiled using the neutral loss scanning acquisition mode of LC-MS/MS. The phosphoproteomic profiling identified 19 phosphorylation sites corresponding to 14 phosphoproteins. The second objective was to analyze urinary exosomes samples isolated from patients with genetic mutations. Polyclonal antibodies were generated to recognize epitopes on the gene products of these genetic mutations, NKCC2 and MRP4. The potential usefulness of urinary exosome analysis was demonstrated using the well-defined renal tubulopathy, Bartter syndrome type I and using the single nucleotide polymorphism in the ABCC4 gene. -

Characterisation of Aspergillus Niger Prolyl Aminopeptidase
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Wageningen University & Research Publications Mol Gen Genomics (2005) 272: 673–679 DOI 10.1007/s00438-004-1094-5 ORIGINAL PAPER Danie¨lle E. J. W. Basten Æ Antoine P. H. A. Moers Albert J. J. van. Ooyen Æ Peter J. Schaap Characterisation of Aspergillus niger prolyl aminopeptidase Received: 29 April 2004 / Accepted: 16 November 2004 / Published online: 15 January 2005 Ó Springer-Verlag 2005 Abstract We have cloned a gene (papA) that encodes a ases and tripeptidases and finally by carboxypeptidases prolyl aminopeptidase from Aspergillus niger. Homolo- and aminopeptidases. The turnover of proteins by pro- gous genes are present in the genomes of the Eurotiales teases provides a ready pool of amino acids as precur- A. nidulans, A. fumigatus and Talaromyces emersonii, sors for the synthesis of new proteins (Bennet and Klich but the gene is not present in the genome of the yeast 1992). Saccharomyces cerevisiae. Cell extracts of strains over- Proteases normally do not hydrolyse bonds adjacent expressing the gene under the control of its own pro- to proline residues. Instead a specialised group of en- moter showed a fourfold to sixfold increase in prolyl zymes has evolved that hydrolyses these bonds. Their aminopeptidase activity, but no change in phenylalanine activity depends on both the isomeric state of the proline or leucine aminopeptidase activity. The overexpressed residue and its position in the peptide chain (Vanhoof enzyme was subsequently purified and characterised. et al. 1995; Cunningham and O’Connor 1997). Proline The enzyme specifically removes N-terminal proline and aminopeptidases (Pap, prolyl iminopeptidase, EC hydroxyproline residues from peptides. -

Catalytic Properties and Inhibition of Proline-Specific Dipeptidyl Peptidases II, IV and VII
UC Berkeley UC Berkeley Previously Published Works Title Catalytic properties and inhibition of proline-specific dipeptidyl peptidases II, IV and VII Permalink https://escholarship.org/uc/item/4nf146zf Journal Biochemical Journal, 371 ISSN 0264-6021 Authors Leiting, B Pryor, K D Wu, J K et al. Publication Date 2003-04-01 Peer reviewed eScholarship.org Powered by the California Digital Library University of California Biochem. J. (2003) 371, 525–532 (Printed in Great Britain) 525 Catalytic properties and inhibition of proline-specific dipeptidyl peptidases II, IV and VII Barbara LEITING*1, KellyAnn D. PRYOR*, Joseph K. WU*, Frank MARSILIO*, Reshma A. PATEL*, Charles S. CRAIK†, Jonathan A. ELLMAN‡, Richard T. CUMMINGS* and Nancy A. THORNBERRY* *Department of Metabolic Disorders, Merck Research Laboratories, Mail code RY50G-236, P.O. Box 2000, Rahway, NJ 07065, U.S.A., †Department of Pharmaceutical Chemistry, University of California, 513 Parnassus Avenue, San Francisco, CA 94143-0446, U.S.A., and ‡Department of Chemistry, University of California, Berkeley, CA 94720, U.S.A. There is currently intense interest in the emerging group of strates and inhibitors for these enzymes, a complete biochemical proline-specific dipeptidases, and their roles in the regulation profile of these enzymes was obtained. The pH profiles, substrate of biological processes. Dipeptidyl peptidase IV (DPP-IV) is specificities as determined by positional scanning, Michaelis– involved in glucose metabolism by contributing to the regulation Menten constants and inhibition profiles for DPP-VII and DPP- of glucagon family peptides and has emerged as a potential target II were shown to be virtually identical, strongly supporting the for the treatment of metabolic diseases. -

Genetic Polymorphism of the Angiotensin-Converting Enzyme (ACE) in Asthmatic Patients
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector RESPIRATORY MEDICINE (1998) 92, 1305-1310 Original Articles Genetic polymorphism of the angiotensin-converting enzyme (ACE) in asthmatic patients H. TOMITA, S. SATO, R. MATSUDA, N. OGISU, T. MORI, T. NIIMI AND S. SHIMIZU 2nd Department of Intemal Medicine, Nagoya City University Medical School, Nagoya, Japan Angiotensin-converting enzyme (ACE) inactivates bradykinin, substance P and neurokinin A, which are believed to play important roles in the pathogenesis of asthma, especially in neurogenic inflammation. It has recently been shown that an insertion (1)ldeletion (D) polymorphism in the ACE gene accounts for variation in serum ACE levels. There are thus three genotypes (insertion homozygote, II; deletion homozygote, DD; heterozygotes, DI). The serum ACE level with the DD type is reported to be about double that of the II type and intermediate in the DI case. In the present study, we examined whether asthma is linked with this ACE gene polymorphism. Seventy-one patients with asthma (27 males and 44 females) and 142 sex- and age-matched healthy controls were determined for their genotype by the polymerase chain reaction (PCR) method. Twenty-five asthmatics demonstrated the II type (352%) 37 the DI type (52.1%), and nine the DD type (12.7%). There were no significant differences in the distributions of genotypes and serum ACE levels between healthy controls and patients. No significant differences were evident in serum IgE levels, age at onset, proportion of atopic type patients and severity of asthma among the three genotypes. -

B Number Gene Name Mrna Intensity Mrna
sample) total list predicted B number Gene name assignment mRNA present mRNA intensity Gene description Protein detected - Membrane protein membrane sample detected (total list) Proteins detected - Functional category # of tryptic peptides # of tryptic peptides # of tryptic peptides detected (membrane b0002 thrA 13624 P 39 P 18 P(m) 2 aspartokinase I, homoserine dehydrogenase I Metabolism of small molecules b0003 thrB 6781 P 9 P 3 0 homoserine kinase Metabolism of small molecules b0004 thrC 15039 P 18 P 10 0 threonine synthase Metabolism of small molecules b0008 talB 20561 P 20 P 13 0 transaldolase B Metabolism of small molecules chaperone Hsp70; DNA biosynthesis; autoregulated heat shock b0014 dnaK 13283 P 32 P 23 0 proteins Cell processes b0015 dnaJ 4492 P 13 P 4 P(m) 1 chaperone with DnaK; heat shock protein Cell processes b0029 lytB 1331 P 16 P 2 0 control of stringent response; involved in penicillin tolerance Global functions b0032 carA 9312 P 14 P 8 0 carbamoyl-phosphate synthetase, glutamine (small) subunit Metabolism of small molecules b0033 carB 7656 P 48 P 17 0 carbamoyl-phosphate synthase large subunit Metabolism of small molecules b0048 folA 1588 P 7 P 1 0 dihydrofolate reductase type I; trimethoprim resistance Metabolism of small molecules peptidyl-prolyl cis-trans isomerase (PPIase), involved in maturation of b0053 surA 3825 P 19 P 4 P(m) 1 GenProt outer membrane proteins (1st module) Cell processes b0054 imp 2737 P 42 P 5 P(m) 5 GenProt organic solvent tolerance Cell processes b0071 leuD 4770 P 10 P 9 0 isopropylmalate -

Methionine Aminopeptidase Emerging Role in Angiogenesis
Chapter 2 Methionine Aminopeptidase Emerging role in angiogenesis Joseph A. Vetro1, Benjamin Dummitt2, and Yie-Hwa Chang2 1Department of Pharmaceutical Chemistry, University of Kansas, 2095 Constant Ave., Lawrence, KS 66047, USA. 2Edward A. Doisy Department of Biochemistry and Molecular Biology, St. Louis University Health Sciences Center, 1402 S. Grand Blvd., St. Louis, MO 63104, USA. Abstract: Angiogenesis, the formation of new blood vessels from existing vasculature, is a key factor in a number of vascular-related pathologies such as the metastasis and growth of solid tumors. Thus, the inhibition of angiogenesis has great potential as a therapeutic modality in the treatment of cancer and other vascular-related diseases. Recent evidence suggests that the inhibition of mammalian methionine aminopeptidase type 2 (MetAP2) catalytic activity in vascular endothelial cells plays an essential role in the pharmacological activity of the most potent small molecule angiogenesis inhibitors discovered to date, the fumagillin class. Methionine aminopeptidase (MetAP, EC 3.4.11.18) catalyzes the non-processive, co-translational hydrolysis of initiator N-terminal methionine when the second residue of the nascent polypeptide is small and uncharged. Initiator Met removal is a ubiquitous and essential modification. Indirect evidence suggests that removal of initiator Met by MetAP is important for the normal function of many proteins involved in DNA repair, signal transduction, cell transformation, secretory vesicle trafficking, and viral capsid assembly and infection. Currently, much effort is focused on understanding the essential nature of methionine aminopeptidase activity and elucidating the role of methionine aminopeptidase type 2 catalytic activity in angiogenesis. In this chapter, we give an overview of the MetAP proteins, outline the importance of initiator Met hydrolysis, and discuss the possible mechanism(s) through which MetAP2 inhibition by the fumagillin class of angiogenesis inhibitors leads to cytostatic growth arrest in vascular endothelial cells. -

Cloud-Clone 16-17
Cloud-Clone - 2016-17 Catalog Description Pack Size Supplier Rupee(RS) ACB028Hu CLIA Kit for Anti-Albumin Antibody (AAA) 96T Cloud-Clone 74750 AEA044Hu ELISA Kit for Anti-Growth Hormone Antibody (Anti-GHAb) 96T Cloud-Clone 74750 AEA255Hu ELISA Kit for Anti-Apolipoprotein Antibodies (AAHA) 96T Cloud-Clone 74750 AEA417Hu ELISA Kit for Anti-Proteolipid Protein 1, Myelin Antibody (Anti-PLP1) 96T Cloud-Clone 74750 AEA421Hu ELISA Kit for Anti-Myelin Oligodendrocyte Glycoprotein Antibody (Anti- 96T Cloud-Clone 74750 MOG) AEA465Hu ELISA Kit for Anti-Sperm Antibody (AsAb) 96T Cloud-Clone 74750 AEA539Hu ELISA Kit for Anti-Myelin Basic Protein Antibody (Anti-MBP) 96T Cloud-Clone 71250 AEA546Hu ELISA Kit for Anti-IgA Antibody 96T Cloud-Clone 71250 AEA601Hu ELISA Kit for Anti-Myeloperoxidase Antibody (Anti-MPO) 96T Cloud-Clone 71250 AEA747Hu ELISA Kit for Anti-Complement 1q Antibody (Anti-C1q) 96T Cloud-Clone 74750 AEA821Hu ELISA Kit for Anti-C Reactive Protein Antibody (Anti-CRP) 96T Cloud-Clone 74750 AEA895Hu ELISA Kit for Anti-Insulin Receptor Antibody (AIRA) 96T Cloud-Clone 74750 AEB028Hu ELISA Kit for Anti-Albumin Antibody (AAA) 96T Cloud-Clone 71250 AEB264Hu ELISA Kit for Insulin Autoantibody (IAA) 96T Cloud-Clone 74750 AEB480Hu ELISA Kit for Anti-Mannose Binding Lectin Antibody (Anti-MBL) 96T Cloud-Clone 88575 AED245Hu ELISA Kit for Anti-Glutamic Acid Decarboxylase Antibodies (Anti-GAD) 96T Cloud-Clone 71250 AEK505Hu ELISA Kit for Anti-Heparin/Platelet Factor 4 Antibodies (Anti-HPF4) 96T Cloud-Clone 71250 CCA005Hu CLIA Kit for Angiotensin II -
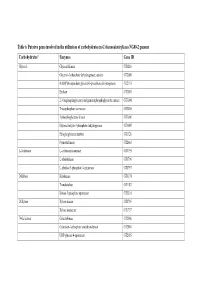
Table 6. Putative Genes Involved in the Utilization of Carbohydrates in G
Table 6. Putative genes involved in the utilization of carbohydrates in G. thermodenitrificans NG80-2 genome Carbohydrates* Enzymes Gene ID Glycerol Glycerol Kinase GT1216 Glycerol-3-phosphate dehydrogenase, aerobic GT2089 NAD(P)H-dependent glycerol-3-phosphate dehydrogenase GT2153 Enolase GT3003 2,3-bisphosphoglycerate-independentphosphoglycerate mutase GT3004 Triosephosphate isomerase GT3005 3-phosphoglycerate kinase GT3006 Glyceraldehyde-3-phosphate dehydrogenase GT3007 Phosphoglycerate mutase GT1326 Pyruvate kinase GT2663 L-Arabinose L-arabinose isomerase GT1795 L-ribulokinase GT1796 L-ribulose 5-phosphate 4-epimerase GT1797 D-Ribose Ribokinase GT3174 Transketolase GT1187 Ribose 5-phosphate epimerase GT3316 D-Xylose Xylose kinase GT1756 Xylose isomerase GT1757 D-Galactose Galactokinase GT2086 Galactose-1-phosphate uridyltransferase GT2084 UDP-glucose 4-epimerase GT2085 Carbohydrates* Enzymes Gene ID D-Fructose 1-phosphofructokinase GT1727 Fructose-1,6-bisphosphate aldolase GT1805 Fructose-1,6-bisphosphate aldolase type II GT3331 Triosephosphate isomerase GT3005 D-Mannose Mannnose-6 phospate isomelase GT3398 6-phospho-1-fructokinase GT2664 D-Mannitol Mannitol-1-phosphate dehydrogenase GT1844 N-Acetylglucosamine N-acetylglucosamine-6-phosphate deacetylase GT2205 N-acetylglucosamine-6-phosphate isomerase GT2204 D-Maltose Alpha-1,4-glucosidase GT0528, GT1643 Sucrose Sucrose phosphorylase GT3215 D-Trehalose Alpha-glucosidase GT1643 Glucose kinase GT2381 Inositol Myo-inositol catabolism protein iolC;5-dehydro-2- GT1807 deoxygluconokinase -

Changes in Proteolysis in Fermented Milk Produced by Streptococcus Thermophilus in Co-Culture with Lactobacillus Plantarum Or Bifidobacterium Animalis Subsp
molecules Article Changes in Proteolysis in Fermented Milk Produced by Streptococcus thermophilus in Co-Culture with Lactobacillus plantarum or Bifidobacterium animalis subsp. lactis During Refrigerated Storage Sining Li 1,2 , Shanhu Tang 1,*, Qiang He 2,*, Jiangxiao Hu 1 and Jing Zheng 1 1 College of Life Science and Technology, Southwest Minzu University, Chengdu 610041, China; [email protected] (S.L.); [email protected] (J.H.); [email protected] (J.Z.) 2 College of Biomass Science and Engineering, Sichuan University, Chengdu 610065, China * Correspondence: [email protected] (S.T.); [email protected] (Q.H.); Tel.: +86-28-85528876 (S.T.); +86-28-85468323 (Q.H.) Received: 1 September 2019; Accepted: 13 October 2019; Published: 15 October 2019 Abstract: Proteolysis in fermented milk, a complex and dynamic process, depends on the starter cultures used. This study aimed to evaluate the influence of Lactobacillus plantarum or Bifidobacterium animalis subsp. lactis, or both, co-fermented with Streptococcus thermophilus, on the changes in the proteolysis profile of fermented milk during 21-day storage at 4 ◦C, including the pH value, proteolytic degree, protease activity, aminopeptidase activity, free amino acid content, and electrophoresis performance. The results showed that the treatments with co-cultures exhibited a higher amount of free amino groups and neutral protease activity at an extracellular level, whereas lower pH values and aminopeptidase activities towards the six substrates at an intracellular level than the ones with a single-strain of S. thermophilus over the refrigerated storage were observed. In co-fermentation with S. thermophilus, B. animalis subsp. lactis did not significantly affect the concentrations of most free amino acids, while contributions of L. -

Characterization of Aminopeptidase in the Free-Living Nematode Panagrellus Redivivus: Subcellular Distribution and Possible Role in Neuropeptide Metabolism E
Journal of Nematology 39(2):153–160. 2007. © The Society of Nematologists 2007. Characterization of Aminopeptidase in the Free-living Nematode Panagrellus redivivus: Subcellular Distribution and Possible Role in Neuropeptide Metabolism E. P. Masler Abstract: Aminopeptidase was detected in homogenates of the free-living nematode Panagrellus redivivus with the aminoacyl substrate L-alanine-4-nitroanilide. Subcellular distribution of activity was 80% soluble and 20% membrane-associated. Aminopep- tidases in the two fractions differed in affinity for Ala-4-NA, with Km’s of 0.65 mM (soluble) and 2.90 mM (membrane). Specific activities (units/mg) at pH 7.8, 27°C were 9.10 (soluble) and 14.30 (membrane). Each enzyme was competitively inhibited by amastatin (90% at 100 µM inhibitor, IC50 = 3.7 µM) and inhibited by puromycin (30% at 500 µM) and 1,10-phenanthroline (IC50’s:; 148 µM, soluble; 89 µM, membrane). Activity was restored by Zn++, with maximum recoveries of 50% (soluble) and 90% (mem- ∼ brane), each at 23 µM ZnCl2. Estimated molecular masses for each were 150 kDa. FMRFamide-like neuropeptides behaved as competitive inhibitors. Modification of the N-terminal F of FMRFamide weakened inhibition by 95%, suggesting that the N-terminus is essential for binding to the enzyme. Two nematode FMRFamides, APKPFIRFa and RNKFEFIRFa, were the most potent tested. This is the first biochemical characterization of aminopeptidase in a free-living nematode other than Caenorhabditis elegans and demon- strates the high selectivity of the P. redivivus enzymes for neuropeptide substrates. Key words: FMRFamide-like peptide, inhibitor; membrane, metallopeptidase, neuropeptide, protease Nematodes, like other eukaryotic organisms, depend reproduction (Day and Maule, 1999; Maule et al., 2002; upon proteolytic enzymes for the regulation of essential Rogers et al., 2003). -

Code Lists Page 1
Code lists Page 1 Code lists AESEV Page 2 AESEV Codelist Name: Severity/Intensity Scale for Adverse Events Description: A scale that defines the degree or state of disease existing in a patient as a result of the occurrence of an adverse event. (NCI) C41338,1; Grade 1 C41339,2; Grade 2 C41340,3; Grade 3 AGEU Page 3 AGEU Codelist Name: Age Unit Description: Those units of time that are routinely used to express the age of a subject. C25301,Days C25529,Hours; h; hr C29846,Month C29844,Week C29848,Year CMDOSFRM Page 4 CMDOSFRM Codelist Name: Concomitant Medication Dose Form Description: A terminology subset of the CDISC SDTM Pharmaceutical Dosage Form codelist created for CDASH Concomitant Medication Dose Form codelist. (NCI) C42887,AEROSOL; aer C25158,CAPSULE; cap C28944,CREAM; C42933,GAS; C42934,GEL; C42966,OINTMENT; oint C42968,PATCH; C42972,POWDER; C42989,SPRAY; C42993,SUPPOSITORY; supp C42994,SUSPENSION; susp C42998,TABLET; tab CMDOSFRQ Page 5 CMDOSFRQ Codelist Name: Concomitant Medication Dosing Frequency per Interval Description: A terminology subset of the CDISC SDTM Frequency codelist created for CDASH Concomitant Medication Dosing Frequency per Interval codelist. (NCI) C64496,BID; BD; Twice per day C64499,PRN; As needed C25473,QD; Daily C64530,QID; 4 times per day C64498,QM; Every Month; Per Month C64525,QOD; Every other day C64527,TID; 3 times per day C17998,UNKNOWN; U; UNK; Unknown CMDOSU Page 6 CMDOSU Codelist Name: Concomitant Medication Dose Units Description: A terminology subset of the CDISC SDTM Unit codelist created for CDASH Concomitant Medication Dose Units codelist. (NCI) C48480,CAPSULE; Capsule Dosing Unit; cap C48155,g; Gram C48579,IU; IE; International Unit C28253,mg; Milligram C28254,mL; Milliliter; cm3 C65060,PUFF; Puff Dosing Unit C48542,TABLET; Tablet Dosing Unit; tab C48152,ug; Microgram; mcg CMROUTE Page 7 CMROUTE Codelist Name: Concomitant Medication Route of Administration Description: A terminology subset of the CDISC SDTM Route codelist created for CDASH Concomitant Medication Route of Administration codelist.