Targeting of the Tail-Anchored Peroxisomal Membrane Proteins PEX26 and PEX15 Occurs Through C-Terminal PEX19-Binding Sites
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
The Peroxisomal PTS1-Import Defect of PEX1- Deficient Cells Is Independent of Pexophagy in Saccharomyces Cerevisiae
International Journal of Molecular Sciences Communication The Peroxisomal PTS1-Import Defect of PEX1- Deficient Cells Is Independent of Pexophagy in Saccharomyces cerevisiae Thomas Mastalski, Rebecca Brinkmeier and Harald W. Platta * Biochemie Intrazellulärer Transportprozesse, Ruhr-Universität Bochum, Universitätsstr. 150, 44801 Bochum, Germany; [email protected] (T.M.); [email protected] (R.B.) * Correspondence: [email protected]; Tel.: +49-234-3224-968 Received: 10 January 2020; Accepted: 27 January 2020; Published: 29 January 2020 Abstract: The important physiologic role of peroxisomes is shown by the occurrence of peroxisomal biogenesis disorders (PBDs) in humans. This spectrum of autosomal recessive metabolic disorders is characterized by defective peroxisome assembly and impaired peroxisomal functions. PBDs are caused by mutations in the peroxisomal biogenesis factors, which are required for the correct compartmentalization of peroxisomal matrix enzymes. Recent work from patient cells that contain the Pex1(G843D) point mutant suggested that the inhibition of the lysosome, and therefore the block of pexophagy, was beneficial for peroxisomal function. The resulting working model proposed that Pex1 may not be essential for matrix protein import at all, but rather for the prevention of pexophagy. Thus, the observed matrix protein import defect would not be caused by a lack of Pex1 activity, but rather by enhanced removal of peroxisomal membranes via pexophagy. In the present study, we can show that the specific block of PEX1 deletion-induced pexophagy does not restore peroxisomal matrix protein import or the peroxisomal function in beta-oxidation in yeast. Therefore, we conclude that Pex1 is directly and essentially involved in peroxisomal matrix protein import, and that the PEX1 deletion-induced pexophagy is not responsible for the defect in peroxisomal function. -

Peroxisomal Disorders and Their Mouse Models Point to Essential Roles of Peroxisomes for Retinal Integrity
International Journal of Molecular Sciences Review Peroxisomal Disorders and Their Mouse Models Point to Essential Roles of Peroxisomes for Retinal Integrity Yannick Das, Daniëlle Swinkels and Myriam Baes * Lab of Cell Metabolism, Department of Pharmaceutical and Pharmacological Sciences, KU Leuven, 3000 Leuven, Belgium; [email protected] (Y.D.); [email protected] (D.S.) * Correspondence: [email protected] Abstract: Peroxisomes are multifunctional organelles, well known for their role in cellular lipid homeostasis. Their importance is highlighted by the life-threatening diseases caused by peroxisomal dysfunction. Importantly, most patients suffering from peroxisomal biogenesis disorders, even those with a milder disease course, present with a number of ocular symptoms, including retinopathy. Patients with a selective defect in either peroxisomal α- or β-oxidation or ether lipid synthesis also suffer from vision problems. In this review, we thoroughly discuss the ophthalmological pathology in peroxisomal disorder patients and, where possible, the corresponding animal models, with a special emphasis on the retina. In addition, we attempt to link the observed retinal phenotype to the underlying biochemical alterations. It appears that the retinal pathology is highly variable and the lack of histopathological descriptions in patients hampers the translation of the findings in the mouse models. Furthermore, it becomes clear that there are still large gaps in the current knowledge on the contribution of the different metabolic disturbances to the retinopathy, but branched chain fatty acid accumulation and impaired retinal PUFA homeostasis are likely important factors. Citation: Das, Y.; Swinkels, D.; Baes, Keywords: peroxisome; Zellweger; metabolism; fatty acid; retina M. Peroxisomal Disorders and Their Mouse Models Point to Essential Roles of Peroxisomes for Retinal Integrity. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Mutations in Novel Peroxin Gene PEX26 That Cause Peroxisome
Am. J. Hum. Genet. 73:233–246, 2003 Mutations in Novel Peroxin Gene PEX26 That Cause Peroxisome-Biogenesis Disorders of Complementation Group 8 Provide a Genotype-Phenotype Correlation Naomi Matsumoto,1,* Shigehiko Tamura,1,* Satomi Furuki,1,* Non Miyata,1 Ann Moser,2 Nobuyuki Shimozawa,3 Hugo W. Moser,2 Yasuyuki Suzuki,3 Naomi Kondo,3 and Yukio Fujiki1,4 1Department of Biology, Faculty of Sciences, Kyushu University Graduate School, Fukuoka, Japan; 2Department of Neurology and Pediatrics, Kennedy-Krieger Institute, Johns Hopkins University, Baltimore; 3Department of Pediatrics, Gifu University School of Medicine, Gifu, Japan; and 4SORST, Japan Science and Technology Corporation, Kawaguchi, Saitama, Japan The human disorders of peroxisome biogenesis (PBDs) are subdivided into 12 complementation groups (CGs). CG8 is one of the more common of these and is associated with varying phenotypes, ranging from the most severe, Zellweger syndrome (ZS), to the milder neonatal adrenoleukodystrophy (NALD) and infantile Refsum disease (IRD). PEX26, encoding the 305-amino-acid membrane peroxin, has been shown to be deficient in CG8. We studied the PEX26 genotype in fibroblasts of eight CG8 patients—four with the ZS phenotype, two with NALD, and two with IRD. Catalase was mostly cytosolic in all these cell lines, but import of the proteins that contained PTS1, the SKL peroxisome targeting sequence, was normal. Expression of PEX26 reestablished peroxisomes in all eight cell lines, confirming that PEX26 defects are pathogenic in CG8 patients. When cells were cultured at 30ЊC, catalase import was restored in the cell lines from patients with the NALD and IRD phenotypes, but to a much lesser extent in those with the ZS phenotype, indicating that temperature sensitivity varied inversely with the severity of the clinical phenotype. -

Table S8. Positively Selected Genes (Psgs) Identified in Glyptosternoid and Yellowhead Catfish Lineages
Table S8. positively selected genes (PSGs) identified in glyptosternoid and yellowhead catfish lineages. Lineage Gene ID Gene name Gene description P-value Corrected P-value G. maculatum ENSDARG00000000001 slc35a5 solute carrier family 35, member A5 0 0 G. maculatum ENSDARG00000000656 psmb9a proteasome (prosome, macropain) subunit, beta type, 9a 0 0 aldo-keto reductase family 7, member A3 (aflatoxin aldehyde G. maculatum ENSDARG00000016649 akr7a3 0 0 reductase) G. maculatum ENSDARG00000017422 apmap adipocyte plasma membrane associated protein 0 0 G. maculatum ENSDARG00000003813 srp54 signal recognition particle 54 0 0 G. maculatum ENSDARG00000016173 cct3 chaperonin containing TCP1, subunit 3 (gamma) 0.012102523 0.049848505 G. maculatum ENSDARG00000018049 sf3b2 splicing factor 3b, subunit 2 0 0 G. maculatum ENSDARG00000004581 sel1l sel-1 suppressor of lin-12-like (C. elegans) 0.004079946 0.01759805 G. maculatum ENSDARG00000020344 slc2a8 solute carrier family 2 (facilitated glucose transporter), member 8 0.00E+00 0 G. maculatum ENSDARG00000011885 mrpl19 mitochondrial ribosomal protein L19 0 0 G. maculatum ENSDARG00000012640 cideb cell death-inducing DFFA-like effector b 0.00112672 0.005077819 G. maculatum ENSDARG00000003127 zgc:123105 zgc:123105 0 0 G. maculatum ENSDARG00000012929 eif2d eukaryotic translation initiation factor 2D 0.001049906 0.004752953 G. maculatum ENSDARG00000012947 SKA2 spindle and kinetochore associated complex subunit 2 0 0 G. maculatum ENSDARG00000015851 pnn pinin, desmosome associated protein 0 0 G. maculatum ENSDARG00000011418 sigmar1 sigma non-opioid intracellular receptor 1 0.00E+00 0 G. maculatum ENSDARG00000006926 btd biotinidase 0 0 G. maculatum ENSDARG00000012674 rpusd4 RNA pseudouridylate synthase domain containing 4 0.00E+00 0 G. maculatum ENSDARG00000017389 igfbp7 insulin-like growth factor binding protein 7 1.05E-02 0.043622349 G. -

Recovery of PEX1-Gly843asp Associated Peroxisome Dysfunction by Flavonoid Compounds in Fibroblasts from Zellweger Spectrum Patients
Recovery of PEX1-Gly843Asp associated peroxisome dysfunction by flavonoid compounds in fibroblasts from Zellweger spectrum patients Gillian Elizabeth MacLean Department of Human Genetics McGill University Montreal, Quebec, Canada Submitted September 2012 A thesis submitted to McGill University in partial fulfillment of the requirements of the degree of Master of Science © Gillian Elizabeth MacLean 2012 TABLE OF CONTENTS TABLE OF CONTENTS .................................................................................................2 ABSTRACT ....................................................................................................................4 RESUME ........................................................................................................................6 ABBREVIATIONS .........................................................................................................8 ACKNOWLEDGEMENTS .............................................................................................9 CHAPTER 1 .................................................................................................................. 11 Introduction ................................................................................................................... 11 Structure and function of PEX1 protein ...................................................................... 13 Recovery of PEX1-G843D by chaperone therapy ....................................................... 14 Objectives of the project............................................................................................ -

Perkinelmer Genomics to Request the Saliva Swab Collection Kit for Patients That Cannot Provide a Blood Sample As Whole Blood Is the Preferred Sample
Zellweger Spectrum Disorder Panel Test Code D4611 Test Summary This panel analyzes 15 genes that have been associated with Zellweger Spectrum Disorders. Turn-Around-Time (TAT)* 3 - 5 weeks Acceptable Sample Types Whole Blood (EDTA) (Preferred sample type) DNA, Isolated Dried Blood Spots Saliva Acceptable Billing Types Self (patient) Payment Institutional Billing Commercial Insurance Indications for Testing This panel may be appropriate for individuals with signs and symptoms of a peroxisomal biogenesis disfuction and/or Zellweger syndrome spectrum disorder and/or a family history of these conditions. Test Description This panel analyzes 15 genes that have been associated with Zellweger Spectrum Disorders. Both sequencing and deletion/duplication (CNV) analysis will be performed on the coding regions of all genes included (unless otherwise marked). All analysis is performed utilizing Next Generation Sequencing (NGS) technology. CNV analysis is designed to detect the majority of deletions and duplications of three exons or greater in size. Smaller CNV events may also be detected and reported, but additional follow-up testing is recommended if a smaller CNV is suspected. All variants are classified according to ACMG guidelines. Condition Description Zellweger syndrome spectrum disorders include Zellweger syndrome (ZS), neonatal adrenoleukodystrophy (NALD), an intermediate form and infantile Refsum disease (IRD). Symptoms may include hypotonia, feeding problems, hearing and vision loss, seizures, distinctive facial characteristics and skeletal abnormalities. Zellweger spectrum disorder is estimated to occur in 1/50,000 individuals. Genes ACOX1, AMACR, HSD17B4, PEX1, PEX10, PEX12, PEX13, PEX14, PEX16, PEX19, PEX2, PEX26, PEX3, PEX5, PEX6 Test Methods and Limitations Sequencing is performed on genomic DNA using an Agilent targeted sequence capture method to enrich for the genes on this panel. -

Hamdan Medical Journal 2012; 5:313–326 (
Hamdan Medical Journal 2012; 5:313–326 (http://dx.doi.org/10.7707/hmj.v5i3.212) REVIEW FOR THE SHEIKH HAMDAN BIN RASHID AL MAKTOUM AWARD FOR MEDICAL SCIENCES Clinical, biochemical and genetic aspects of peroxisomal disorders – an expanding group of genetic diseases in humans Ronald JA Wanders University of Amsterdam, Academic Medical Centre, Department of Clinical Chemistry and Paediatrics, Emma Children’s Hospital, Laboratory Genetic Metabolic Diseases, Amsterdam, the Netherlands Abstract clitoris hypertrophy, camptodactyly and simian creases. Unaware of this publication Smith, Opitz and Zellweger syndrome (ZS) in its classic form is an autosomal recessive lethal 2 disease characterized by the absence of morphologically recognizable Inhorn in 1965 described ‘a syndrome of multiple peroxisomes. Detailed studies on ZS in the early 1980s have led to developmental defects including polycystic kidneys the discovery of a set of peroxisomal biomarkers in blood which has and intrahepatic biliary dysgenesis in two sibs’, revolutionized our knowledge about peroxisomes and peroxisomal disorders, who presented with a large number of comparable and formed the basis for the discovery of the group of peroxisomal diseases defects including severe hypotonia, high forehead, known at present. Peroxisomal disorders are classi!ed into two distinct shallow supraorbital ridges, camptodactyly, minor groups including the disorders of peroxisome biogenesis (group 1) and anomalies of the eyes, ears, palate and hands, peroxisome function (group 2). The enzymatic and molecular basis of most and failure to thrive. Two years later, Passarge and peroxisomal disorders has been identi!ed through the years and pre- and McAdams3 described #ve sisters with similar clinical post-natal diagnostic methods have been established. -

Downloaded Per Proteome Cohort Via the Web- Site Links of Table 1, Also Providing Information on the Deposited Spectral Datasets
www.nature.com/scientificreports OPEN Assessment of a complete and classifed platelet proteome from genome‑wide transcripts of human platelets and megakaryocytes covering platelet functions Jingnan Huang1,2*, Frauke Swieringa1,2,9, Fiorella A. Solari2,9, Isabella Provenzale1, Luigi Grassi3, Ilaria De Simone1, Constance C. F. M. J. Baaten1,4, Rachel Cavill5, Albert Sickmann2,6,7,9, Mattia Frontini3,8,9 & Johan W. M. Heemskerk1,9* Novel platelet and megakaryocyte transcriptome analysis allows prediction of the full or theoretical proteome of a representative human platelet. Here, we integrated the established platelet proteomes from six cohorts of healthy subjects, encompassing 5.2 k proteins, with two novel genome‑wide transcriptomes (57.8 k mRNAs). For 14.8 k protein‑coding transcripts, we assigned the proteins to 21 UniProt‑based classes, based on their preferential intracellular localization and presumed function. This classifed transcriptome‑proteome profle of platelets revealed: (i) Absence of 37.2 k genome‑ wide transcripts. (ii) High quantitative similarity of platelet and megakaryocyte transcriptomes (R = 0.75) for 14.8 k protein‑coding genes, but not for 3.8 k RNA genes or 1.9 k pseudogenes (R = 0.43–0.54), suggesting redistribution of mRNAs upon platelet shedding from megakaryocytes. (iii) Copy numbers of 3.5 k proteins that were restricted in size by the corresponding transcript levels (iv) Near complete coverage of identifed proteins in the relevant transcriptome (log2fpkm > 0.20) except for plasma‑derived secretory proteins, pointing to adhesion and uptake of such proteins. (v) Underrepresentation in the identifed proteome of nuclear‑related, membrane and signaling proteins, as well proteins with low‑level transcripts. -
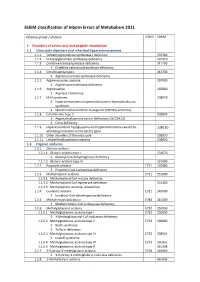
SSIEM Classification of Inborn Errors of Metabolism 2011
SSIEM classification of Inborn Errors of Metabolism 2011 Disease group / disease ICD10 OMIM 1. Disorders of amino acid and peptide metabolism 1.1. Urea cycle disorders and inherited hyperammonaemias 1.1.1. Carbamoylphosphate synthetase I deficiency 237300 1.1.2. N-Acetylglutamate synthetase deficiency 237310 1.1.3. Ornithine transcarbamylase deficiency 311250 S Ornithine carbamoyltransferase deficiency 1.1.4. Citrullinaemia type1 215700 S Argininosuccinate synthetase deficiency 1.1.5. Argininosuccinic aciduria 207900 S Argininosuccinate lyase deficiency 1.1.6. Argininaemia 207800 S Arginase I deficiency 1.1.7. HHH syndrome 238970 S Hyperammonaemia-hyperornithinaemia-homocitrullinuria syndrome S Mitochondrial ornithine transporter (ORNT1) deficiency 1.1.8. Citrullinemia Type 2 603859 S Aspartate glutamate carrier deficiency ( SLC25A13) S Citrin deficiency 1.1.9. Hyperinsulinemic hypoglycemia and hyperammonemia caused by 138130 activating mutations in the GLUD1 gene 1.1.10. Other disorders of the urea cycle 238970 1.1.11. Unspecified hyperammonaemia 238970 1.2. Organic acidurias 1.2.1. Glutaric aciduria 1.2.1.1. Glutaric aciduria type I 231670 S Glutaryl-CoA dehydrogenase deficiency 1.2.1.2. Glutaric aciduria type III 231690 1.2.2. Propionic aciduria E711 232000 S Propionyl-CoA-Carboxylase deficiency 1.2.3. Methylmalonic aciduria E711 251000 1.2.3.1. Methylmalonyl-CoA mutase deficiency 1.2.3.2. Methylmalonyl-CoA epimerase deficiency 251120 1.2.3.3. Methylmalonic aciduria, unspecified 1.2.4. Isovaleric aciduria E711 243500 S Isovaleryl-CoA dehydrogenase deficiency 1.2.5. Methylcrotonylglycinuria E744 210200 S Methylcrotonyl-CoA carboxylase deficiency 1.2.6. Methylglutaconic aciduria E712 250950 1.2.6.1. Methylglutaconic aciduria type I E712 250950 S 3-Methylglutaconyl-CoA hydratase deficiency 1.2.6.2. -

Lineage-Specific Effector Signatures of Invariant NKT Cells Are Shared Amongst Δγ T, Innate Lymphoid, and Th Cells
Downloaded from http://www.jimmunol.org/ by guest on September 26, 2021 δγ is online at: average * The Journal of Immunology , 10 of which you can access for free at: 2016; 197:1460-1470; Prepublished online 6 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600643 http://www.jimmunol.org/content/197/4/1460 Lineage-Specific Effector Signatures of Invariant NKT Cells Are Shared amongst T, Innate Lymphoid, and Th Cells You Jeong Lee, Gabriel J. Starrett, Seungeun Thera Lee, Rendong Yang, Christine M. Henzler, Stephen C. Jameson and Kristin A. Hogquist J Immunol cites 41 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription http://www.jimmunol.org/content/suppl/2016/07/06/jimmunol.160064 3.DCSupplemental This article http://www.jimmunol.org/content/197/4/1460.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 26, 2021. The Journal of Immunology Lineage-Specific Effector Signatures of Invariant NKT Cells Are Shared amongst gd T, Innate Lymphoid, and Th Cells You Jeong Lee,* Gabriel J. -

Peroksisomale Sykdommer V01
2/1/2021 Peroksisomale sykdommer v01 Avdeling for medisinsk genetikk Peroksisomale sykdommer Genpanel, versjon v01 * Enkelte genomiske regioner har lav eller ingen sekvensdekning ved eksomsekvensering. Dette skyldes at de har stor likhet med andre områder i genomet, slik at spesifikk gjenkjennelse av disse områdene og påvisning av varianter i disse områdene, blir vanskelig og upålitelig. Disse genetiske regionene har vi identifisert ved å benytte USCS segmental duplication hvor områder større enn 1 kb og ≥90% likhet med andre regioner i genomet, gjenkjennes (https://genome.ucsc.edu). Vi gjør oppmerksom på at ved identifiseringav ekson oppstrøms for startkodon kan eksonnummereringen endres uten at transkript ID endres. Avdelingens websider har en full oversikt over områder som er affisert av segmentale duplikasjoner. ** Transkriptets kodende ekson. Ekson Gen Gen affisert (HGNC (HGNC Transkript Ekson** Fenotype av symbol) ID) segdup* ABCD1 61 NM_000033.3 7-10 1-10 X-linked adrenoleukodystrophy OMIM ABCD3 67 NM_002858.3 1-23 PMP70 deficiency OMIM ACBD5 23338 NM_145698.4 1-13 Acyl-CoA-binding domain-containing protein 5 deficiency OMIM ACOX1 119 NM_004035.6 1-14 Peroxisomal straight-chain acyl-CoA oxidase deficiency OMIM Pseudo-neonatal adrenoleukodystrophy OMIM ACOX2 120 NM_003500.3 2-15 Peroxisomal branched-chain acyl-CoA oxidase deficiency OMIM Congenital bile acid synthesis defect type 6 OMIM file:///data/Peroksisom_v01-web.html 1/4 2/1/2021 Peroksisomale sykdommer v01 Ekson Gen Gen affisert (HGNC (HGNC Transkript Ekson** Fenotype av symbol) ID) segdup*