Comprehensive Bioinformatics Analysis Identifies POLR2I As a Key
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

POLR2L Antibody Cat
POLR2L Antibody Cat. No.: XW-7445 POLR2L Antibody Specifications HOST SPECIES: Chicken SPECIES REACTIVITY: Human, Mouse, Rat IMMUNOGEN: 1-67 TESTED APPLICATIONS: WB POLR2L antibody can be used for the detection of POLR2L by Western blot, may also work APPLICATIONS: for IHC and ICC. PREDICTED MOLECULAR 7.6 kDa (calculated) WEIGHT: Properties PURIFICATION: Antigen affinity-purified CLONALITY: Polyclonal CONJUGATE: Unconjugated PHYSICAL STATE: Liquid BUFFER: Phosphate-Buffered Saline. No preservatives added. CONCENTRATION: 1 mg/mL October 1, 2021 1 https://www.prosci-inc.com/polr2l-antibody-7445.html POLR2L antibody can be stored at 4˚C for short term (weeks). Long term storage should STORAGE CONDITIONS: be at -20˚C. As with all antibodies care should be taken to avoid repeated freeze thaw cycles. Antibodies should not be exposed to prolonged high temperatures. Additional Info OFFICIAL SYMBOL: POLR2L DNA-directed RNA polymerases I, II, and III subunit RPABC5, DNA-directed RNA ALTERNATE NAMES: polymerase III subunit L, RNA polymerases I, and III subunit ABC5, RBP10, RPB10, RPABC5, RPB7.6, hRPB7.6, RPB10beta, POLR2L ACCESSION NO.: NP_066951.1 PROTEIN GI NO.: 10863925 GENE ID: 5441 USER NOTE: Optimal dilutions for each application to be determined by the researcher. Background and References DNA directed RNA polymerase II polypeptide L; polymerase (RNA) II (DNA directed) polypeptide L (7.6kD); RNA polymerase II subunit. This protein is a subunit of RNA polymerase II, the polymerase responsible for synthesizing messenger RNA in eukaryotes. BACKGROUND: It contains four conserved cysteines characteristic of an atypical zinc-binding domain. Like its counterpart in yeast, this subunit may be shared by the other two DNA-directed RNA polymerases. -

A Network Propagation Approach to Prioritize Long Tail Genes in Cancer
bioRxiv preprint doi: https://doi.org/10.1101/2021.02.05.429983; this version posted February 8, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. A Network Propagation Approach to Prioritize Long Tail Genes in Cancer Hussein Mohsen1,*, Vignesh Gunasekharan2, Tao Qing2, Sahand Negahban3, Zoltan Szallasi4, Lajos Pusztai2,*, Mark B. Gerstein1,5,6,3,* 1 Computational Biology & Bioinformatics Program, Yale University, New Haven, CT 06511, USA 2 Breast Medical Oncology, Yale School of Medicine, New Haven, CT 06511, USA 3 Department of Statistics & Data Science, Yale University, New Haven, CT 06511, USA 4 Children’s Hospital Informatics Program, Harvard-MIT Division of Health Sciences and Technology, Harvard Medical School, Boston, MA 02115, USA 5 Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06511, USA 6 Department of Computer Science, Yale University, New Haven, CT 06511, USA * Corresponding author Abstract Introduction. The diversity of genomic alterations in cancer pose challenges to fully understanding the etiologies of the disease. Recent interest in infrequent mutations, in genes that reside in the “long tail” of the mutational distribution, uncovered new genes with significant implication in cancer development. The study of these genes often requires integrative approaches with multiple types of biological data. Network propagation methods have demonstrated high efficacy in uncovering genomic patterns underlying cancer using biological interaction networks. Yet, the majority of these analyses have focused their assessment on detecting known cancer genes or identifying altered subnetworks. -

Download Download
Supplementary Figure S1. Results of flow cytometry analysis, performed to estimate CD34 positivity, after immunomagnetic separation in two different experiments. As monoclonal antibody for labeling the sample, the fluorescein isothiocyanate (FITC)- conjugated mouse anti-human CD34 MoAb (Mylteni) was used. Briefly, cell samples were incubated in the presence of the indicated MoAbs, at the proper dilution, in PBS containing 5% FCS and 1% Fc receptor (FcR) blocking reagent (Miltenyi) for 30 min at 4 C. Cells were then washed twice, resuspended with PBS and analyzed by a Coulter Epics XL (Coulter Electronics Inc., Hialeah, FL, USA) flow cytometer. only use Non-commercial 1 Supplementary Table S1. Complete list of the datasets used in this study and their sources. GEO Total samples Geo selected GEO accession of used Platform Reference series in series samples samples GSM142565 GSM142566 GSM142567 GSM142568 GSE6146 HG-U133A 14 8 - GSM142569 GSM142571 GSM142572 GSM142574 GSM51391 GSM51392 GSE2666 HG-U133A 36 4 1 GSM51393 GSM51394 only GSM321583 GSE12803 HG-U133A 20 3 GSM321584 2 GSM321585 use Promyelocytes_1 Promyelocytes_2 Promyelocytes_3 Promyelocytes_4 HG-U133A 8 8 3 GSE64282 Promyelocytes_5 Promyelocytes_6 Promyelocytes_7 Promyelocytes_8 Non-commercial 2 Supplementary Table S2. Chromosomal regions up-regulated in CD34+ samples as identified by the LAP procedure with the two-class statistics coded in the PREDA R package and an FDR threshold of 0.5. Functional enrichment analysis has been performed using DAVID (http://david.abcc.ncifcrf.gov/) -

Protein Purification Protein Localization in Vivo Fluorescent Imaging Protein Arrays Real Time Imaging Protein Interactions Protein Trafficking Protein Turnover
Overcoming Challenges of Protein Analysis in Mammalian Systems Danette L. Daniels, Ph.D. Current Technologies for Protein Analysis Biochemical/ In Vivo Proteomic Cell Based Animal Analysis Analysis Models Fluorescent proteins Affinity tags Antibodies How about a system applicable to the all approaches that also addresses limitations of current methods? • Minimal interference with protein of interest • Efficient capture/isolation • Detection/real-time imaging • Differential labeling • High Signal/background HaloTag Platform Biochemical/ In Vivo Proteomic Cell Based Animal Analysis Analysis Models Protein purification Protein localization In vivo fluorescent imaging Protein arrays Real time imaging Protein interactions Protein trafficking Protein turnover HaloTag® HaloCHIP™ HaloLink™ HaloTag® Fluorescent Purification Protein:DNA Protein Arrays Pull-Down Ligands HaloTag is a Genetically Engineered Protein Fusion Tag O Functional Protein of Cl O Interest HT + group Protein of Functional HT O O Interest group . A monomeric , 34 kDa, modified bacterial dehalogenase genetically engineered to covalently bind specific, synthetic HaloTag® ligands . Irreversible, covalent attachment of chemical functionalities . Suitable as either N- or C- terminal fusion Mutagenized HaloTag® Protein Enables Covalent HaloTag®-Ligand Complex Hydrolase (DhaA) HaloTag® Catalytic process Facilitated bond formation T r p 1 0 7 T r p 1 0 7 HaloTag®: • 34kDa protein • Monomeric N N H N 4 1 H N A s n - H H • Single change: C l 4 1 A s n C l 2 1 His272Phe for covalent O R O O - C C bond. 3 R O 1 0 6 A s p 1 0 6 A s p O H O H Covalent bond: H H O O • Stable after N C G l u 1 3 0 C G l u 1 3 0 N - O denaturation. -

Produktinformation
Produktinformation Diagnostik & molekulare Diagnostik Laborgeräte & Service Zellkultur & Verbrauchsmaterial Forschungsprodukte & Biochemikalien Weitere Information auf den folgenden Seiten! See the following pages for more information! Lieferung & Zahlungsart Lieferung: frei Haus Bestellung auf Rechnung SZABO-SCANDIC Lieferung: € 10,- HandelsgmbH & Co KG Erstbestellung Vorauskassa Quellenstraße 110, A-1100 Wien T. +43(0)1 489 3961-0 Zuschläge F. +43(0)1 489 3961-7 [email protected] • Mindermengenzuschlag www.szabo-scandic.com • Trockeneiszuschlag • Gefahrgutzuschlag linkedin.com/company/szaboscandic • Expressversand facebook.com/szaboscandic SANTA CRUZ BIOTECHNOLOGY, INC. POLR2G CRISPR/Cas9 KO Plasmid (h): sc-406059 BACKGROUND APPLICATIONS The Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and POLR2G CRISPR/Cas9 KO Plasmid (h) is recommended for the disruption of CRISPR-associated protein (Cas9) system is an adaptive immune response gene expression in human cells. defense mechanism used by archea and bacteria for the degradation of foreign genetic material (4,6). This mechanism can be repurposed for other 20 nt non-coding RNA sequence: guides Cas9 functions, including genomic engineering for mammalian systems, such as to a specific target location in the genomic DNA gene knockout (KO) (1,2,3,5). CRISPR/Cas9 KO Plasmid products enable the U6 promoter: drives gRNA scaffold: helps Cas9 identification and cleavage of specific genes by utilizing guide RNA (gRNA) expression of gRNA bind to target DNA sequences derived from the Genome-scale CRISPR Knock-Out (GeCKO) v2 library developed in the Zhang Laboratory at the Broad Institute (3,5). Termination signal Green Fluorescent Protein: to visually REFERENCES verify transfection CRISPR/Cas9 Knockout Plasmid CBh (chicken β-Actin 1. Cong, L., et al. -

Molecular Pharmacology of Cancer Therapy in Human Colorectal Cancer by Gene Expression Profiling1,2
[CANCER RESEARCH 63, 6855–6863, October 15, 2003] Molecular Pharmacology of Cancer Therapy in Human Colorectal Cancer by Gene Expression Profiling1,2 Paul A. Clarke,3 Mark L. George, Sandra Easdale, David Cunningham, R. Ian Swift, Mark E. Hill, Diana M. Tait, and Paul Workman Cancer Research UK Centre for Cancer Therapeutics, Institute of Cancer Research, Sutton, Surrey SM2 5NG [P. A. C., M. L. G., S. E., P. W.]; Department of Gastrointestinal Oncology, Royal Marsden Hospital, Sutton, Surrey [D. C., M. E. H., D. M. T.]; and Department of Surgery, Mayday Hospital, Croydon, Surrey [M. L. G., R. I. S.], United Kingdom ABSTRACT ment with a single dose of MMC4 and during a continuous infusion of 5FU. In this study, we report for the first time gene expression Global gene expression profiling has potential for elucidating the com- profiling in cancer patients before, and critically, during the period of plex cellular effects and mechanisms of action of novel targeted anticancer exposure to chemotherapy. We have demonstrated that the approach agents or existing chemotherapeutics for which the precise molecular is feasible, and we have detected a novel molecular response that mechanism of action may be unclear. In this study, decreased expression would not have been predicted from in vitro studies and that would of genes required for RNA and protein synthesis, and for metabolism were have otherwise been missed by conventional approaches. The results detected in rectal cancer biopsies taken from patients during a 5-fluorou- also suggest a possible new therapeutic approach. Overall our obser- racil infusion. Our observations demonstrate that this approach is feasible and can detect responses that may have otherwise been missed by con- vations suggest that gene expression profiling in response to treatment ventional methods. -

POLR2I Antibody (Monoclonal) (M01) Mouse Monoclonal Antibody Raised Against a Partial Recombinant POLR2I
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 POLR2I Antibody (monoclonal) (M01) Mouse monoclonal antibody raised against a partial recombinant POLR2I. Catalog # AT3377a Specification POLR2I Antibody (monoclonal) (M01) - Product Information Application WB, E Primary Accession P36954 Other Accession BC017112 Reactivity Human Host mouse Clonality Monoclonal Isotype IgG1 Kappa Calculated MW 14523 POLR2I Antibody (monoclonal) (M01) - Additional Information Antibody Reactive Against Recombinant Protein.Western Blot detection against Gene ID 5438 Immunogen (36.74 KDa) . Other Names DNA-directed RNA polymerase II subunit RPB9, RNA polymerase II subunit B9, DNA-directed RNA polymerase II subunit I, RNA polymerase II 145 kDa subunit, RPB145, POLR2I Target/Specificity POLR2I (AAH17112, 26 a.a. ~ 125 a.a) partial recombinant protein with GST tag. MW of the GST tag alone is 26 KDa. Detection limit for recombinant GST tagged Dilution POLR2I is approximately 0.03ng/ml as a WB~~1:500~1000 capture antibody. Format Clear, colorless solution in phosphate POLR2I Antibody (monoclonal) (M01) - buffered saline, pH 7.2 . Background Storage This gene encodes a subunit of RNA Store at -20°C or lower. Aliquot to avoid polymerase II, the polymerase responsible for repeated freezing and thawing. synthesizing messenger RNA in eukaryotes. This subunit, in combination with two other Precautions polymerase subunits, forms the DNA binding POLR2I Antibody (monoclonal) (M01) is for domain of the polymerase, a groove in which research use only and not for use in the DNA template is transcribed into RNA. The diagnostic or therapeutic procedures. product of this gene has two zinc finger motifs with conserved cysteines and the subunit does possess zinc binding activity. -
Attachment PDF Icon
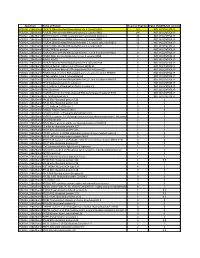
Spectrum Name of Protein Count of Peptides Ratio (POL2RA/IgG control) POLR2A_228kdBand POLR2A DNA-directed RNA polymerase II subunit RPB1 197 NOT IN CONTROL IP POLR2A_228kdBand POLR2B DNA-directed RNA polymerase II subunit RPB2 146 NOT IN CONTROL IP POLR2A_228kdBand RPAP2 Isoform 1 of RNA polymerase II-associated protein 2 24 NOT IN CONTROL IP POLR2A_228kdBand POLR2G DNA-directed RNA polymerase II subunit RPB7 23 NOT IN CONTROL IP POLR2A_228kdBand POLR2H DNA-directed RNA polymerases I, II, and III subunit RPABC3 19 NOT IN CONTROL IP POLR2A_228kdBand POLR2C DNA-directed RNA polymerase II subunit RPB3 17 NOT IN CONTROL IP POLR2A_228kdBand POLR2J RPB11a protein 7 NOT IN CONTROL IP POLR2A_228kdBand POLR2E DNA-directed RNA polymerases I, II, and III subunit RPABC1 8 NOT IN CONTROL IP POLR2A_228kdBand POLR2I DNA-directed RNA polymerase II subunit RPB9 9 NOT IN CONTROL IP POLR2A_228kdBand ALMS1 ALMS1 3 NOT IN CONTROL IP POLR2A_228kdBand POLR2D DNA-directed RNA polymerase II subunit RPB4 6 NOT IN CONTROL IP POLR2A_228kdBand GRINL1A;Gcom1 Isoform 12 of Protein GRINL1A 6 NOT IN CONTROL IP POLR2A_228kdBand RECQL5 Isoform Beta of ATP-dependent DNA helicase Q5 3 NOT IN CONTROL IP POLR2A_228kdBand POLR2L DNA-directed RNA polymerases I, II, and III subunit RPABC5 5 NOT IN CONTROL IP POLR2A_228kdBand KRT6A Keratin, type II cytoskeletal 6A 3 NOT IN CONTROL IP POLR2A_228kdBand POLR2K DNA-directed RNA polymerases I, II, and III subunit RPABC4 2 NOT IN CONTROL IP POLR2A_228kdBand RFC4 Replication factor C subunit 4 1 NOT IN CONTROL IP POLR2A_228kdBand RFC2 -

A High Resolution Physical and RH Map of Pig Chromosome 6Q1.2 And
A high resolution physical and RH map of pig chromosome 6q1.2 and comparative analysis with human chromosome 19q13.1 Flávia Martins-Wess, Denis Milan, Cord Drögemüller, Rodja Voβ-Nemitz, Bertram Brenig, Annie Robic, Martine Yerle, Tosso Leeb To cite this version: Flávia Martins-Wess, Denis Milan, Cord Drögemüller, Rodja Voβ-Nemitz, Bertram Brenig, et al.. A high resolution physical and RH map of pig chromosome 6q1.2 and comparative analysis with human chromosome 19q13.1. BMC Genomics, BioMed Central, 2003, 4, pp.435-444. 10.1186/1471-2164-4- 20. hal-02680244 HAL Id: hal-02680244 https://hal.inrae.fr/hal-02680244 Submitted on 31 May 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. BMC Genomics BioMed Central Research article Open Access A high resolution physical and RH map of pig chromosome 6q1.2 and comparative analysis with human chromosome 19q13.1 Flávia Martins-Wess1, Denis Milan*2, Cord Drögemüller1, Rodja Voβ- Nemitz1, Bertram Brenig3, Annie Robic2, Martine Yerle2 and Tosso Leeb*1 Address: 1Institute of Animal Breeding and Genetics, School of Veterinary Medicine Hannover, Bünteweg 17p, 30559 Hannover, Germany, 2Institut National de la Recherche Agronomique (INRA), Laboratoire de Génétique Cellulaire, BP27, 31326 Castanet Tolosan Cedex, France and 3Institute of Veterinary Medicine, University of Göttingen, Groner Landstr. -

The Human Genome Project
TO KNOW OURSELVES ❖ THE U.S. DEPARTMENT OF ENERGY AND THE HUMAN GENOME PROJECT JULY 1996 TO KNOW OURSELVES ❖ THE U.S. DEPARTMENT OF ENERGY AND THE HUMAN GENOME PROJECT JULY 1996 Contents FOREWORD . 2 THE GENOME PROJECT—WHY THE DOE? . 4 A bold but logical step INTRODUCING THE HUMAN GENOME . 6 The recipe for life Some definitions . 6 A plan of action . 8 EXPLORING THE GENOMIC LANDSCAPE . 10 Mapping the terrain Two giant steps: Chromosomes 16 and 19 . 12 Getting down to details: Sequencing the genome . 16 Shotguns and transposons . 20 How good is good enough? . 26 Sidebar: Tools of the Trade . 17 Sidebar: The Mighty Mouse . 24 BEYOND BIOLOGY . 27 Instrumentation and informatics Smaller is better—And other developments . 27 Dealing with the data . 30 ETHICAL, LEGAL, AND SOCIAL IMPLICATIONS . 32 An essential dimension of genome research Foreword T THE END OF THE ROAD in Little has been rapid, and it is now generally agreed Cottonwood Canyon, near Salt that this international project will produce Lake City, Alta is a place of the complete sequence of the human genome near-mythic renown among by the year 2005. A skiers. In time it may well And what is more important, the value assume similar status among molecular of the project also appears beyond doubt. geneticists. In December 1984, a conference Genome research is revolutionizing biology there, co-sponsored by the U.S. Department and biotechnology, and providing a vital of Energy, pondered a single question: Does thrust to the increasingly broad scope of the modern DNA research offer a way of detect- biological sciences. -

Termination of RNA Polymerase II Transcription by the 5’-3’ Exonuclease Xrn2
TERMINATION OF RNA POLYMERASE II TRANSCRIPTION BY THE 5’-3’ EXONUCLEASE XRN2 by MICHAEL ANDRES CORTAZAR OSORIO B.S., Universidad del Valle – Colombia, 2011 A thesis submitted to the Faculty of the Graduate School of the University of Colorado in partial fulfillment of the requirements for the degree of Doctor of Philosophy Molecular Biology Program 2018 This thesis for the Doctor of Philosophy degree by Michael Andrés Cortázar Osorio has been approved for the Molecular Biology Program by Mair Churchill, Chair Richard Davis Jay Hesselberth Thomas Blumenthal James Goodrich David Bentley, Advisor Date: Aug 17, 2018 ii Cortázar Osorio, Michael Andrés (Ph.D., Molecular Biology) Termination of RNA polymerase II transcription by the 5’-3’ exonuclease Xrn2 Thesis directed by Professor David L. Bentley ABSTRACT Termination of transcription occurs when RNA polymerase (pol) II dissociates from the DNA template and releases a newly-made mRNA molecule. Interestingly, an active debate fueled by conflicting reports over the last three decades is still open on which of the two main models of termination of RNA polymerase II transcription does in fact operate at 3’ ends of genes. The torpedo model indicates that the 5’-3’ exonuclease Xrn2 targets the nascent transcript for degradation after cleavage at the polyA site and chases pol II for termination. In contrast, the allosteric model asserts that transcription through the polyA signal induces a conformational change of the elongation complex and converts it into a termination-competent complex. In this thesis, I propose a unified allosteric-torpedo mechanism. Consistent with a polyA site-dependent conformational change of the elongation complex, I found that pol II transitions at the polyA site into a mode of slow transcription elongation that is accompanied by loss of Spt5 phosphorylation in the elongation complex. -

POLR2I (F-11): Sc-398049
SANTA CRUZ BIOTECHNOLOGY, INC. POLR2I (F-11): sc-398049 BACKGROUND APPLICATIONS RNA polymerase II (Pol II) is a multi-subunit enzyme responsible for the tran- POLR2I (F-11) is recommended for detection of POLR2I of mouse, rat and scription of protein-coding genes. Transcription initiation requires recruitment human origin by Western Blotting (starting dilution 1:100, dilution range of the complete transcription machinery to a promoter via solicitation by 1:100-1:1000), immunoprecipitation [1-2 µg per 100-500 µg of total protein activators and chromatin remodeling factors. Pol II can coordinate 10 to 14 (1 ml of cell lysate)], immunofluorescence (starting dilution 1:50, dilution subunits. This complex interacts with the promoter regions of genes and a range 1:50-1:500), immunohistochemistry (including paraffin-embedded variety of elements and transcription factors. POLR2I (polymerase (RNA) II sections) (starting dilution 1:50, dilution range 1:50-1:500) and solid phase (DNA directed) polypeptide I), also known as RPB9 or hRPB14.5, is a 125 ELISA (starting dilution 1:30, dilution range 1:30-1:3000). amino acid nuclear protein belonging to the archaeal rpoM/eukaryotic RPA12/ POLR2I (F-11) is also recommended for detection of POLR2I in additional RPB9/RPC11 RNA polymerase family. Component of RNA polymerase II, POLR2I species, including equine, canine, bovine and porcine. catalyzes the transcription of DNA into RNA using the four ribonucleoside triphosphates as substrates. POLR2I is part of the upper jaw surrounding the Suitable for use as control antibody for POLR2I siRNA (h): sc-97881, POLR2I central large cleft and is thought to grab the incoming DNA template.