ELOF1 Is a Transcription-Coupled DNA Repair Factor That Directs RNA Polymerase II Ubiquitylation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

POLR2I Antibody (Monoclonal) (M01) Mouse Monoclonal Antibody Raised Against a Partial Recombinant POLR2I
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 POLR2I Antibody (monoclonal) (M01) Mouse monoclonal antibody raised against a partial recombinant POLR2I. Catalog # AT3377a Specification POLR2I Antibody (monoclonal) (M01) - Product Information Application WB, E Primary Accession P36954 Other Accession BC017112 Reactivity Human Host mouse Clonality Monoclonal Isotype IgG1 Kappa Calculated MW 14523 POLR2I Antibody (monoclonal) (M01) - Additional Information Antibody Reactive Against Recombinant Protein.Western Blot detection against Gene ID 5438 Immunogen (36.74 KDa) . Other Names DNA-directed RNA polymerase II subunit RPB9, RNA polymerase II subunit B9, DNA-directed RNA polymerase II subunit I, RNA polymerase II 145 kDa subunit, RPB145, POLR2I Target/Specificity POLR2I (AAH17112, 26 a.a. ~ 125 a.a) partial recombinant protein with GST tag. MW of the GST tag alone is 26 KDa. Detection limit for recombinant GST tagged Dilution POLR2I is approximately 0.03ng/ml as a WB~~1:500~1000 capture antibody. Format Clear, colorless solution in phosphate POLR2I Antibody (monoclonal) (M01) - buffered saline, pH 7.2 . Background Storage This gene encodes a subunit of RNA Store at -20°C or lower. Aliquot to avoid polymerase II, the polymerase responsible for repeated freezing and thawing. synthesizing messenger RNA in eukaryotes. This subunit, in combination with two other Precautions polymerase subunits, forms the DNA binding POLR2I Antibody (monoclonal) (M01) is for domain of the polymerase, a groove in which research use only and not for use in the DNA template is transcribed into RNA. The diagnostic or therapeutic procedures. product of this gene has two zinc finger motifs with conserved cysteines and the subunit does possess zinc binding activity. -
Attachment PDF Icon
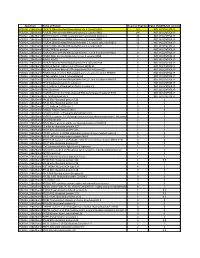
Spectrum Name of Protein Count of Peptides Ratio (POL2RA/IgG control) POLR2A_228kdBand POLR2A DNA-directed RNA polymerase II subunit RPB1 197 NOT IN CONTROL IP POLR2A_228kdBand POLR2B DNA-directed RNA polymerase II subunit RPB2 146 NOT IN CONTROL IP POLR2A_228kdBand RPAP2 Isoform 1 of RNA polymerase II-associated protein 2 24 NOT IN CONTROL IP POLR2A_228kdBand POLR2G DNA-directed RNA polymerase II subunit RPB7 23 NOT IN CONTROL IP POLR2A_228kdBand POLR2H DNA-directed RNA polymerases I, II, and III subunit RPABC3 19 NOT IN CONTROL IP POLR2A_228kdBand POLR2C DNA-directed RNA polymerase II subunit RPB3 17 NOT IN CONTROL IP POLR2A_228kdBand POLR2J RPB11a protein 7 NOT IN CONTROL IP POLR2A_228kdBand POLR2E DNA-directed RNA polymerases I, II, and III subunit RPABC1 8 NOT IN CONTROL IP POLR2A_228kdBand POLR2I DNA-directed RNA polymerase II subunit RPB9 9 NOT IN CONTROL IP POLR2A_228kdBand ALMS1 ALMS1 3 NOT IN CONTROL IP POLR2A_228kdBand POLR2D DNA-directed RNA polymerase II subunit RPB4 6 NOT IN CONTROL IP POLR2A_228kdBand GRINL1A;Gcom1 Isoform 12 of Protein GRINL1A 6 NOT IN CONTROL IP POLR2A_228kdBand RECQL5 Isoform Beta of ATP-dependent DNA helicase Q5 3 NOT IN CONTROL IP POLR2A_228kdBand POLR2L DNA-directed RNA polymerases I, II, and III subunit RPABC5 5 NOT IN CONTROL IP POLR2A_228kdBand KRT6A Keratin, type II cytoskeletal 6A 3 NOT IN CONTROL IP POLR2A_228kdBand POLR2K DNA-directed RNA polymerases I, II, and III subunit RPABC4 2 NOT IN CONTROL IP POLR2A_228kdBand RFC4 Replication factor C subunit 4 1 NOT IN CONTROL IP POLR2A_228kdBand RFC2 -

A High Resolution Physical and RH Map of Pig Chromosome 6Q1.2 And
A high resolution physical and RH map of pig chromosome 6q1.2 and comparative analysis with human chromosome 19q13.1 Flávia Martins-Wess, Denis Milan, Cord Drögemüller, Rodja Voβ-Nemitz, Bertram Brenig, Annie Robic, Martine Yerle, Tosso Leeb To cite this version: Flávia Martins-Wess, Denis Milan, Cord Drögemüller, Rodja Voβ-Nemitz, Bertram Brenig, et al.. A high resolution physical and RH map of pig chromosome 6q1.2 and comparative analysis with human chromosome 19q13.1. BMC Genomics, BioMed Central, 2003, 4, pp.435-444. 10.1186/1471-2164-4- 20. hal-02680244 HAL Id: hal-02680244 https://hal.inrae.fr/hal-02680244 Submitted on 31 May 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. BMC Genomics BioMed Central Research article Open Access A high resolution physical and RH map of pig chromosome 6q1.2 and comparative analysis with human chromosome 19q13.1 Flávia Martins-Wess1, Denis Milan*2, Cord Drögemüller1, Rodja Voβ- Nemitz1, Bertram Brenig3, Annie Robic2, Martine Yerle2 and Tosso Leeb*1 Address: 1Institute of Animal Breeding and Genetics, School of Veterinary Medicine Hannover, Bünteweg 17p, 30559 Hannover, Germany, 2Institut National de la Recherche Agronomique (INRA), Laboratoire de Génétique Cellulaire, BP27, 31326 Castanet Tolosan Cedex, France and 3Institute of Veterinary Medicine, University of Göttingen, Groner Landstr. -

The Human Genome Project
TO KNOW OURSELVES ❖ THE U.S. DEPARTMENT OF ENERGY AND THE HUMAN GENOME PROJECT JULY 1996 TO KNOW OURSELVES ❖ THE U.S. DEPARTMENT OF ENERGY AND THE HUMAN GENOME PROJECT JULY 1996 Contents FOREWORD . 2 THE GENOME PROJECT—WHY THE DOE? . 4 A bold but logical step INTRODUCING THE HUMAN GENOME . 6 The recipe for life Some definitions . 6 A plan of action . 8 EXPLORING THE GENOMIC LANDSCAPE . 10 Mapping the terrain Two giant steps: Chromosomes 16 and 19 . 12 Getting down to details: Sequencing the genome . 16 Shotguns and transposons . 20 How good is good enough? . 26 Sidebar: Tools of the Trade . 17 Sidebar: The Mighty Mouse . 24 BEYOND BIOLOGY . 27 Instrumentation and informatics Smaller is better—And other developments . 27 Dealing with the data . 30 ETHICAL, LEGAL, AND SOCIAL IMPLICATIONS . 32 An essential dimension of genome research Foreword T THE END OF THE ROAD in Little has been rapid, and it is now generally agreed Cottonwood Canyon, near Salt that this international project will produce Lake City, Alta is a place of the complete sequence of the human genome near-mythic renown among by the year 2005. A skiers. In time it may well And what is more important, the value assume similar status among molecular of the project also appears beyond doubt. geneticists. In December 1984, a conference Genome research is revolutionizing biology there, co-sponsored by the U.S. Department and biotechnology, and providing a vital of Energy, pondered a single question: Does thrust to the increasingly broad scope of the modern DNA research offer a way of detect- biological sciences. -

POLR2I (F-11): Sc-398049
SANTA CRUZ BIOTECHNOLOGY, INC. POLR2I (F-11): sc-398049 BACKGROUND APPLICATIONS RNA polymerase II (Pol II) is a multi-subunit enzyme responsible for the tran- POLR2I (F-11) is recommended for detection of POLR2I of mouse, rat and scription of protein-coding genes. Transcription initiation requires recruitment human origin by Western Blotting (starting dilution 1:100, dilution range of the complete transcription machinery to a promoter via solicitation by 1:100-1:1000), immunoprecipitation [1-2 µg per 100-500 µg of total protein activators and chromatin remodeling factors. Pol II can coordinate 10 to 14 (1 ml of cell lysate)], immunofluorescence (starting dilution 1:50, dilution subunits. This complex interacts with the promoter regions of genes and a range 1:50-1:500), immunohistochemistry (including paraffin-embedded variety of elements and transcription factors. POLR2I (polymerase (RNA) II sections) (starting dilution 1:50, dilution range 1:50-1:500) and solid phase (DNA directed) polypeptide I), also known as RPB9 or hRPB14.5, is a 125 ELISA (starting dilution 1:30, dilution range 1:30-1:3000). amino acid nuclear protein belonging to the archaeal rpoM/eukaryotic RPA12/ POLR2I (F-11) is also recommended for detection of POLR2I in additional RPB9/RPC11 RNA polymerase family. Component of RNA polymerase II, POLR2I species, including equine, canine, bovine and porcine. catalyzes the transcription of DNA into RNA using the four ribonucleoside triphosphates as substrates. POLR2I is part of the upper jaw surrounding the Suitable for use as control antibody for POLR2I siRNA (h): sc-97881, POLR2I central large cleft and is thought to grab the incoming DNA template. -

Connectivity Analysis of Single-Cell RNA-Seq Derived Transcriptional
Connectivity analysis of single-cell RNA- seq derived transcriptional signatures A dissertation submitted to the Graduate School of the University of Cincinnati in partial fulfillment of the requirement for the degree of Doctor of Philosophy by Naim Al Mahi M.Sc. In Statistics, Ball State University, IN, 2014 B.Sc. In Statistics, University of Dhaka, Bangladesh, 2011 Committee Members: Mario Medvedovic, PhD (Chair) Jaroslaw Meller, PhD Siva Sivaganesan, PhD Jane Yu, PhD November 7, 2020 Division of Biostatistics and Bioinformatics Department of Environmental and Public Health Sciences University of Cincinnati College of Medicine Cincinnati, OH Abstract With the recent progress in high-throughput sequencing technologies, unprecedented amount of data from multiple omics modalities including genomics, transcriptomics, proteomics, and small-molecule data have become available. Accessibility and reusability of these biomedical big data provide immense opportunities and challenges as well. For example, integrating and connecting multiple data modalities such as, connecting single-cell transcriptomics data to small-molecule perturbational data can enhance the understanding of complex mechanisms underlie a disease and identify potential therapeutics. Recently released LINCS (Library of Integrated Network-based Cellular Signatures)-L1000 transcriptional signatures of chemical perturbations has opened new avenues to study cellular responses to existing drugs and new bioactive compounds. Connecting transcriptional signature of a disease to these chemical perturbation signatures to identify bioactive chemicals that can “revert” the disease signatures can lead to novel drug discovery and drug repurposing. Although, considerable research has been devoted to utilize bulk assay transcriptional data to study the relationship between diseases, genes, and drugs, considerably less attention has been paid to study the connection at the single cell level. -

CREB-Dependent Transcription in Astrocytes: Signalling Pathways, Gene Profiles and Neuroprotective Role in Brain Injury
CREB-dependent transcription in astrocytes: signalling pathways, gene profiles and neuroprotective role in brain injury. Tesis doctoral Luis Pardo Fernández Bellaterra, Septiembre 2015 Instituto de Neurociencias Departamento de Bioquímica i Biologia Molecular Unidad de Bioquímica y Biologia Molecular Facultad de Medicina CREB-dependent transcription in astrocytes: signalling pathways, gene profiles and neuroprotective role in brain injury. Memoria del trabajo experimental para optar al grado de doctor, correspondiente al Programa de Doctorado en Neurociencias del Instituto de Neurociencias de la Universidad Autónoma de Barcelona, llevado a cabo por Luis Pardo Fernández bajo la dirección de la Dra. Elena Galea Rodríguez de Velasco y la Dra. Roser Masgrau Juanola, en el Instituto de Neurociencias de la Universidad Autónoma de Barcelona. Doctorando Directoras de tesis Luis Pardo Fernández Dra. Elena Galea Dra. Roser Masgrau In memoriam María Dolores Álvarez Durán Abuela, eres la culpable de que haya decidido recorrer el camino de la ciencia. Que estas líneas ayuden a conservar tu recuerdo. A mis padres y hermanos, A Meri INDEX I Summary 1 II Introduction 3 1 Astrocytes: physiology and pathology 5 1.1 Anatomical organization 6 1.2 Origins and heterogeneity 6 1.3 Astrocyte functions 8 1.3.1 Developmental functions 8 1.3.2 Neurovascular functions 9 1.3.3 Metabolic support 11 1.3.4 Homeostatic functions 13 1.3.5 Antioxidant functions 15 1.3.6 Signalling functions 15 1.4 Astrocytes in brain pathology 20 1.5 Reactive astrogliosis 22 2 The transcription -

Using Network Inference to Discover Molecular Pathways Underlying Cytokine Synergism and Age-Related Neurodegeneration by Bryce Hwang S.B., M.I.T
Using network inference to discover molecular pathways underlying cytokine synergism and age-related neurodegeneration by Bryce Hwang S.B., M.I.T. (2018) Submitted to the Department of Electrical Engineering and Computer Science in partial fulfillment of the requirements for the degree of Master of Engineering in Computer Science and Molecular Biology at the MASSACHUSETTS INSTITUTE OF TECHNOLOGY June 2018 ○c Bryce Hwang. All rights reserved. The author hereby grants to M.I.T. permission to reproduce and to distribute publicly paper and electronic copies of this thesis document in whole or in part in any medium now known or hereafter created. Author.............................................................. Department of Electrical Engineering and Computer Science May 25, 2018 Certified by. Ernest Fraenkel Professor Thesis Supervisor Accepted by . Katrina LaCurts Chair, Master of Engineering Thesis Committee Using network inference to discover molecular pathways underlying cytokine synergism and age-related neurodegeneration by Bryce Hwang Submitted to the Department of Electrical Engineering and Computer Science on May 25, 2018, in partial fulfillment of the requirements for the degree of Master of Engineering in Computer Science and Molecular Biology Abstract New high-throughput “omic” methods can help shed light on molecular pathways underpinning diseases ranging from cancers to neurodegenerative disorders. However, effectively integrating information across these diverse data types is challenging. Network modeling approaches can help bridge this gap. In particular, the Prize- Collecting Steiner Forest approach (PCSF) is a network modeling method that provides high-confidence subnetworks of physically interacting molecules by integrating diverse “omics” data with prior knowledge from protein-protein interaction networks (PPIs). However, PCSF is sensitive to initial parameterization and generating biological hypotheses from the resulting subnetworks can often be difficult. -

POLR2I Human Shrna Plasmid Kit (Locus ID 5438
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TR316665 RNA Polymerase II p14.5 (POLR2I) Human shRNA Plasmid Kit (Locus ID 5438) Product data: Product Type: shRNA Plasmids Product Name: RNA Polymerase II p14.5 (POLR2I) Human shRNA Plasmid Kit (Locus ID 5438) Locus ID: 5438 Synonyms: hRPB14.5; RPB9 Vector: pRS (TR20003) Format: Retroviral plasmids Components: POLR2I - Human, 4 unique 29mer shRNA constructs in retroviral untagged vector(Gene ID = 5438). 5µg purified plasmid DNA per construct Non-effective 29-mer scrambled shRNA cassette in pRS Vector, TR30012, included for free. RefSeq: NM_006233, NM_006233.1, NM_006233.2, NM_006233.3, NM_006233.4, BC067794, BC067794.1, BC017112, BC017860, NM_006233.5 Summary: This gene encodes a subunit of RNA polymerase II, the polymerase responsible for synthesizing messenger RNA in eukaryotes. This subunit, in combination with two other polymerase subunits, forms the DNA binding domain of the polymerase, a groove in which the DNA template is transcribed into RNA. The product of this gene has two zinc finger motifs with conserved cysteines and the subunit does possess zinc binding activity. [provided by RefSeq, Jul 2008] shRNA Design: These shRNA constructs were designed against multiple splice variants at this gene locus. To be certain that your variant of interest is targeted, please contact [email protected]. If you need a special design or shRNA sequence, please utilize our custom shRNA service. This product is to be used for laboratory only. -

Systematic Identification of Transcriptional and Post-Transcriptional
Liu et al. BMC Bioinformatics 2014, 15:336 http://www.biomedcentral.com/1471-2105/15/336 METHODOLOGY ARTICLE Open Access Systematic identification of transcriptional and post-transcriptional regulations in human respiratory epithelial cells during influenza A virus infection Zhi-Ping Liu1, Hulin Wu2, Jian Zhu3* and Hongyu Miao2* Abstract Background: Respiratory epithelial cells are the primary target of influenza virus infection in human. However, the molecular mechanisms of airway epithelial cell responses to viral infection are not fully understood. Revealing genome-wide transcriptional and post-transcriptional regulatory relationships can further advance our understanding of this problem, which motivates the development of novel and more efficient computational methods to simultaneously infer the transcriptional and post-transcriptional regulatory networks. Results: Here we propose a novel framework named SITPR to investigate the interactions among transcription factors (TFs), microRNAs (miRNAs) and target genes. Briefly, a background regulatory network on a genome-wide scale (~23,000 nodes and ~370,000 potential interactions) is constructed from curated knowledge and algorithm predictions, to which the identification of transcriptional and post-transcriptional regulatory relationships is anchored. To reduce the dimension of the associated computing problem down to an affordable size, several topological and data-based approaches are used. Furthermore, we propose the constrained LASSO formulation and combine it with the dynamic Bayesian network (DBN) model to identify the activated regulatory relationships from time-course expression data. Our simulation studies on networks of different sizes suggest that the proposed framework can effectively determine the genuine regulations among TFs, miRNAs and target genes; also, we compare SITPR with several selected state-of-the-art algorithms to further evaluate its performance. -

GSGS: a Computational Framework to Reconstruct Signaling Pathways
GSGS: A Computational Framework to Reconstruct Signaling Pathways from Gene Sets Lipi Acharya§, Thair Judeh∗,†, Zhansheng Duan♭, Michael Rabbat‡ and Dongxiao Zhu∗,†,¶ ∗Department of Computer Science, Wayne State University, 5057 Woodward Avenue, Detroit, MI 48202, USA. §Department of Computer Science, University of New Orleans, 2000 Lakeshore Drive, New Orleans, LA 70148, USA. †Research Institute for Children, Children’s Hospital, New Orleans, LA 70118, USA. ♭Center for Information Engineering Science Research, Xi’an Jiaotong University, 28 Xianning West Road, Xi’an, Shaanxi 710049, China. ‡Department of Electrical and Computer Engineering, McGill University, 3480 University Street, Montr´eal, Qu´ebec H3A 2A7, Canada. ¶To whom correspondence should be addressed. abstract We propose a novel two-stage Gene Set Gibbs Sampling (GSGS) framework, to reverse engineer signaling pathways from gene sets inferred from molecular profiling data. We hy- pothesize that signaling pathways are structurally an ensemble of overlapping linear signal transduction events which we encode as Information Flow Gene Sets (IFGS’s). We infer pathways from gene sets corresponding to these events subjected to a random permutation of genes within each set. In Stage I, we use a source separation algorithm to derive un- ordered and overlapping IFGS’s from molecular profiling data, allowing cross talk among IFGS’s. In Stage II, we develop a Gibbs sampling like algorithm, Gene Set Gibbs Sampler, arXiv:1101.3983v3 [q-bio.QM] 13 Oct 2011 to reconstruct signaling pathways from the latent IFGS’s derived in Stage I. The novelty of this framework lies in the seamless integration of the two stages and the hypothesis of IFGS’s as the basic building blocks for signal pathways. -

Comprehensive Bioinformatics Analysis Identifies POLR2I As a Key
fgene-12-698570 July 30, 2021 Time: 16:49 # 1 ORIGINAL RESEARCH published: 05 August 2021 doi: 10.3389/fgene.2021.698570 Comprehensive Bioinformatics Analysis Identifies POLR2I as a Key Gene in the Pathogenesis of Hypertensive Nephropathy Shilong You, Jiaqi Xu, Boquan Wu, Shaojun Wu, Ying Zhang*, Yingxian Sun* and Naijin Zhang* Department of Cardiology, The First Hospital of China Medical University, Shenyang, China Hypertensive nephropathy (HN), mainly caused by chronic hypertension, is one of the major causes of end-stage renal disease. However, the pathogenesis of HN remains unclarified, and there is an urgent need for improved treatments. Gene expression profiles for HN and normal tissue were obtained from the Gene Expression Omnibus Edited by: database. A total of 229 differentially co-expressed genes were identified by weighted Zhi-Ping Liu, Shandong University, China gene co-expression network analysis and differential gene expression analysis. These Reviewed by: genes were used to construct protein–protein interaction networks to search for hub Mohadeseh Zarei Ghobadi, genes. Following validation in an independent external dataset and in a clinical database, University of Tehran, Iran POLR2I, one of the hub genes, was identified as a key gene related to the pathogenesis Menghui Yao, First Affiliated Hospital of Zhengzhou of HN. The expression level of POLR2I is upregulated in HN, and the up-regulation of University, China POLR2I is positively correlated with renal function in HN. Finally, we verified the protein *Correspondence: levels of POLR2I in vivo to confirm the accuracy of our analysis. In conclusion, our Naijin Zhang [email protected] study identified POLR2I as a key gene related to the pathogenesis of HN, providing new Yingxian Sun insights into the molecular mechanisms underlying HN.