Connectivity Analysis of Single-Cell RNA-Seq Derived Transcriptional
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Essentiality Landscape and Druggable Oncogenic Dependencies in Herpesviral Primary Effusion Lymphoma
ARTICLE DOI: 10.1038/s41467-018-05506-9 OPEN Gene essentiality landscape and druggable oncogenic dependencies in herpesviral primary effusion lymphoma Mark Manzano1, Ajinkya Patil1, Alexander Waldrop2, Sandeep S. Dave2, Amir Behdad3 & Eva Gottwein1 Primary effusion lymphoma (PEL) is caused by Kaposi’s sarcoma-associated herpesvirus. Our understanding of PEL is poor and therefore treatment strategies are lacking. To address this 1234567890():,; need, we conducted genome-wide CRISPR/Cas9 knockout screens in eight PEL cell lines. Integration with data from unrelated cancers identifies 210 genes as PEL-specific oncogenic dependencies. Genetic requirements of PEL cell lines are largely independent of Epstein-Barr virus co-infection. Genes of the NF-κB pathway are individually non-essential. Instead, we demonstrate requirements for IRF4 and MDM2. PEL cell lines depend on cellular cyclin D2 and c-FLIP despite expression of viral homologs. Moreover, PEL cell lines are addicted to high levels of MCL1 expression, which are also evident in PEL tumors. Strong dependencies on cyclin D2 and MCL1 render PEL cell lines highly sensitive to palbociclib and S63845. In summary, this work comprehensively identifies genetic dependencies in PEL cell lines and identifies novel strategies for therapeutic intervention. 1 Department of Microbiology-Immunology, Feinberg School of Medicine, Northwestern University, Chicago, IL 60611, USA. 2 Duke Cancer Institute and Center for Genomic and Computational Biology, Duke University, Durham, NC 27708, USA. 3 Department of Pathology, Feinberg School of Medicine, Northwestern University, Chicago, IL 60611, USA. Correspondence and requests for materials should be addressed to E.G. (email: [email protected]) NATURE COMMUNICATIONS | (2018) 9:3263 | DOI: 10.1038/s41467-018-05506-9 | www.nature.com/naturecommunications 1 ARTICLE NATURE COMMUNICATIONS | DOI: 10.1038/s41467-018-05506-9 he human oncogenic γ-herpesvirus Kaposi’s sarcoma- (IRF4), a critical oncogene in multiple myeloma33. -

Large-Scale Image-Based Profiling of Single-Cell Phenotypes in Arrayed CRISPR-Cas9 Gene Perturbation Screens
Published online: January 23, 2018 Method Large-scale image-based profiling of single-cell phenotypes in arrayed CRISPR-Cas9 gene perturbation screens Reinoud de Groot1 , Joel Lüthi1,2 , Helen Lindsay1 , René Holtackers1 & Lucas Pelkmans1,* Abstract this reason, CRISPR-Cas9 has been used in large-scale functional genomic screens (Shalem et al, 2015). Most screens performed to High-content imaging using automated microscopy and computer date employ a pooled screening strategy, which can identify genes vision allows multivariate profiling of single-cell phenotypes. Here, that cause differential growth in screening conditions (Koike-Yusa we present methods for the application of the CISPR-Cas9 system et al, 2013; Shalem et al, 2014; Wang et al, 2014). However, in large-scale, image-based, gene perturbation experiments. We pooled screening precludes multivariate profiling of single-cell show that CRISPR-Cas9-mediated gene perturbation can be phenotypes. This can be partially overcome by combining pooled achieved in human tissue culture cells in a timeframe that is screening with single-cell RNA-seq, but this does not easily scale compatible with image-based phenotyping. We developed a pipe- to the profiling of thousands of single cells from thousands of line to construct a large-scale arrayed library of 2,281 sequence- perturbations, and is limited to features that can be read from verified CRISPR-Cas9 targeting plasmids and profiled this library RNA transcript profiles (Adamson et al, 2016; Dixit et al, 2016; for genes affecting cellular morphology and the subcellular local- Jaitin et al, 2016; Datlinger et al, 2017). Moreover, sequencing- ization of components of the nuclear pore complex (NPC). -

Comparative Proteomics Analysis of Human Liver Microsomes and S9
DMD Fast Forward. Published on November 7, 2019 as DOI: 10.1124/dmd.119.089235 This article has not been copyedited and formatted. The final version may differ from this version. DMD # 89235 Comparative Proteomics Analysis of Human Liver Microsomes and S9 Fractions Xinwen Wang, Bing He, Jian Shi, Qian Li, and Hao-Jie Zhu Department of Clinical Pharmacy, University of Michigan, Ann Arbor, Michigan (X.W., B.H., J.S., H.-J.Z.); and School of Life Science and Technology, China Pharmaceutical University, Nanjing, Jiangsu, 210009 (Q.L.) Downloaded from dmd.aspetjournals.org at ASPET Journals on October 2, 2021 1 DMD Fast Forward. Published on November 7, 2019 as DOI: 10.1124/dmd.119.089235 This article has not been copyedited and formatted. The final version may differ from this version. DMD # 89235 Running title: Comparative Proteomics of Human Liver Microsomes and S9 Corresponding author: Hao-Jie Zhu Ph.D. Department of Clinical Pharmacy University of Michigan College of Pharmacy 428 Church Street, Room 4565 Downloaded from Ann Arbor, MI 48109-1065 Tel: 734-763-8449, E-mail: [email protected] dmd.aspetjournals.org Number of words in Abstract: 250 at ASPET Journals on October 2, 2021 Number of words in Introduction: 776 Number of words in Discussion: 2304 2 DMD Fast Forward. Published on November 7, 2019 as DOI: 10.1124/dmd.119.089235 This article has not been copyedited and formatted. The final version may differ from this version. DMD # 89235 Non-standard ABBreviations: DMEs, drug metabolism enzymes; HLM, human liver microsomes; HLS9, -

POLR2I Antibody (Monoclonal) (M01) Mouse Monoclonal Antibody Raised Against a Partial Recombinant POLR2I
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 POLR2I Antibody (monoclonal) (M01) Mouse monoclonal antibody raised against a partial recombinant POLR2I. Catalog # AT3377a Specification POLR2I Antibody (monoclonal) (M01) - Product Information Application WB, E Primary Accession P36954 Other Accession BC017112 Reactivity Human Host mouse Clonality Monoclonal Isotype IgG1 Kappa Calculated MW 14523 POLR2I Antibody (monoclonal) (M01) - Additional Information Antibody Reactive Against Recombinant Protein.Western Blot detection against Gene ID 5438 Immunogen (36.74 KDa) . Other Names DNA-directed RNA polymerase II subunit RPB9, RNA polymerase II subunit B9, DNA-directed RNA polymerase II subunit I, RNA polymerase II 145 kDa subunit, RPB145, POLR2I Target/Specificity POLR2I (AAH17112, 26 a.a. ~ 125 a.a) partial recombinant protein with GST tag. MW of the GST tag alone is 26 KDa. Detection limit for recombinant GST tagged Dilution POLR2I is approximately 0.03ng/ml as a WB~~1:500~1000 capture antibody. Format Clear, colorless solution in phosphate POLR2I Antibody (monoclonal) (M01) - buffered saline, pH 7.2 . Background Storage This gene encodes a subunit of RNA Store at -20°C or lower. Aliquot to avoid polymerase II, the polymerase responsible for repeated freezing and thawing. synthesizing messenger RNA in eukaryotes. This subunit, in combination with two other Precautions polymerase subunits, forms the DNA binding POLR2I Antibody (monoclonal) (M01) is for domain of the polymerase, a groove in which research use only and not for use in the DNA template is transcribed into RNA. The diagnostic or therapeutic procedures. product of this gene has two zinc finger motifs with conserved cysteines and the subunit does possess zinc binding activity. -
Attachment PDF Icon
Spectrum Name of Protein Count of Peptides Ratio (POL2RA/IgG control) POLR2A_228kdBand POLR2A DNA-directed RNA polymerase II subunit RPB1 197 NOT IN CONTROL IP POLR2A_228kdBand POLR2B DNA-directed RNA polymerase II subunit RPB2 146 NOT IN CONTROL IP POLR2A_228kdBand RPAP2 Isoform 1 of RNA polymerase II-associated protein 2 24 NOT IN CONTROL IP POLR2A_228kdBand POLR2G DNA-directed RNA polymerase II subunit RPB7 23 NOT IN CONTROL IP POLR2A_228kdBand POLR2H DNA-directed RNA polymerases I, II, and III subunit RPABC3 19 NOT IN CONTROL IP POLR2A_228kdBand POLR2C DNA-directed RNA polymerase II subunit RPB3 17 NOT IN CONTROL IP POLR2A_228kdBand POLR2J RPB11a protein 7 NOT IN CONTROL IP POLR2A_228kdBand POLR2E DNA-directed RNA polymerases I, II, and III subunit RPABC1 8 NOT IN CONTROL IP POLR2A_228kdBand POLR2I DNA-directed RNA polymerase II subunit RPB9 9 NOT IN CONTROL IP POLR2A_228kdBand ALMS1 ALMS1 3 NOT IN CONTROL IP POLR2A_228kdBand POLR2D DNA-directed RNA polymerase II subunit RPB4 6 NOT IN CONTROL IP POLR2A_228kdBand GRINL1A;Gcom1 Isoform 12 of Protein GRINL1A 6 NOT IN CONTROL IP POLR2A_228kdBand RECQL5 Isoform Beta of ATP-dependent DNA helicase Q5 3 NOT IN CONTROL IP POLR2A_228kdBand POLR2L DNA-directed RNA polymerases I, II, and III subunit RPABC5 5 NOT IN CONTROL IP POLR2A_228kdBand KRT6A Keratin, type II cytoskeletal 6A 3 NOT IN CONTROL IP POLR2A_228kdBand POLR2K DNA-directed RNA polymerases I, II, and III subunit RPABC4 2 NOT IN CONTROL IP POLR2A_228kdBand RFC4 Replication factor C subunit 4 1 NOT IN CONTROL IP POLR2A_228kdBand RFC2 -

A High Resolution Physical and RH Map of Pig Chromosome 6Q1.2 And
A high resolution physical and RH map of pig chromosome 6q1.2 and comparative analysis with human chromosome 19q13.1 Flávia Martins-Wess, Denis Milan, Cord Drögemüller, Rodja Voβ-Nemitz, Bertram Brenig, Annie Robic, Martine Yerle, Tosso Leeb To cite this version: Flávia Martins-Wess, Denis Milan, Cord Drögemüller, Rodja Voβ-Nemitz, Bertram Brenig, et al.. A high resolution physical and RH map of pig chromosome 6q1.2 and comparative analysis with human chromosome 19q13.1. BMC Genomics, BioMed Central, 2003, 4, pp.435-444. 10.1186/1471-2164-4- 20. hal-02680244 HAL Id: hal-02680244 https://hal.inrae.fr/hal-02680244 Submitted on 31 May 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. BMC Genomics BioMed Central Research article Open Access A high resolution physical and RH map of pig chromosome 6q1.2 and comparative analysis with human chromosome 19q13.1 Flávia Martins-Wess1, Denis Milan*2, Cord Drögemüller1, Rodja Voβ- Nemitz1, Bertram Brenig3, Annie Robic2, Martine Yerle2 and Tosso Leeb*1 Address: 1Institute of Animal Breeding and Genetics, School of Veterinary Medicine Hannover, Bünteweg 17p, 30559 Hannover, Germany, 2Institut National de la Recherche Agronomique (INRA), Laboratoire de Génétique Cellulaire, BP27, 31326 Castanet Tolosan Cedex, France and 3Institute of Veterinary Medicine, University of Göttingen, Groner Landstr. -

Functional Gene Clusters in Global Pathogenesis of Clear Cell Carcinoma of the Ovary Discovered by Integrated Analysis of Transcriptomes
International Journal of Environmental Research and Public Health Article Functional Gene Clusters in Global Pathogenesis of Clear Cell Carcinoma of the Ovary Discovered by Integrated Analysis of Transcriptomes Yueh-Han Hsu 1,2, Peng-Hui Wang 1,2,3,4,5 and Chia-Ming Chang 1,2,* 1 Department of Obstetrics and Gynecology, Taipei Veterans General Hospital, Taipei 112, Taiwan; [email protected] (Y.-H.H.); [email protected] (P.-H.W.) 2 School of Medicine, National Yang-Ming University, Taipei 112, Taiwan 3 Institute of Clinical Medicine, National Yang-Ming University, Taipei 112, Taiwan 4 Department of Medical Research, China Medical University Hospital, Taichung 440, Taiwan 5 Female Cancer Foundation, Taipei 104, Taiwan * Correspondence: [email protected]; Tel.: +886-2-2875-7826; Fax: +886-2-5570-2788 Received: 27 April 2020; Accepted: 31 May 2020; Published: 2 June 2020 Abstract: Clear cell carcinoma of the ovary (ovarian clear cell carcinoma (OCCC)) is one epithelial ovarian carcinoma that is known to have a poor prognosis and a tendency for being refractory to treatment due to unclear pathogenesis. Published investigations of OCCC have mainly focused only on individual genes and lack of systematic integrated research to analyze the pathogenesis of OCCC in a genome-wide perspective. Thus, we conducted an integrated analysis using transcriptome datasets from a public domain database to determine genes that may be implicated in the pathogenesis involved in OCCC carcinogenesis. We used the data obtained from the National Center for Biotechnology Information (NCBI) Gene Expression Omnibus (GEO) DataSets. We found six interactive functional gene clusters in the pathogenesis network of OCCC, including ribosomal protein, eukaryotic translation initiation factors, lactate, prostaglandin, proteasome, and insulin-like growth factor. -

Comparative Transcriptomics Identifies Potential Stemness-Related Markers for Mesenchymal Stromal/Stem Cells
bioRxiv preprint doi: https://doi.org/10.1101/2021.05.25.445659; this version posted May 26, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Comparative Transcriptomics Identifies Potential Stemness-Related Markers for Mesenchymal Stromal/Stem Cells Authors Myret Ghabriel 1, Ahmed El Hosseiny 1, 2, Ahmed Moustafa*1, 2 and Asma Amleh*1, 2 Affiliations 1Biotechnology Program, American University in Cairo, New Cairo 11835, Egypt 2Department of Biology, American University in Cairo, New Cairo 11835, Egypt *Corresponding authors: Ahmed Moustafa [email protected] Asma Amleh [email protected]. Abstract Mesenchymal stromal/stem cells (MSCs) are multipotent cells residing in multiple tissues with the capacity for self-renewal and differentiation into various cell types. These properties make them promising candidates for regenerative therapies. MSC identification is critical in yielding pure populations for successful therapeutic applications; however, the criteria for MSC identification proposed by the International Society for Cellular Therapy (ISCT) is inconsistent across different tissue sources. In this study, we aimed to identify potential markers to be used together with the ISCT’s criteria to provide a more accurate means of MSC identification. Thus, we carried out a comparative analysis of the expression of human and mouse MSCs derived from multiple tissues to identify the common differentially expressed genes. We show that six members of the proteasome degradation system are similarly expressed across MSCs derived from bone marrow, adipose tissue, amnion, and umbilical cord. -

Genome-Wide Transcript and Protein Analysis Reveals Distinct Features of Aging in the Mouse Heart
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.28.272260; this version posted April 21, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Genome-wide transcript and protein analysis reveals distinct features of aging in the mouse heart Isabela Gerdes Gyuricza1, Joel M. Chick2, Gregory R. Keele1, Andrew G. Deighan1, Steven C. Munger1, Ron Korstanje1, Steven P. Gygi3, Gary A. Churchill1 1The Jackson Laboratory, Bar Harbor, Maine 04609 USA; 2Vividion Therapeutics, San Diego, California 92121, USA; 3Harvard Medical School, Boston, Massachusetts 02115, USA Corresponding author: [email protected] Key words for online indexing: Heart Aging Transcriptomics Proteomics eQTL pQTL Stoichiometry ABSTRACT Investigation of the molecular mechanisms of aging in the human heart is challenging due to confounding factors, such as diet and medications, as well limited access to tissues. The laboratory mouse provides an ideal model to study aging in healthy individuals in a controlled environment. However, previous mouse studies have examined only a narrow range of the genetic variation that shapes individual differences during aging. Here, we analyzed transcriptome and proteome data from hearts of genetically diverse mice at ages 6, 12 and 18 months to characterize molecular changes that occur in the aging heart. Transcripts and proteins reveal distinct biological processes that are altered through the course of natural aging. Transcriptome analysis reveals a scenario of cardiac hypertrophy, fibrosis, and reemergence of fetal gene expression patterns. -
Supplementary Figures and Tables
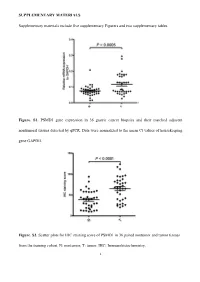
SUPPLEMENTARY MATERIALS Supplementary materials include five supplementary Figurers and two supplementary tables. Figure. S1. PSMD1 gene expression in 36 gastric cancer biopsies and their matched adjacent nontumoral tissues detected by qPCR. Data were normalized to the mean Ct values of housekeeping gene GAPDH. Figure. S2. Scatter plots for IHC staining score of PSMD1 in 36 paired nontumor and tumor tissues from the training cohort. N: nontumor; T: tumor. IHC: Immunohistochemistry. 1 Figure. S3. Kaplan-Meier survival curve analysis of DFS for all 411 patients with gastric cancer according to the PSMD1 expression stratified by clinicopathological risk factors. P-values were calculated by the log-rank test. 2 Figure. S4. Kaplan-Meier survival curve analysis of OS for all 411 patients with gastric cancer according to the PSMD1 expression stratified by clinicopathological risk factors. P-values were calculated by the log-rank test. 3 Figure. S5. Extended model for prediction of DFS and OS in patients with GC based on PSMD1 expression. ROC analyses of sensitivity and specificity for prediction of a, b DFS and c, d OS by the combined model (TNM stage and PSMD1 expression combination), TNM stage model, andPSMD1 expression model. 4 Figure. S6. X-tile plots of the DFS and OS nomograms. Color intensity of the plot represents low (dark, black) to high (bright, red, or green) strength of association at each division. Red represents the inverse association between the PSMD1 expression and survival (DFS or OS), whereas green represents direct association. a DFS nomogram, b OS nomogram. 5 Table S1. Clinical characteristics of the 36 patients with gastric cancer. -

A Peripheral Blood Gene Expression Signature to Diagnose Subclinical Acute Rejection
CLINICAL RESEARCH www.jasn.org A Peripheral Blood Gene Expression Signature to Diagnose Subclinical Acute Rejection Weijia Zhang,1 Zhengzi Yi,1 Karen L. Keung,2 Huimin Shang,3 Chengguo Wei,1 Paolo Cravedi,1 Zeguo Sun,1 Caixia Xi,1 Christopher Woytovich,1 Samira Farouk,1 Weiqing Huang,1 Khadija Banu,1 Lorenzo Gallon,4 Ciara N. Magee,5 Nader Najafian,5 Milagros Samaniego,6 Arjang Djamali ,7 Stephen I. Alexander,2 Ivy A. Rosales,8 Rex Neal Smith,8 Jenny Xiang,3 Evelyne Lerut,9 Dirk Kuypers,10,11 Maarten Naesens ,10,11 Philip J. O’Connell,2 Robert Colvin,8 Madhav C. Menon,1 and Barbara Murphy1 Due to the number of contributing authors, the affiliations are listed at the end of this article. ABSTRACT Background In kidney transplant recipients, surveillance biopsies can reveal, despite stable graft function, histologic features of acute rejection and borderline changes that are associated with undesirable graft outcomes. Noninvasive biomarkers of subclinical acute rejection are needed to avoid the risks and costs associated with repeated biopsies. Methods We examined subclinical histologic and functional changes in kidney transplant recipients from the prospective Genomics of Chronic Allograft Rejection (GoCAR) study who underwent surveillance biopsies over 2 years, identifying those with subclinical or borderline acute cellular rejection (ACR) at 3 months (ACR-3) post-transplant. We performed RNA sequencing on whole blood collected from 88 indi- viduals at the time of 3-month surveillance biopsy to identify transcripts associated with ACR-3, developed a novel sequencing-based targeted expression assay, and validated this gene signature in an independent cohort. -

The Human Genome Project
TO KNOW OURSELVES ❖ THE U.S. DEPARTMENT OF ENERGY AND THE HUMAN GENOME PROJECT JULY 1996 TO KNOW OURSELVES ❖ THE U.S. DEPARTMENT OF ENERGY AND THE HUMAN GENOME PROJECT JULY 1996 Contents FOREWORD . 2 THE GENOME PROJECT—WHY THE DOE? . 4 A bold but logical step INTRODUCING THE HUMAN GENOME . 6 The recipe for life Some definitions . 6 A plan of action . 8 EXPLORING THE GENOMIC LANDSCAPE . 10 Mapping the terrain Two giant steps: Chromosomes 16 and 19 . 12 Getting down to details: Sequencing the genome . 16 Shotguns and transposons . 20 How good is good enough? . 26 Sidebar: Tools of the Trade . 17 Sidebar: The Mighty Mouse . 24 BEYOND BIOLOGY . 27 Instrumentation and informatics Smaller is better—And other developments . 27 Dealing with the data . 30 ETHICAL, LEGAL, AND SOCIAL IMPLICATIONS . 32 An essential dimension of genome research Foreword T THE END OF THE ROAD in Little has been rapid, and it is now generally agreed Cottonwood Canyon, near Salt that this international project will produce Lake City, Alta is a place of the complete sequence of the human genome near-mythic renown among by the year 2005. A skiers. In time it may well And what is more important, the value assume similar status among molecular of the project also appears beyond doubt. geneticists. In December 1984, a conference Genome research is revolutionizing biology there, co-sponsored by the U.S. Department and biotechnology, and providing a vital of Energy, pondered a single question: Does thrust to the increasingly broad scope of the modern DNA research offer a way of detect- biological sciences.